Abstract
Osteoarthritis (OA) is a common joint disease affecting people worldwide, notably impacting quality of life due to joint pain and functional limitations. This study explores the potential of radiomics — quantitative image analysis combined with machine learning — to enhance knee OA diagnosis. Using a multimodal dataset of MRI and CT scans from 138 knees, radiomic features were extracted from cartilage segments. Machine learning algorithms were employed to classify degenerated and healthy knees based on radiomic features. Feature selection, guided by correlation and importance analyses, revealed texture and shape-related features as key predictors. Robustness analysis, assessing feature stability across segmentation variations, further refined feature selection. Results demonstrate high accuracy in knee OA classification using radiomics, showcasing its potential for early disease detection and personalized treatment approaches. This work contributes to advancing OA assessment and is part of the European SINPAIN project aimed at developing new OA therapies.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
1.1 Knee Osteoarthritis
Osteoarthritis (OA) is a prevalent joint disorder that significantly impacts individuals’ quality of life and poses challenges to the healthcare systems. It is a common condition, particularly affecting middle age and older people: a substantial percentage of individuals aged 60 and older is affected by it worldwide. OA leads to substantial disability, with symptoms like joint pain and stiffness affecting daily activities such as climbing stairs and walking, but also sleep disturbance due to night pain. In the knee, OA affects the three compartments of the knee joint (medial, lateral and patellofemoral) Lespasio et al. (2017) and usually develops slowly over 10 to 15 years, depending on its origin (Roos & Arden, 2016).
OA involves the breakdown of hyaline cartilage, as well as changes in the bone beneath it and the synovium (joint lining). Hyaline cartilage is crucial because of its ability to resist compression, primarily due to its unique structure and water-retaining properties. In OA, the collagen network within the cartilage deteriorates, leading to increased water content. This breakdown makes the cartilage less capable of withstanding mechanical stresses (Hussain et al., 2016; Heinegård & Saxne, 2011). As a result, the subchondral bone responds with new bone formation beneath the cartilage i.e. subchondral sclerosis and at the sides as bony spurs i.e osteophytes. Also, the synovial membrane becomes irritated and inflamed causing increased joint fluid production. Pain in OA significantly impacts quality of life and functional limitations. The pain experienced in OA is multifaceted, transitioning from intermittent weight-bearing pain to persistent chronic pain (Zhang & Jordan, 2010; Neogi, 2013). OA has a multifactorial etiology involving both systemic and local factors. Systemic risk factors include age (strongest risk factor for all joints), gender (women are more susceptible), race/ethnicity (varying prevalence among different groups), genetics (heritable component estimated at 50-65%), and hormonal influences (menopause-related changes in women). Local risk factors encompass obesity (particularly influential for knee OA) Coggon et al. (2001), joint injury or surgery, occupation (repetitive joint use), physical activity (elite sports activities may increase risk), mechanical factors (muscle weakness, joint misalignment, ligament laxity), and limb length inequality (Roos & Arden, 2016; Murphy et al., 2008; Zhang & Jordan, 2010; Neogi, 2013).
1.2 Knee OA Diagnosis
The main diagnostic tools for the assessment of human joints are X-ray, computed tomography (CT), and more commonly also magnetic resonance imaging (MRI). The severity of OA can be evaluated by assessing joint space narrowing and damage to cartilage and underlying bone. Various scales are available for this purpose. The Kellgren–Lawrence (KL) grading system is used to rate OA on plain X-rays. The definite presence of an osteophyte (Kellgren–Lawrence grade 2) confirms a structural diagnosis of OA. However, this system combines an overall grade for OA from joint space narrowing and the presence of osteophytes, assuming these structural changes appear continuously (Roemer et al., 2020; Kellgren et al., 1957). Other grading systems, like the OA Research Society International (OARSI) Atlas system (Roemer et al., 2020), separate joint space narrowing from osteophyte presence but focus solely on the tibiofemoral joint, which underestimates the patellofemoral contribution to the disease. The Ahlbäck scale, developed in 1968, is based on joint space narrowing measurements (Ahlback, 1968), but it presents challenges in its application due to variations influenced by observer experience, age, and personal attitudes. This scale also highlights difficulties in distinguishing between low and moderate grades of arthritis (Galli et al., 2003).
MRI has become a critical tool in diagnosing, especially in research settings, due to its ability to detect detailed soft-tissue and structural changes not visible on conventional X-rays. Semi-quantitative MRI-based grading systems, such as the Whole Organ Magnetic Resonance Imaging Score (WORMS) and Knee OA Scoring System (KOSS), consider various features of the MR image from the entire knee joint, offering increased sensitivity compared to traditional grading systems. However, interpreting pre-radiographic MRI findings remains challenging, as distinguishing between normal age-related changes and predictive signs of knee OA progression is not straightforward in clinical practice (Demehri & Shakoor, 2018).
Treatment modalities for knee OA include physical therapy, medication, injections, osteotomies and, in the most severe cases, partial or total knee replacement operation (Ringdahl & Pandit, 2011). Therefore, early diagnosis is crucial to timely intervention and potentially avoiding the need for surgery. Assessing the severity of osteoarthritis presents challenges because of the various classification systems available, each with its own limitations. Early diagnosis remains critical, given the progressive nature of osteoarthritis, as it allows early intervention with less invasive treatments such as physiotherapy and drug therapy, potentially reducing the need for surgery. Therefore, further advances in imaging technology and diagnostic criteria are essential to optimize the management and outcomes of patients with knee osteoarthritis.
1.3 Use of Radiomics in Medical Imaging
Radiomics involves the extraction of quantitative metrics known as radiomic features from medical images to capture tissue properties. It can reveal quantitative information and properties often imperceptible to the human eye, that can result to be relevant in clinical analysis (combined with various types of data) and can offer valuable insights into the characteristics and progression of diseases. The impact of radiomics can be enhanced by the use of machine learning: integration of artificial intelligence, machine learning, and advanced image analysis techniques to discern meaningful information from medical images. By integrating radiomic data with clinical, genomic and biological information, researchers can unravel complex disease pathways and refine patient stratification strategies (Mayerhoefer et al., 2020). Radiomics has been used for a large number of medical purposes, such as the detection of disease risk factors (Angelone et al., 2022, 2023; Sansone et al., 2023; Cetin et al., 2020), detection of genomic patterns (Liu et al., 2023; Saxena et al., 2022), the detection of soft tissue masses, bone tumors and calcifications (Eslami et al., 2020; Lei et al., 2019) for various diagnostic imaging techniques. Radiomic features extracted from CT scans and analyzed using machine learning (ML) techniques have proven to be valid and robust across various oncological diagnostic applications, including liver tumors (Sánchez et al., 2021) and head and neck cancer (Parmar et al., 2015a). A study focusing on lung cancer emphasized the necessity of identifying optimal machine-learning methods for radiomic applications to generate stable and clinically relevant radiomic biomarkers applicable in clinical practice (Parmar et al., 2015b).
