Abstract
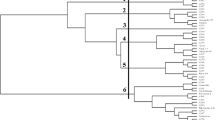
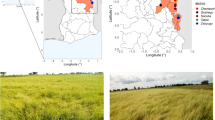
Understanding the existing genetic variation in the production system’s germplasm pool is the first step in any program aimed at improving crop genetic diversity. Regardless of its economic and sociocultural values in Ethiopia, the lack of research attention on ginger has limited its ability to improve genetically. Thus, the purpose of this study was to evaluate the genetic diversity among 100 ginger accessions collected from different agro-ecologies by using 12 polymorphic simple sequence repeat (SSR) markers. The polymorphism rate indicated that 97.2% of bands are polymeric out of the 139 distinct bands produced across all loci. The result also showed an average of 3.64 different alleles (Na), 1.53 number of effective alleles (Ne), and 0.55 Shannon information index (I). The observed heterozygosity was 0.13, and the expected heterozygosity was 0.28. Analysis of molecular variance revealed a 4% variation between populations and a 96% variation within populations. The 100 accessions were grouped into three clusters based on population structure analysis. Phylogenetic tree analysis has created three major tree branches and identified a significant number of identical duplicates. The experiment suggested that there might be potential markers associated with high rhizome yield and disease tolerance, but more research is necessary to confirm this. The experiment’s findings can serve as a foundation for Ethiopia’s efforts to improve the genetic conservation and improvement of ginger.





Similar content being viewed by others
Data availability
We declare that data of the experimental works at all stages and findings are available.
Abbreviations
- AMOVA:
-
Analysis of molecular variance
- CBT:
-
Center for Biotechnology
- CTAB:
-
Ctyl triacetate bromide
- DNA:
-
Deoxyribonucleic acid
- EDTA:
-
Ethylene diamine tetra acetate
- EST:
-
Expressed sequence tag
- HWE:
-
Hardy-Weinberg equilibrium
- PAG:
-
Polyacrylamide gel
- PCoA:
-
Principal coordinate analysis
- SSR:
-
Simple sequence repeat
- TE:
-
Tris-Hcl ethylene diamine tetra acetate
- UPGMA:
-
Unwighted pair group method with arthimetic mean
- ZOF:
-
Zingeber officinale
References
Akshitha HJ, Prasath D, Umesha K, Mohammed FP, Venkataravanappa V (2022) Molecular characterization of ginger genotypes using RAPD and SSR markers. J Hortl Sci 17(1):95–102. https://doi.org/10.24154/jhs.v17i1.1052
Chandrasekar A, Riju A, Sathyanath NV, Eapen SJ (2009) Spic EST is an annotated database on expressed sequence tags of spices (special issue 1). Global Science Books 3:50–53. http://220.227.138.213/spiceest/contprime.php
Das A, Gaur M, Barik DP, Subudhi E (2016) Genetic diversity analysis of 60 ginger germplasm accession using ISSR and SSR markers. Plant Biosyst 151(5):822–832. https://doi.org/10.1080/11263504.2016.1211197
Devi KD, Kshetrimayum P, Nandeibam SS, Huidrom S (2013) An efficient protocol for total DNA extraction from the members of order Zingiberales suitable for diverse PCR-based downstream applications. https://doi.org/10.1186/2193-1801-2-669
Earl DA, von Holdt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
FAOSTAT (2020) Food and Agriculture Organization of the United Nations. Rome Italy. http://faostat.fao.org/site/567/desktopdefault. Aspx page ID 567
Girma H, Kindie T (2008) The effect of seed rhizome size on the growth, yield, and economic return of Ginger (Zingiber officinale Rosc.) Asian J Plant Sci 7(2):213–217
Girma H, Mahammedsani Z, Melaku A (2022) Genetic conservation and importance of ginger in Ethiopia. A book chapter on genetically modified plants and beyond. https://doi.org/10.5772/intechopen.10307
Hervé N, Mahama O, Souleymane B, Serge FZ, Aimé SK, Mahamadou S (2017) Genetic diversity analysis of burkinafaso ginger (Zingibir offinale Rosc.) landraces using microsatilite markers. Int J Curr Res 9(05):501221–50126
Ismail NA, Rafii MY, Mahmud TMM, Hanafi MM, Gous M (2016) Molecular markers: a potential resource for ginger genetic diversity studies. Mol Biol Rep 43:1347–1358. https://doi.org/10.1007/s11033-016-4070-3
