Abstract
A bred plant variety is a valuable tool for enhancing crop production and total yield. Recently, our breeding team developed three high seed yielding synthetic cultivars of bitter fennel. To characterize these newly breeding populations and differentiate them from each other and their parental populations, we conducted a Distinctness, Uniformity, and Stability (DUS) assessment using morphological traits evaluated over two consecutive years, as well as 20 Start Codon Targeted (SCoT) markers. The evaluation of morphological traits revealed that, five of qualitative traits exhibited monomorphism, while two other qualitative traits displayed polymorphism. Additionally, the investigation of eight quantitative traits showcased significant diversity. The Combined Over Years Distinctiveness analysis at a 1% significance level, based on eight quantitative characteristics, indicated distinctiveness for two breeding populations, namely Early and Medium Maturity Synthetic cultivars, compared to other populations. The Combined Over Years Uniformity analysis identified that all of the breeding populations and their parental populations were uniform in terms of the measured traits. The differences between Phenotypic Coefficients of Variation and Genotypic Coefficients of Variation were not particularly noticeable in the experimental material used currently. The majority of the measured traits showed high heritability, highlighting their stability over time. PCA utilized for quantitative morphological features, the first two principal components collectively accounted for more than 86.8% of the phenotypic variation. Subsequently, a dendrogram was constructed, which clustered breeding and parental populations into three distinct groups. The 15 SCoT markers produced a total of 164 extended fragments, with 139 (85%) of them showed polymorphic.The genetic similarity between the populations exhibited a range of 0.08–0.81, and cluster analysis utilizing Jaccard similarity coefficients and the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) technique revealed clear distinctions among the three breeding populations and their parental populations. Altogether, the findings indicate that morphological DUS descriptors effectively distinguish some the populations, while others with overlapping expressions cannot be discriminated. Notably, SCoT markers demonstrate a fine ability to detect distinctions among fennel populations, suggesting the possibility of utilizing them as a complement to morphological traits in the DUS test. As a conclusion, these three newly breeding populations of fennel are distinct from each other and from their parental populations, and they can advance for registration and seed purity testing.
Similar content being viewed by others
Introduction
Discoveries in Kurdistan’s Shanidar Cave suggest that the history of human use of medicinal plants dates back as far as 60,000 years ago (Lietava 1992). Bitter fennel (Foeniculum vulgare Mill. var. vulgare), as the source subspecies of fennel derived products, is one of the earliest medicinal plants, with numerous applications in food, beverage, and pharmaceutical industries. Bitter fennel, hereafter referred to simply as fennel, originated from the Mediterranean regionbut nowadays it is cultivated almost worldwide. The most economically valuable product of fennel plants is the seeds, which are mostly used for essential oil extraction (Rather et al. 2016; Barros et al. 2010). Numerous research indicated that fennel herbal medicines possess valuable curative properties such as antifungal, anti-cancer, antibacterial, anti-inflammatory, and antioxidant effects (Hamdy Roby et al. 2013; Ouariachi et al. 2014; Oliveira et al. 2015; Salami et al. 2016; Afifi et al. 2021; Akbari et al. 2023). Additionally, various pharmaceutical products in the market are formulated based on the phytochemical substances found in fennel such as trans anethol and methyl chavicole (Kooti et al. 2015; Edoardo et al. 2010; Elagayyar et al. 2001).
The global fennel seed production is estimated at nearly 800 thousand tons, with Iran emerging as a major contributor (FAO 2021). Iranian fennels exhibit remarkable diversity in habitats, and their adaptation to local environments over an extended period suggests the presence of distinct fennel landraces in each region. This has been proved through morphological research (Bahmani et al. 2012a, b, 2015, 2024; Shojaiefar et al. 2015; Akbari et al. 2015), phytochemical combinations (Akbari et al. 2023, 2024; Izadi Darbandi et al. 2023; Bahmani et al. 2016), genetic properties (Sheidai et al. 2007; Bahmani et al. 2012a, b, 2013; Shojaiefar et al. 2015).