To date, radiomics has not been extensively used in the field of knee OA. Jiang et al. (2024) discuss how radiomics can enhance accurate OA diagnosis by converting medical images into quantitative data. The study highlights the limitations of current imaging techniques like X-ray, CT, and MRI in early OA detection and stresses the importance of radiomics in addressing this issue. Furthermore, it reviews current research on radiomics in OA, detailing studies that classify, detect, and predict OA in various joints (e.g., knee, hip, hand, spine). Radiomics offers advantages over traditional clinical variables in detecting specific forms of OA and shows promise in improving diagnostic accuracy and prognosis prediction. In Xie et al. (2021), they extracted radiomic features from T2-weighted MRI images of subjects with post-traumatic osteoarthritis and a control group, focusing on subchondral bone and femoral and tibial cartilage. Radiomic analysis revealed a distinctive signature in the cartilage and subchondral bone of various compartments, showing excellent performance in distinguishing between patients and controls. The articles (Xue et al., 2022) and Hirvasniemi et al. (2021) explore the use of MRI-based radiomics approaches for diagnosing knee OA by analyzing subchondral bone. Both approaches achieve high performance in distinguishing between OA subjects and healthy control class.
1.4 Objectives
The primary objective of this work is to explore a novel approach using radiomic analysis applied to CT and MRI imaging modalities to assess cartilage degradation in knee OA. Cartilage degradation is a hallmark of OA and has a significant impact on patients’ quality of life. Traditional diagnostic approaches often provide limited insight into early-stage changes. By combining CT and MRI modalities and using radiomics, a technique whose application in cartilage studies is limited, we seek to establish a robust methodology for in-depth evaluation of cartilage. Our goal is to employ radiomics, which involves the extraction of quantitative image features, to identify reliable biomarkers for early detection and characterization of OA-related cartilage changes. In addition, we seek to evaluate the performance and robustness of radiomic features under different imaging conditions, highlighting the potential of this approach to improve diagnostic accuracy and inform personalized treatment strategies for patients with OA.
2 Methods
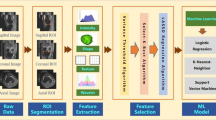
The workflow followed in the present study is shown in Fig. 1.
2.1 Subjects Recruitment
For this study, patients diagnosed with OA and healthy individuals were recruited for knee scanning. The recruitment is part of the EU projects RESTOREFootnote 1 and SINPAINFootnote 2, aimed at developing new therapies for cartilage regeneration. The RESTORE project ended in 2022, while SINPAIN will end in 2026.
Approval for the study was obtained from the Icelandic Bioethics Commission (approval number: VSN-19-050V1) in March 2022. Prior to any scans, subjects provided comprehensive written consent, ensuring transparency and accountability. To ensure patient privacy, all data are anonymized and securely stored on a dedicated hospital server approved by the Data Protection Unit.
Led by the chief of Orthopedics at Landspitali University Hospital in Reykjavik, Iceland, subject recruitment adheres to specific criteria: 1) Subjects with degenerative cartilage (D), evaluated clinically with Ahlback grades between 2 and 4, awaiting total knee arthroplasty. 2) Healthy subjects (C) with no prior medical history of knee cartilage diseases or trauma.
Scans, including CT, MRI and X-rays, are performed at Landspitali University Hospital in Reykjavik, Iceland. Standardized protocols and positioning with a splint are used. The Toshiba Aquillion One CT scanner (320 slice) covers an area of 16 cm2, while the Siemens Healthcare Prism MRI scanner (3T) uses 3D volumetric sequences with 0.6-mm isotropic voxels.
2.2 Dataset Description
A total of 138 knees of 95 subjects were scanned; about half of the cohort scanned both their right and left legs in two consecutive acquisitions. Of the 138 knees, 53 belong to class D, while 84 belong to class C. Information about subjects’ gender and age - average (avg) and standard deviation(std) - are reported in Table 1.
The study centered on the examination of knee cartilages, including the femoral, lateral tibial, medial tibial, and patellar cartilages. Figure 2 shows a three-dimensional representation of the four different cartilage segments for a degenerative and a control subject for both left and right legs. It is important to note that half of the knees represent the right and left knees of the same patient. This aspect is taken into consideration in subsequent analyses.
2.3 Cartilage Segmentation and Registration
The images were processed using Mimics (Materialise’s Interactive Medical Image Control System), a medical imaging software specialized in generating 3D models from medical data. By loading a stack of images in the XY plane, Mimics produced images in the XZ and YZ directions, offering a comprehensive three-dimensional representation. To ensure accuracy, both cartilages and bones were subjected to segmentation, a process for identifying and delineating these structures within the images. Cartilages were segmented from MRI images due to MRI’s suitability for soft tissue examination, while bones were segmented from CT images, following a uniform protocol. The segmentation process involved creating masks to isolate specific elements based on density thresholds. Editing operations, such as cropping, refined the masks, and region-growing isolated parts by removing unconnected pixels. This meticulous segmentation aimed to extract meaningful and consistent information from the images. The resulting masks were then converted into highly accurate 3D objects with Mimics. In the registration process, 3D objects from different CT and MRI masks were combined to reconstruct a realistic 3D model of the knee compartment. MRI data were aligned with CT data through image registration, facilitated by acquiring images at the exact same location. Four landmarks were taken on the anatomical parts to ensure accurate overlap. The newly created image integrated the CT data with the segmented bones and cartilage from the MRI. Manual realignment of the segmented parts ensured accurate positioning. To eliminate overlaps, refinements were performed on the masks, such as Boolean subtraction operations. The final 3D model provided a realistic overview of the knee compartment. The same methodology has already been used in Ciliberti et al. (2022) and Aubonnet et al. (2023), affirming the relevance and reliability of the proposed approach. Hence, we now have segmented cartilage from both MRI, where it was initially delineated, and CT. This allows us to conduct a comparative analysis and perform evaluations of radiomics features on the two separate imaging modalities.