Jansen PM (1981) Spices, condiments and medicinal plants in Ethiopia, their taxonomy and agricultural significance. Centre for Agricultural Publishing and Documentation, Wageningen, the Netherlands. Serratia J Gen Microbiol 98:39–66
Jatoi SA, Kikuchi A, Yi SS, Watanabe JA (2006) Use of rice SSR markers as RAPD markers for genetic diversity analysis in Zingiberaceae. Breed Sci 56:107–111
Kaewsri W, Paisooksanti Y, Veesommai U, Eiadthong W, Vadrodaya S (2007) Phylogenetic analysis of Thai Amomum (Alpinioideae: Zingiberaceae) using AFLP markers. Kasetsart J (Nat Sci) 41:213–222
Kifle A, Merga J, Genene G, Abukiya G (2021) Technical guideline for management of ginger bacterial wilt and leaf spot diseases in Ethiopia. Farm Africa, Ethiopia. https://www.farmafrica.org/downloads/2022/
Langner E, Greifenberg S, Gruenwald J (1998) Ginger: history and use. Adv Ther 15(1):25–44. PMID: 10178636
Mahdi HJ, Andayani R, Aziz I (2013) Determination of phylogenetic and molecular characteristics of three Malaysian ginger cultivars (Zingiber officinale Roscoe) using microsatellite DNA. Trop Life Sci Res 24(2):65–76
Momina A, Sentayehu A, Girma H, Abush T (2011) Variability of ginger (Zingiber officinal Rosco) accessions for morphological and some quality traits in Ethiopia. Int J Agric Res 6(6):444–457
Neeta S (2019) Biotechnology and crop improvement of ginger (Zingiber officinale Rosc.). https://doi.org/10.5772/intechopen.88574
Nor AI, Rafii MY, Mahmud TM, Hanafi MM, Gous M (2019) Genetic diversity of torch ginger (Etlingera elatior) germplasm revealed by ISSR and SSR markers. Bio Med Res Int. https://doi.org/10.1155/2019/5904804
Peakall R, Smouse PE (2006) GenAlEx 6.5: Genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research-update. Bioinformatics 28(19):2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Prasanna KP, Subramanian S, Suresh J, Kannan JR, Gnanam R (2016) Effect of gamma rays on induction of chlorophyll mutants in ginger genotypes. Int J Agric 6(5):105–110
Perrier X, Jacquemoud-Collet JP (2006) DARwin software http://darwin.cirad.fr/darwin
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multi-locus genotype data. Genetics 155:945–959
Tamura K, Stecher G, Kumar S (2021) MEGA11: molecular evolutionary genetics analysis version 11. Mol Biol Evol 38:3022–3027
Wada E, Feyissa T, Tesfaye K, Asfaw Z, Potter D (2021) Genetic diversity of Ethiopian cocoyam (Xanthosoma sagittifolium (L.) Schott) accessions as revealed by morphological traits and SSR markers. PLoS ONE 16(1):e0245120. https://doi.org/10.1371/journal.pone.0245120
Wahyuni S, Xu DH, Bermawie N, Tsunematsu H, Ban T (2003) Genetic relationships among ginger accessions based on AFLP marker. J Biotechnol Part 8(2):60–68
Yeh FC, Yang R, Boyle T (1999) Microsoft window-based free ware for population genetic analysis (POPGENE Ver. 1.31). Canada, AB: University of Alberta
Acknowledgements
This research was conducted with funding from Southern Agricultural Research Institute and Addis Ababa University.
Funding
This research report was part of the first author’s PhD work and it was supported by budgets from the Southern Agricultural Research Institute and Addis Ababa University; Institute of Biotechnology. However, the authors declare that no funds or other supports were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the planning and implementation of the work. Genene Gezahegn performed material preparation, data collection, analysis, and first draft. Yayis Rezene and Tileye Feyissa did technical support, supervision, and edition of the manuscript versions. Hence, all authors read and approved the final manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gezahegn, G., Rezene, Y. & Feyissa, T. Genetic diversity analysis of Ethiopian ginger (Zingiber officinale Roscoe) accessions using simple sequence repeat (SSR) markers. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01972-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01972-x