To meet the rising global demand for fennel seeds, it is imperative to increase fennel seed yield. To achieve this, our options are applying optimized agronomic management and along with developing high yielding fennel cultivars. To improve agronomic management in fennel farms, some researchers reported different kinds of chemical and organic fertilizers, irrigation plans, sowing date, and depth, and plant density to get more seeds and essential oil per area (Ehsanipour et al. 2012; Khorshidi et al. 2010; Mohamed and Abdu 2004). All of these factors can increase fennel yields but not to the level that can fulfill the world’s need, while plant breeding has almost endless prospects in developing high yielding cultivars.
Intentional exploitation of heterosis through breeding efforts has led to significant yield increases in numerous crops with hybrid breeding playing a pivotal role. Successful hybrid breeding relies on effective pollination control, and for this, male sterility systems are highly useful. In the absence of such system for hybrid cultivars, synthetic cultivar arises as an alternative approach to utalize hybrid vigor (Liu et al. 2020; Maydell et al. 2021). Heterosis exhibits notable efficacy in outcrossing crops such as fennel with cross-pollination rates ranging from 82.2 to 91.4% (Ramanujam et al. 1964; Sawargaonkar et al. 2018). Population improvement through methods such as synthetic cultivar development is highly recommended for fennel breeding programs (Poehlman and Sleper 1995).
Any newly developed cultivars must be distinct (D) from previously released cultivars, uniform (U) withing the population, and stable (S) in terms of morphological and other examined features (Pourabed et al. 2015). The International Union for the Protection of New Varieties of Plants (UPOV) criteria require the fundamental Distinctness, Uniformity, and Stability (DUS) test for identifying and registering newly developed cultivars. The establishment of distinctiveness is the most important aspect of DUS testing. If the differences in traits are constant and obvious, a variety might be deemed distinctive (UPOV 2002). The DUS tests are mostly founded on morpho-physiological features recorded from plants grown in the same environment in a close vicinity (Cooke 1995). There are 27 morphological DUS descriptors that are suggested for performing DUS test for fennel (UPOV 2018). Because there are so many existing cultivars, the traditional method of DUS experiments is time-consuming and costly, necessitating high expenses and specialized workers, as well as often subjective decisions (Cooke 1995). Furthermore, most of those characters are quantitative, and their expression is influenced by environment, so replicated assessment is highly necessary. Compared to morphological traits, incorporating molecular markers into DUS testing has been demonstrated to be faster and more cost-effective (Mailer et al. 1994). Molecular markers, consisting of nucleotide sequences displaying polymorphism among individuals due to variations like insertion, deletion, point mutations, duplication, and translocation, and they do not necessarily impact their corresponding gene’s activity. An optimal DNA marker should showcase co-dominance, uniform genomic distribution, high reproducibility, and the ability to detect significant levels of polymorphism (Nadeem et al. 2018). Various types of molecular markers, including microsatellites (Wang et al. 2015; Sousa et al. 2017; Tiwari et al. 2018; Gupta et al. 2022), have been assessed for their efficacy in DUS tests. Restriction Fragment Length Polymorphism (RFLP) (Gebhardt et al. 1989), Amplified Fragment Length Polymorphism (AFLP) (De Riek et al. 2001), Sequence tagged Microsatellite Site (STMS) (Singh et al. 2004), Random Amplified Polymorphic DNA (RAPD) (Yadav and Sciences 2010), Inter Simple Sequence Repeats (ISSR) (Prevost and Wilkinson 1999), RAPD and ISSR (Shukla et al. 2011; Lu et al. 2009) are reliable and effective methods of cultivar identification.
The Start Codon Targeted (SCoT) polymorphism is widely used in PCR-based molecular markers due to its rapidity, cost-effectiveness, minimal DNA quantity needed, and the absence of a prerequisite for DNA sequence knowledge. SCoT markers produce more reliable and uniform bands, establishing them as influential polymorphic markers that effectively highlight distinctions among individuals. Hence, this method can find applied in diverse fields, encompassing genetic mapping and population research in the realm of plants (Collard and Mackill 2009).
Our research group has developed innovative fennel breeding populations, each consisting of three synthetic cultivars strategically designed to improve yield performance. For any newly developed population to be considered a new variety, distinctness, uniformity, and stability are crucial factors. This study aimed to evaluate the second-generation breeding populations against their parental populations over two years, utilizing the DUS test through morphological traits. Additionally, 20 SCoT markers were employed to enhance the discrimination and identification of fennel breeding populations.