2.4 Radiomics Features Extraction
Radiomics features were extracted using the PyRadiomics library (Van Griethuysen et al., 2017) based on cartilage segmentations obtained from both MRI and CT, as previously described in Section 2.3. The extracted features describe shape, pixel statistics based on gray value histograms and texture and can be summarized as follows:
-
First Order Statistics (19 features)
-
Shape-based (3D) (16 features)
-
Shape-based (2D) (10 features)
-
Gray Level Co-occurrence Matrix (GLCM) (24 features)
-
Gray Level Run Length Matrix (GLRLM) (16 features)
-
Gray Level Zone Size Matrix (GLSZM) (16 features)
-
Neighbouring Gray Tone Difference Matrix (NGTDM) (5 features)
-
Gray Level Dependence Matrix (GLDM) (14 features)
In addition to the original image, the described features are also calculated for the wavelet images. The wavelet transform can be used to decompose the original image into different frequencies or scales. This allows textures and other information within specific frequency bands to be examined. This could potentially improve the ability to distinguish between different conditions. Then a total of 875 features for each of the four cartilage considered were extracted.
2.5 Standardization and Features Selection
Once the radiomic features were extracted, a first phase of feature normalization was carried out. Given the high number of features extracted compared to the number of samples considered, an accurate feature selection is a fundamental step, improving computational efficiency, model performance, and avoiding overfitting. In this work, the correlation matrix, a statistical approach to quantify the linear correlation between variables, was first evaluated using the Pearson correlation coefficient (Cohen et al., 2009), excluding all features with a correlation greater than 90%. Afterward, a feature importance analysis was performed by selecting only the most important features for the classification.
2.6 Robustness Analysis
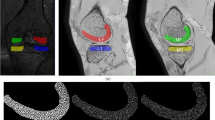
Following the statistical approach proposed in Ponsiglione et al. (2023), a robustness analysis of the radiomics features was also conducted, to improve the reliability of the analysis. In particular, the variability and robustness of radiomics features as a function of cartilage segmentation both in MRI and CT images was investigated, varying the shape of the segmentation through morphological operators and simulating an erosion or dilation of the area. The Fig. 3 shows a visual representation of the effects of morphological operations, specifically dilation and erosion, on segmented cartilages. In Fig. 3 each panel demonstrates the effect of different levels of dilation and erosion on what could be interpreted as variation in segmenting biological structures. Starting from the top, the first red block shows the effect of dilation, expanding the boundaries of the ROI from 5 to 1 pixels, showing progressively lower levels of dilation. The middle row labeled ’Original’ represents the baseline. Following, in the blue block, there are the effects of erosion from 1 to 5 pixels. Such manipulations are crucial for exploring the robustness of radiomics features since changing the shape and size of ROIs reflects intra- and inter-operator variability in segmentation, which is common in practical clinical applications.
Following these alterations, a full extraction of radiomic features is performed on both the original and modified ROIs. Then, a Shapiro-Wilk test (\(\alpha \) = 0.05) was performed to evaluate the normality of the distribution of radiomic features, and a t-test or Mann-Whitney statistical test was performed (depending on the normality result) to compare the distributions of radiomic features between original ROI and dilated ROI and between original ROI and eroded ROI. Based on the p-value obtained, a robustness score in the range between 0 and 1 was obtained, as detailed in Ponsiglione et al. (2023). Robustness score values closer to 1 indicate greater robustness of the feature classes examined.
Boxplots and trend lines were represented to see how the robustness score varies as the amplitude of the erosion or dilation varies.
An innovative method was then developed to integrate robustness into the calculation of feature importance for ML models. This method prioritizes features not only based on model significance but also on their stability amid data variations, such as those introduced by noise or changes to ROI segmentation.
Typically, feature importance in ML is determined by each feature’s contribution to the model’s predictive accuracy, but this traditional approach often overlooks the stability or robustness of features to fluctuations in the input data, especially when the sample of data is not very large. In this new methodology, robustness is quantified using the negative logarithm of the average p-value of the robustness score, which inversely correlates a feature’s weight with its robustness: a lower p-value, suggesting a lower likelihood that the feature’s robustness is random, results in a higher weight, and the negative logarithm transformation scales the p-values to a more understandable range, where smaller p-values (indicating stronger evidence against the null hypothesis and affirming the robustness of the feature) produce larger weights. By multiplying the original feature importance by this robustness weight, importance scores are recalibrated to favor features that are both predictive and robust. Features with lower robustness (higher p-value) suffer a penalty, decreasing their modified importance score. The reweighted system may reveal that some features previously perceived as crucial are less important when robustness is taken into account, while others gain importance due to their enhanced robustness, suggesting a more reliable set of predictors.
Statistical tests were carried out in Python using the SciPy library (Virtanen et al., 2020).
2.7 Machine Learning Analysis
Based on the most significant features calculated, a first ML analysis was conducted for the classification of D and C subjects, considering all features extracted from all cartilages. The following algorithms have been used: Random Forest, Linear Support Vector Machine (SVM), Logistic Regression, Gradient Boosting, Linear Perceptron. In this context, particularly when dealing with limited data samples, research to identify optimal models and their corresponding hyperparameters is of paramount importance. A prevalent method to achieve this is through the use of grid search, meticulously coupled with cross-validation (CV) techniques. In our work we used a "nested grid search" coupled with a K-fold CV. This approach involves an initial layer of external stratified cross-validation which is crucial as it maintains class balance when partitioning the dataset, thus providing a more representative cross-sectional framework for evaluation. During each iteration of this external cross-validation, an internal grid search is conducted, accompanied by its cross-validation process, since the nested structure serves a dual purpose: it not only tunes the hyperparameters but also offers a more objective estimate of the model’s performance on unseen data. The algorithms tested and the hyperparameters tuned are shown in Table 2. To ensure comprehensive evaluation, both the mean and standard deviation were considered in all cases, this approach produces a more generalized and reliable performance metric, effectively mitigating the risks associated with overfitting, particularly on the validation set.
By constantly rotating the validation set in each external iteration, the method ensures a robust and unbiased evaluation, ultimately leading to more reliable and generalizable model performance.
All the ML analyses were performed with the Scikit-Learn library in Python (Kramer & Kramer, 2016).
Figure 4 shows the steps of the workflow from feature extraction to the final classification.
2.8 Evaluation Metrics
The following metrics were used to evaluate the performances of trained models:
-
Accuracy: This is the percentage of correct predictions out of the total number of predictions. It is calculated as the number of correct predictions divided by the total number of predictions:
$$ \text {Accuracy} = \frac{\textrm {CorrPred}}{\textrm {TotPred}} $$ -
Precision: Represents the proportion of positive identifications that are actually correct. It is calculated as the number of true positives divided by the sum of true positives and false positives:
$$ \text {Precision} = \frac{\textrm {True positives (TP)}}{\textrm {True positives (TP)} + \textrm{False positives (FP)}} $$ -
Recall (Sensitivity): Represents the proportion of positive identifications that are actually correct. It is calculated as the number of true positives divided by the sum of true positives and false positives:
$$ \text {Recall} = \frac{\text {True positives (TP)}}{\textrm {True positives (TP)} + \textrm{False negatives (FN)}} $$ -
F1-Score: This is the harmonic mean of precision and recall, this score takes into account both precision and recall to provide a single measure of model performance. It is especially useful when you want a balance between precision and recall and when dealing with unbalanced datasets:
$$ \text {F1 score} = 2 \times \frac{\text {Precision} \times \text {Recall}}{\textrm {Precision} + \textrm {Recall}} $$
3 Results
3.1 Binary Classification Control-Degenerative
3.1.1 Binary classification using the least correlated radiomic features
The correlation matrix was calculated as the first feature selection method to eliminate all highly correlated features, especially with a linear correlation greater than 90%. It is shown in Fig. 5 and allowed us to reduce the dataset from 3500 initial features, relating to all four cartilages, to 1272 features in MRI imaging and 958 features in CT imaging. Using a 10-fold nested CV, as explained in Section 2.7, the evaluation metrics of four different ML algorithms are shown in Table 3 for MRI imaging and in Table 4 for CT imaging. As can be seen, the best performances in the classification between D and C patients can be obtained with the SVM Linear algorithm for the radiomic features extracted from the MRI, obtaining an average accuracy of 90.25%, and with the logistic regression for the radiomic features extracted from the CT, obtaining an average accuracy of 91.33%.
3.1.2 Binary Classification Using the least correlated Radiomic Features after Feature Importance
The set of features considered for model training was further reduced by also carrying out a feature importance analysis following the correlation matrix. In Fig. 6a it is possible to observe that the first most important features for MRI, using SVM algorithm, mainly refer to the patellar cartilage and are mainly belonging to the shape and GLSZM classes. For the CT images, using the Logistic Regression algorithm, a pattern of features is not so evident, in fact, there is an almost similar distribution between the patellar, femoral, and tibial cartilage, and mainly shape, GLSZM and GLCM features appear, as can be seen in Fig. 6b. In Fig. 7 the distribution of the 50 most important features is shown based on the classes of radiomic features per cartilage area considered, both for CT and MRI images.
The analysis reveals that among the most important features are those related to texture matrices, particularly those derived from the wavelet transform, and those related to shape. This underlines the relevance of specific texture patterns and morphological aspects in the analysed cartilage for the clinical conditions of interest.
The predominance of texture features derived from the wavelet transform may reflect their ability to describe the complexity of cartilage tissue at different scales of analysis, which is essential for identifying subtle but clinically relevant patterns. Furthermore, the importance of shape-related features highlights how the shape and geometric structure of cartilage may play a critical role in the assessment of joint health. By training the models with the first 50 most important features we obtain the results shown in Tables 5 and 6. The performances, as expected, improved, reaching very high results for all the evaluation metrics considered, with an average of around 99% for all in the MRI and 96% in the CT.
3.1.3 Robustness Analysis Results
To make the analysis even more reliable and robust, the robustness of the radiomic features was evaluated with respect to shape variations in the segmentation, as described in Section 2.6. Therefore, after introducing a controlled noise by dilating and eroding the edges of the segmentation and extracting the radiomic features for each dilated, eroded and original ROI, a robustness score was produced based on the p-value resulting from the Mann-Whitney test. In particular, boxplots were produced for variations of pixels from 1 to 5 which facilitate robustness comparisons between classes of features described in Section 2.3. In particular, in Figs. 8 and 9 it is possible to observe the boxplots of robustness score for the segmentations with edge dilation from 1 to 5 pixels, respectively for MRI and CT. In Figs. 10 and 11, however, the boxplots of robustness score for the segmentations with edge erosion from 1 to 5 pixels are shown. For both erosion and dilation, considering that a robustness score closer to 1 indicates greater robustness of the feature classes, it is possible to observe that the robustness decreases as the degree of increase of the morphological operation increases. This is an expected result because greater alterations of the ROI determine a higher variability in the extracted radiomic features. However, it is interesting to note how erosion is more affected by this phenomenon. Observing Figs. 10 and 11, for an alteration of 5 pixels, the robustness score drops much more than the analog in Figs. 8 and 9. Trend lines, shown in Fig. 12, have also been graphed to show the trend of the robustness score as the degree of variation in pixels in erosion/dilation varies. In these plots, a more robust feature class would show a flatter trend line, indicating that its robustness score is less affected by changes in ROI.
As can be seen from the boxplots and trend plots, the first-order features and the GLCM features remain more robust for all dilation amplitudes, with greater robustness for small variations. In the case of erosion, however, we can observe greater robustness in the case of first-order, NGDTM and GLCM features. By combining the robustness analysis in the feature importance analysis as described in Section 2.6, we obtain new weighted feature importance scores of which the best 50 are shown in the barplots in Fig. 13.
By retraining the ML algorithms with these new more important and robust features, we obtain the metrics summarized in the Tables 7 and 8. As expected, the tables show lower results than previous performances, in which the importance of features was evaluated solely on the basis of meticulously executed segmentation data. However, moving away from an ideal situation and considering the great variability of results during the segmentation process, having more generalizable and more robust features becomes crucial, despite lower but still optimal results.