Materials and methods
Plant material
Our breeding team developed breeding populations comprising three synthetic cultivars by crossing various populations possessing enhanced general combining ability to improve yield performance (Bahmani et al. 2023). The breeding populations, alongside selected elite parents were utilized in this research. Specifically, Early Maturity Synthetic involving two of its parental populations (Hashtgerd and Fasa), Medium Maturity Synthetic involving four of its parental populations (Moghan, Fozve, Meshkinshahr, Khash), and Late Maturity Synthetic involving two of its parental populations (Chahestan and Hajiabad) were employed in this research. The study included DUS testing based on morphological traits recorded in 2018 and 2019, as well as the analysis of 20 populations using SCoT markers (Table 1).
Morphological characterization
Field experiment
The investigation took place in the 2018 and 2019 growing seasons, where seeds of the populations were sown in an open field testing operation under a randomized complete block design (RCBD) with three replications. The field had sandy loam soil and was located in Pakdasht (Tehran province, Iran), with a semi-arid climate (Tables 2). Each population was planted in a 3 m2 plot, with an interrow distance of 25 cm to achieve a plant density of 10 plants per m2 in both years (Khorshidi et al. 2010). No diseases or pests were detected throughout the growing seasons.
Data collection
For the two-year data collection, we randomly selected twenty plants in each plot and assessed a range of morphological traits. These traits comprised eight quantitative and seven qualitative characteristics recommended by the DUS test guidelines (Table 3).
Statistical analysis for the morphological data
To establish distinctiveness based on visually assessed traits, we considered distinctiveness between two populations evident if the expression of one or more traits distinctly fell into two states, as outlined in the test guidelines. The analysis of the eight quantitative DUS traits was performed using DUSNT software (Watson 2004). Following UPOV guidelines (2008a and 2008b), we conducted analyses for Combined Over Years Distinctiveness (COYD) and Combined Over Years Uniformity (COYU). Comparing the genotypic and phenotypic coefficients of variation (GCV and PCV), as well as heritability using pooled data of quantifiable variables collected over two years, allowed for the assessment of stability. To provide more information, Principal Component Analysis and Clusters Analysis were executed utilizing R statistical software (version 4.2.3).
PCR amplification through SCoT markers
DNA extraction
Genomic DNA was extracted from young leaves of the 20 populations using the CTAB method as described by Doyle and Doyle (1987) with specific modifications, as outlined by Kafkas et al. (2006). The quality of the DNA samples was verified using a Qubit Fluorometer (Invitrogen) and electrophoresis. Subsequently, the concentration of DNA samples were adjusted to 5–10 ng/µl using Tris–EDTA buffer, and the samples were then stored at −20 °C for further analysis.
Genetic assessment
As indicated in Table 4, 20 SCoT primers were used for amplification to genetically evaluate the 20 populations (Collard and Mackill 2009; Yan et al. 2016). Polymerase chain reaction (PCR) was carried out in a final volume of 20 µl using 10 µl of master mix (2X amplicon), 2 µl of primer (10 M), 2 µl of genomic DNA, and 6 µl of deionized water after adjusting the amplification conditions. The following temperature condition was used to amplify PCR: initial denaturation at 94 °C for 4 min, followed by 35 cycles including DNA denaturation at 94 °C for 1 min and DNA annealing at 49–62 °C for 1 min, then extension at 72 °C for 2 min and a final extension at 72 °C for five minutes. The PCR product was separated using electrophoresis for 1 h at 80 V in a %1.5 agarose gel with TAE buffer. The separated bands were observed using ethidium bromide and UV light.