4 Discussion
The present work shows a classification of knees with cartilage degradation compared to a control group, using radiomics features and ML algorithms with a multimodal approach. In the first instance, a classic ML approach was followed, paying attention to an accurate subdivision of the dataset for training and an accurate selection of features. Analyzing medical imaging data, we find a striking pattern: about half of patients have images for both their right and left knees. Despite the natural differences between these two joints, there is remarkable symmetry in their anatomy and bone structures, leading to parallel formations in the cartilage. This symmetry becomes even more pronounced in patients suffering from degenerative conditions, where similar patterns of cartilage degradation are observable in both knees. While this is not a universal trend across all samples, it has critical implications for the ML application in medical imaging. The main challenge is avoiding overfitting, which occurs when a model, after being trained on a specific dataset, becomes over-adapted to its peculiarities, thus losing its predictive power on new and invisible data. In this work, a great effort was made to make the model as reliable and generalizable as possible. There is a risk that the model inadvertently learns to recognize knee-specific images, particularly if images of both knees from the same patient are included in separate training and testing subsets. To mitigate this problem, a refined approach to data preparation is essential, so knees from the same patient were grouped in the same fold of the dataset. This method ensures that the model’s learning process remains uncontaminated by patient-specific traits, without bias. The correlation between the features was then evaluated and a feature importance analysis was subsequently carried out to select the non-redundant and most predictive features for the ML models.
The predominance of texture features derived from the wavelet transform may reflect their ability to describe the complexity of cartilage tissue at different scales of analysis, which is essential for identifying subtle but clinically relevant patterns. Furthermore, the importance of shape-related features highlights how the shape and geometric structure of cartilage may play a critical role in the assessment of joint health. Features related to wavelet texture matrices capture information regarding the variation of pixel intensity levels across different scales and orientations. This can be clinically interpreted as a measure of the heterogeneity of cartilage composition and internal structure, potentially reflecting microscopic changes due to pathological processes such as osteoarthritis. The presence of particular texture patterns may indicate changes in cartilage composition that precede visible morphological changes. Whereas, shape features are representative of cartilage geometry and size. These may reflect structural changes in cartilage, such as thickening, deformation or volume reduction, which are hallmarks of degenerative disease or injury. It is known that OA cartilages present changes in composition due to increased water content and reduction in collagen and proteoglycan network (Cooke et al., 2018; Danalache et al., 2019; Saarakkala et al., 2010; Wu et al., 2019). This would explain both the structural changes and the texture differences in our cohort. Cartilage thickness tends to decrease in OA subjects (Aubonnet et al., 2023; Kubakaddi et al., 2013). In clinical terms, the importance of these features supports the idea that radiomics may serve as a powerful non-invasive tool for early detection of joint disease and monitoring disease progression. In particular, greater variability in texture features might be associated with early stages of cartilage damage, while significant alterations in shape features might indicate a more advanced condition. However, to further improve the generalizability of the models, a robustness analysis was introduced, following the statistical approach proposed by Ponsiglione et al. (2023) and developing an innovative method to take into account the robustness of features in the features selection process.
The objective of the robustness analysis is to enhance the reliability of radiomic features by assessing their stability against segmentation variations in MRI and CT images. Indeed, segmentation is affected by observer variability: individual observers (either within the same observer over time - intra-observer, or between different observers - inter-observer) may interpret and delineate structures in medical images in different ways. Assessing robustness against such variability is critical to ensuring the reliability of radiometric features in clinical applications. This was achieved by evaluating how changes in segmentation, induced by morphological operations such as dilation and erosion, affect the stability of radiomic features. Robustness scores were calculated using statistical tests and integrated into the feature importance calculation for machine learning models. The findings reveal that first-order and GLCM features are the most robust, whereas shape features are highly sensitive to segmentation changes. Ideally, robustness is indicated by a p-value greater than 0.5 across all morphological changes (from 1 to 5 pixels). Figures 8 and 10 illustrate this trend for GLCM and first-order features under both erosion and dilation on MRI. Figures 9 and 11 show the same positive trend for first-order, NGTDM and GLDM features on CT. Other features exhibit a different trend, with p-values decreasing to 0 as the dilation/erosion pixels increase.
More specifically, all feature classes show a downward trend, indicating a decrease in robustness scores as the degree of morphological operation increases. This is expected since more extensive morphological alterations can lead to greater changes in the features extracted from the altered ROIs. Initial robustness scores to 1-pixel change provide information on the intrinsic stability of each feature class to slight morphological changes.
Based on the trend plots and the previous discussed boxplots, the first-order and GLCM feature classes appear to be the most robust in the analysis of dilation and erosion. This would indicate that these classes maintain their feature values more consistently despite morphological changes applied to the images. First-order features, which describe the distribution of individual pixel values without considering spatial relationships (such as energy, entropy, or average pixel intensity), may be inherently less sensitive to minor changes in ROI delineations because they capture the global statistics of the ROI rather than local models. This may explain why they appear to remain more stable across different levels of morphological operations. GLCM features, on the other hand, are based on the frequency of pixel values occurring at a certain distance and orientation within the image, which describes the texture, this suggests that the structural information they encapsulate is preserved even when the ROI is modified. This could mean that the texture patterns are sufficiently distinct and not easily disrupted by ROI dilation. In particular, a strong difference can be observed between the robustness scores of dilation and erosion; in fact, features remain more robust for dilation. In general, this is easily explained by the fact that, compared to dilation, in erosion, information is removed by reducing the region of interest. In particular, with very high erosion (3 to 5 pixels), as shown in Fig. 3, the features change significantly, affecting the results of the analysis. These features are likely to provide reliable and consistent information for the diagnosis of conditions such as osteoarthritis in CT and MRI scans, despite variations in image segmentation. From the boxplots in Figs. 8, 10, 9, and 11, it appears that first-order features are generally the most robust against segmentation variations. This robustness suggests that pixel intensity values within the ROI are reliable despite small changes in segmentation, while texture features are more sensitive to changes in segmentation. Interestingly, some texture features, such as GLCM and NGDTM, hold up better than others, such as GLRLM and GLSZM, when these small changes occur. In contrast, shape features are much more sensitive to segmentation, meaning that inaccurate segmentation could significantly affect the results. As expansion and erosion increase from 1 to 5 pixels, the variance of robustness of almost all features increases. This indicates higher sensitivity and potentially lower reliability under conditions of more significant variation in ROIs.
Given the proposed multimodal approach, particular considerations can also be made on the differences between the classification on MRI and CT images. For a purely visual evaluation by a clinician, diagnoses based on CT only consider space between joints and bony structures, but it does not allow clear visualization of the cartilaginous structure. From our study, however, it emerges that with radiomic techniques the classification of degenerated knees is not only possible but also very efficient from CT, almost comparable to the results obtained with radiomic features extracted from MRI. This gives space to new perspectives.
Comparison of our results with the existing literature is challenging for several reasons. As previously underlined, existing studies on using radiomics to predict knee degeneration are limited. Among them, most of the studies either focused on different ROIs compared to ours or only considered MRI data. Xie et al. (2021) proposed a methodology that used combined radiomics models for the cartilage and subchondral bone regions across various knee compartments (lateral femur, lateral tibia, medial femur, and medial tibia) on T2 mapping images. Their approach demonstrated excellent performance in distinguishing post-traumatic OA patients from healthy controls. However, this study focused exclusively on patients predisposed to post-traumatic OA after anterior cruciate ligament reconstruction, whereas our cohort does not specifically include or differentiate subjects based on post-traumatic OA status. In Xue et al. (2022) the authors aimed to develop an MRI-based radiomics predictive model for identifying knee OA by analyzing the tibial and femoral subchondral bone. Radiomic features were extracted from different regions of the lateral and medial compartments of the femur condyle and tibial plateau. The model reached an area under the curve (AUC) score of 0.961. In Hirvasniemi et al. (2021) the authors assessed tibiofemoral OA extracting MRI–based radiomic features (in particular, shape and texture features) from proximal tibia ROI, reaching an AUC score of 0.80. Cui et al. (2023) extracted radiomic features from knee cartilage portion manually segmented on MRI. The considered ROI were the medial and lateral compartments of the tibiofemoral joints and the patellofemoral joints. The accuracy of ML model on the training set reached 94% of accuracy.
In contrast to these studies, our innovative approach incorporates segmented knee cartilage on both CT and MRI, achieving high results, particularly on MRI features. The precise segmentation process on MRI and subsequent accurate registration on CT resulted in a unique multimodal dataset. This approach facilitates new comparisons and represents a significant advance in the field. It is the first study available to extract radiomic features of cartilage from CT and the first to compare the two imaging modalities for predicting knee OA after a precise segmentation process. This paves the way for future research in the field of radiomics, exploring the combined use of features extracted from CT and MRI.
4.1 Limitations
Some limitations should be acknowledged. Firstly, the imbalance between the classes C and D within the dataset may have influenced the performance of the machine learning algorithms. Class imbalance can lead to biased model training, where the classifier tends to favor the majority class (in this case, the healthy class), potentially resulting in reduced sensitivity to detect the minority class (patients with knee osteoarthritis). Strategies such as oversampling the minority class or adjusting class weights during model training could be explored to address this issue in future studies.
Secondly, the quality of the medical imaging data, particularly MRI and CT scans, is crucial for accurate radiomics analysis. Variations in image resolution, noise levels, and artifacts can impact the extraction and reliability of radiomic features. Ensuring standardized imaging protocols and rigorous quality control measures during image acquisition can help mitigate these challenges.
Furthermore, the study’s limited sample size and the number of subjects pose potential limitations. With fewer subjects, there is a risk of overfitting the machine learning models to the specific characteristics of the dataset, which may limit the generalizability of the findings to broader patient populations. Increasing the sample size and subjects’ diversity could enhance future investigations’ robustness and external validity.
Lastly, the segmentation and registration processes are performed manually, which is time-consuming. Although manual segmentation offers advantages, particularly given the non-homogeneous shapes and varying morphology of osteoarthritic cartilage across patients, developing an automated process that equals the accuracy of manual segmentation is essential, albeit challenging. Optimizing and accelerating these initial steps are crucial for providing quicker and more accessible results for patients undergoing knee scans. Such advancements will enhance the efficiency of early diagnosis and improve patient outcomes.
5 Conclusion
This study presents a novel approach by integrating CT and MRI data and applying radiomics, a relatively unexplored technique in the literature on cartilage evaluation. The demonstration of consistent results between the two imaging modalities underscores the efficacy and potential of this new methodology in the evaluation of cartilage degeneration, contributing to the early diagnosis of the disease. In addition, the robustness analysis applied in our study provides further validation of the reliability and consistency of our results. The generalization of the results gives room for new perspectives in diagnosis. This work represents a significant step forward in the use of radiomics for cartilage assessment, demonstrating that it is a promising tool in clinical practice, in line with the goal of the European SINPAIN project (https://www.osteoarthritis-sinpain.eu/), which aims to develop new therapies for the treatment of knee OA.
Availability of data and material
An online database with some of the data used and processed for this study is available at the following website: https://restore-project.ru.is/, accessed on 30/04/2024. The database will contain the full cohort of used data by the year 2025.
Notes
https://restoreproject.eu/ - Grant agreement ID: 814558
https://www.osteoarthritis-sinpain.eu/ - Grant agreement ID: 101057778
References
Ahlback, S. (1968). Osteoarthrosis of the knee. a radiographic investigation. Acta Radiol Diagn (Suppl), 277, 7–72.
Angelone, F., Ricciardi, C., Gatta, G., Sansone, M., Ponsiglione, A. M., Belfiore, M. P., Amato, F., & Grassi, R. (2022). Breast density analysis on mammograms: Application of machine learning with textural features. In: 2022 IEEE International Conference on Metrology for Extended Reality, Artificial Intelligence and Neural Engineering (MetroXRAINE), pp. 295–300 (2022). IEEE
Angelone, F., Ponsiglione, A. M., Belfiore, M. P., Gatta, G., Grassi, R., Amato, F., & Sansone, M. (2023). 2023. Evaluation of breast density variability between right and left breasts. In: Atti del Convegno Nazionale di Bioingegneria.
Aubonnet, R., Ramos, J., Recenti, M., Jacob, D., Ciliberti, F., Guerrini, L., Gislason, M. K., Sigurjonsson, O., Tsirilaki, M., Jónsson, H., Jr., amp, et al. (2023). Toward new assessment of knee cartilage degeneration. Cartilage,14(3), 351–374.
Cetin, I., Raisi-Estabragh, Z., Petersen, S. E., Napel, S., Piechnik, S. K., Neubauer, S., Gonzalez Ballester, M. A., Camara, O., & Lekadir, K. (2020). Radiomics signatures of cardiovascular risk factors in cardiac mri: results from the uk biobank. Frontiers in Cardiovascular Medicine, 7, 591368.
Ciliberti, F. K., Guerrini, L., Gunnarsson, A. E., Recenti, M., Jacob, D., Cangiano, V., Tesfahunegn, Y. A., Islind, A. S., Tortorella, F., Tsirilaki, M., amp, et al. (2022). Ct-and mri-based 3d reconstruction of knee joint to assess cartilage and bone. Diagnostics,12(2), 279.
Coggon, D., Reading, I., Croft, P., McLaren, M., Barrett, D., & Cooper, C. (2001). Knee osteoarthritis and obesity. International Journal of Obesity, 25(5), 622–627.
Cohen, I., Huang, Y., Chen, J., Benesty, J., Benesty, J., Chen, J., Huang, Y., & Cohen, I. (2009). Pearson correlation coefficient. Noise reduction in speech processing, 1–4
Cooke, M., Lawless, B., Jones, S. W., & Grover, L. (2018). Matrix degradation in osteoarthritis primes the superficial region of cartilage for mechanical damage. Acta Biomaterialia, 78, 320–328. https://doi.org/10.1016/j.actbio.2018.07.037
Cui, T., Liu, R., Jing, Y., Fu, J., & Chen, J. (2023). Development of machine learning models aiming at knee osteoarthritis diagnosing: an mri radiomics analysis. Journal of Orthopaedic Surgery and Research, 18(1), 375.
Danalache, M., Kleinert, R., Schneider, J., Erler, A. L., Schwitalle, M., Riester, R., Traub, F., & Hofmann, U. K. (2019). Changes in stiffness and biochemical composition of the pericellular matrix as a function of spatial chondrocyte organisation in osteoarthritic cartilage. Osteoarthritis and Cartilage, 27(5), 823–832. https://doi.org/10.1016/j.joca.2019.01.008
Demehri, S., & Shakoor, D. (2018). Structural changes in aging-knee vs early-knee osteoarthritis: review of current evidence and future challenges. Osteoarthritis and Cartilage, 26(11), 1412–1414.
Eslami, P., Parmar, C., Foldyna, B., Scholtz, J. -E., Ivanov, A., Zeleznik, R., Lu, M. T., Ferencik, M., Vasan, R. S., Baltrusaitis, K., & et al. (2020). Radiomics of coronary artery calcium in the framingham heart study. Radiology: Cardiothoracic Imaging, 2(1), 190119
Galli, M., De Santis, V., & Tafuro, L. (2003). Reliability of the ahlbäck classification of knee osteoarthritis. Osteoarthritis and Cartilage, 11(8), 580–584. https://doi.org/10.1016/S1063-4584(03)00095-5
Heinegård, D., & Saxne, T. (2011). The role of the cartilage matrix in osteoarthritis. Nature Reviews Rheumatology, 7(1), 50–56.
Hirvasniemi, J., Klein, S., Bierma-Zeinstra, S., Vernooij, M. W., Schiphof, D., & Oei, E. H. (2021). A machine learning approach to distinguish between knees without and with osteoarthritis using mri-based radiomic features from tibial bone. European Radiology, 31, 8513–8521.
Hussain, S., Neilly, D., Baliga, S., Patil, S., & Meek, R. (2016). Knee osteoarthritis: a review of management options. Scottish Medical Journal, 61(1), 7–16.
Jiang, T., Lau, S.-H., Zhang, J., Chan, L.-C., Wang, W., Chan, P.-K., Cai, J., & Wen, C. (2024). Radiomics signature of osteoarthritis: Current status and perspective. Journal of Orthopaedic Translation, 45, 100–106. https://doi.org/10.1016/j.jot.2023.10.003
Kellgren, J. H., Lawrence, J., amp, et al. (1957). Radiological assessment of osteo-arthrosis. Ann Rheum Dis, 16(4), 494–502.
Kramer, O., & Kramer, O. (2016). Scikit-learn. Machine learning for evolution strategies, 45–53
Kubakaddi, S., Ravikumar, K., & Harini, D. (2013). Measurement of cartilage thickness for early detection of knee osteoarthritis (koa). In: 2013 IEEE Point-of-Care Healthcare Technologies (PHT), pp. 208–211, IEEE
Lei, C., Wei, W., Liu, Z., Xiong, Q., Yang, C., Yang, M., Zhang, L., Zhu, T., Zhuang, X., Liu, C., amp, et al. (2019). Mammography-based radiomic analysis for predicting benign bi-rads category 4 calcifications. European Journal of Radiology,121, 108711.
Lespasio, M. J., Piuzzi, N. S., Husni, M. E., Muschler, G. F., Guarino, A., & Mont, M. A. (2017). Knee osteoarthritis: a primer. The Permanente Journal, 21
Liu, Z., Duan, T., Zhang, Y., Weng, S., Xu, H., Ren, Y., Zhang, Z., & Han, X. (2023). Radiogenomics: a key component of precision cancer medicine. British Journal of Cancer, 129(5), 741–753.
Mayerhoefer, M., Materka, A., Langs, G., Häggström, I., Szczypiński, P., Gibbs, P., & Cook, G. (2020). Introduction to radiomics. The Journal of Nuclear Medicine, 61, 488–495. https://doi.org/10.2967/jnumed.118.222893
Murphy, L., Schwartz, T. A., Helmick, C. G., Renner, J. B., Tudor, G., Koch, G., Dragomir, A., Kalsbeek, W. D., Luta, G., & Jordan, J. M. (2008). Lifetime risk of symptomatic knee osteoarthritis. Arthritis Care & Research: Official Journal of the American College of Rheumatology, 59(9), 1207–1213.
Neogi, T. (2013). The epidemiology and impact of pain in osteoarthritis. Osteoarthritis and Cartilage, 21(9), 1145–1153. https://doi.org/10.1016/j.joca.2013.03.018. Pain in Osteoarthritis
Parmar, C., Grossmann, P., Bussink, J., Lambin, P., & Aerts, H. (2015b). Machine learning methods for quantitative radiomic biomarkers. Scientific Reports,5. https://doi.org/10.1038/srep13087
Parmar, C., Grossmann, P., Rietveld, D., Rietbergen, M., Lambin, P., & Aerts, H. (2015a). Radiomic machine-learning classifiers for prognostic biomarkers of head and neck cancer. Frontiers in Oncology,5. https://doi.org/10.3389/fonc.2015.00272
Ponsiglione, A. M., Angelone, F., Amato, F., & Sansone, M. (2023). A statistical approach to assess the robustness of radiomics features in the discrimination of mammographic lesions. Journal of Personalized Medicine, 13(7), 1104.
Ringdahl, E., & Pandit, S. (2011). Treatment of knee osteoarthritis. American Family Physician, 83(11), 1287–1292.
Roemer, F. W., Demehri, S., Omoumi, P., Link, T. M., Kijowski, R., Saarakkala, S., Crema, M. D., & Guermazi, A. (2020). State of the art: imaging of osteoarthritis-revisited 2020. Radiology,296(1), 5–21.
Roos, E. M., & Arden, N. K. (2016). Strategies for the prevention of knee osteoarthritis. Nature Reviews Rheumatology, 12(2), 92–101.
Saarakkala, S., Julkunen, P., Kiviranta, P., Mäkitalo, J., Jurvelin, J. S., & Korhonen, R. K. (2010). Depth-wise progression of osteoarthritis in human articular cartilage: investigation of composition, structure and biomechanics. Osteoarthritis and Cartilage, 18(1), 73–81. https://doi.org/10.1016/j.joca.2009.08.003
Sánchez, L. E., Rundo, L., Gill, A., Hoare, M., Serrão, E. M., & Sala, E. (2021). Robustness of radiomic features in ct images with different slice thickness, comparing liver tumour and muscle. Scientific Reports,11. https://doi.org/10.1038/s41598-021-87598-w
Sansone, M., Fusco, R., Grassi, F., Gatta, G., Belfiore, M. P., Angelone, F., Ricciardi, C., Ponsiglione, A. M., Amato, F., Galdiero, R., amp, et al. (2023). Machine learning approaches with textural features to calculate breast density on mammography. Current Oncology,30(1), 839–853.
Saxena, S., Jena, B., Gupta, N., Das, S., Sarmah, D., Bhattacharya, P., Nath, T., Paul, S., Fouda, M. M., Kalra, M., amp, et al. (2022). Role of artificial intelligence in radiogenomics for cancers in the era of precision medicine. Cancers,14(12), 2860.
Van Griethuysen, J. J., Fedorov, A., Parmar, C., Hosny, A., Aucoin, N., Narayan, V., Beets-Tan, R. G., Fillion-Robin, J.-C., Pieper, S., & Aerts, H. J. (2017). Computational radiomics system to decode the radiographic phenotype. Cancer Research, 77(21), 104–107.
Virtanen, P., Gommers, R., Oliphant, T. E., Haberland, M., Reddy, T., Cournapeau, D., Burovski, E., Peterson, P., Weckesser, W., Bright, J., & et al. (2020). Scipy 1.0: fundamental algorithms for scientific computing in python. Nature Methods, 17(3), 261–272
Wu, P.-J., Masouleh, M. I., Dini, D., Paterson, C., Török, P., Overby, D. R., & Kabakova, I. V. (2019). Detection of proteoglycan loss from articular cartilage using brillouin microscopy, with applications to osteoarthritis. Biomed. Opt. Express, 10(5), 2457–2466. https://doi.org/10.1364/BOE.10.002457
Xie, Y., Dan, Y., Tao, H., Wang, C., Zhang, C., Wang, Y., Yang, J., Yang, G., & Chen, S. (2021). Radiomics feature analysis of cartilage and subchondral bone in differentiating knees predisposed to posttraumatic osteoarthritis after anterior cruciate ligament reconstruction from healthy knees. BioMed Research International,2021. https://doi.org/10.1155/2021/4351499
Xue, Z., Wang, L., Sun, Q., Xu, J., Liu, Y., Ai, S., Zhang, L., & Liu, C. (2022). Radiomics analysis using mr imaging of subchondral bone for identification of knee osteoarthritis. Journal of Orthopaedic Surgery and Research, 17(1), 414.
Zhang, Y., & Jordan, J. M. (2010). Epidemiology of osteoarthritis. Clinics in Geriatric Medicine, 26(3), 355–369. https://doi.org/10.1016/j.cger.2010.03.001. Osteoarthritis
Acknowledgements
The authors would like to thank the projects RESTORE (https://restoreproject.eu/) and SINPAIN (https://www.osteoarthritis-sinpain.eu/) for their contribution to this study. The authors also would like to thank Marco Ghiselli and Kristjá¡n Ö-rn Jóhannesson from the National University Hospital of Iceland for the medical image acquisition and Vicenzo Cangiano for his help in medical image segmentation.
Funding
This study is funded by European Union (CORDIS Grant agreement ID: 101057778) and Landspitali research fund
Author information
Authors and Affiliations
Contributions
All authors (F.A., F.K.C., G.P.T., H.J.Jr., A.M.P., M.K.G., F.T., F.AM. and P.G.) contributed to the design of the manuscript. Methodology, F.A., F.K.C. and G.P.T.; software, F.A., F.K.C. and G.P.T; validation, F.A. and P.G.; formal analysis, F.A., F.K.C. and G.P.T.; investigation, F.A., F.K.C. and G.P.T.; resources, P.G.; data curation, P.G.; writing—original draft preparation, F.A., F.K.C. and G.P.T.; writing—review and editing, H.J.Jr., A.M.P., M.K.G., F.T., F.A. and P.G.; supervision, F.A. and P.G.; project administration, P.G.; funding acquisition, P.G. and H.J.Jr.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Ethics approval and consent to participate
Approval for the study was obtained from the Icelandic Bioethics Commission (approval number: VSN-19-050V1) in March 2022. Subjects provided comprehensive written consent, ensuring transparency and accountability.
Consent for publication
Informed consent was obtained from all subjects involved in the study. Written informed consent has been obtained from the patient(s) to publish this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Angelone, F., Ciliberti, F.K., Tobia, G.P. et al. Innovative Diagnostic Approaches for Predicting Knee Cartilage Degeneration in Osteoarthritis Patients: A Radiomics-Based Study. Inf Syst Front (2024). https://doi.org/10.1007/s10796-024-10527-5
Accepted:
Published:
DOI: https://doi.org/10.1007/s10796-024-10527-5