Statistical analysis for the SCoT markers
After electrophoresis of the PCR products, the clear and sharp bands were scored so that each allele amplified with the primers was scored as 1 for presence, while absence was indicated by 0. To analyze the genetic data, the number of total bands (TB), the number of polymorphic bands (PB), and the percentage of polymorphism (% PL) were determined. Marker performance measures such as polymorphic information content (PIC), marker index (MI), and resolving power (RP) were calculated. The PIC value for each primer was calculated as PICi = 2 fi (1-fi), where PICi represents the PIC of the ith marker, fi denotes the frequency of the fragments that are present, and 1-fi denotes the frequency of absent fragments (Roldán-Ruiz et al. 2000). Prevost and Wilkinson (1999) determined the marker index (MI) employing the following equation: MI = PIC × number of polymorphism bands. According to Gilbert et al. (1999), the equation Rp = ΣIb represents the Resolving power (Rp), where the formula for Ib is 1 - 2 ×|0.5 - p|. In this equation, p denotes the frequency of the presence of a specific band (or allele) in the population. To determine similarity level among the populations, Jaccard’s coefficient of similarity was estimated (Jaccard 1908). Cluster analysis based on the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) technique was conducted using R statistical software, version 4.2.3.
Results
Morphological characterization for DUS test
All eleven fennel populations exhibited consistent scores for seven qualitative traits measured over two years of study. Among these traits, foliage attitude, curvature tip of the leaf, foliage color, plant grumolo formation, and male sterility showed no variation, while the remaining two traits displayed only minor variation (Table 5, Fig. 1).
On the other hand, the eight quantitative traits revealed significant variability among the populations over the two years. For instance, leaf length ranged from 26.62 cm (Early Maturity Synthetic) to 29.963 cm (Chahestan). Cotyledon length varied from a maximum of 6.829 cm (Fasa) to a minimum of 4.981 cm (Hajiabad). Fasa displayed the earliest onset of flowering at 75.14 days, while Chahestan took the longest at 120.48 days. The tallest height at flowering time was recorded in Hajiabad (148.858 cm), while the shortest was from Hashtgerd (91.75 cm). The primary umbel diameter ranged from 6.50 cm in Hajiabad to 12.79 cm in Hashtgerd. Moghan exhibited the highest thousand-seed weight at 4.086 g, whereas Chahestan recorded the lowest weight at 2.527 g. Petiole length of the first leaf varied from 8.671 cm in Hashtgerd to 11.238 cm in Meshkinshahr. Additionally, the main umbel appeared earliest in Early Maturity Synthetic at 74.4 days and latest in Late Maturity Synthetic at 119.01 days. Tables 5 and 6 present the observed qualitative and quantitative morphological characters.
COY-D and COY-U analysis
The data collected from the eight quantitative traits underwent COY-D statistical analysis at a 1% significance level. Each population was treated as a candidate variety and compared to the remaining 10 populations, serving as reference populations, to generate a pairwise distinctiveness matrix through COY-D analysis (Table 7). Results from comparing the Early Maturity Synthetic with other populations revealed distinctiveness across all examined quantitative traits. Similarly, the Medium Maturity Synthetic, when compared with other populations, demonstrated distinctiveness across all examined quantitative traits. Furthermore, the comparison of the Late-Maturity Synthetic with other populations indicated distinctiveness across most traits compared to the other populations. However, non-distinctiveness was observed from its parental population, Hajiabad (Table 7). Additionally, a COY-D analysis utilizing the MJRA model was performed on the eight quantitative traits (Table 8). For all traits, the F1 ratio (Ratio of Genotype MS to Genotype × Year MS) was significant, suggesting that the environment had less of an impact on the expression of the trait. The following traits: time of main umbel appearance, length of petiole, length of cotyledons, time of beginning of flowering, and umbel diameter showed a significant F2 ratio (Ratio of Genotype × Year MS to Genotype × Rep MS). This indicates that the consistency in distinctiveness patterns for these traits varied across the two years of the experiment. In summary, the environment significantly influenced the distinctiveness patterns among the populations for these traits, limiting their ability to establish varietal distinctiveness within the current set of populations.
COY-U analysis was conducted for eight quantitative traits using pairwise comparisons of the standard deviations (SDs) for each trait (Table 9). A thorough examination revealed that among the 11 fennel populations, Khash and Late Maturity Synthetic exhibited uniformity for all eight characters, followed by Chahestan, Fozveh, and Medium Maturity Synthetic, which showed uniformity for seven characters. Moreover, there was a lack of uniformity observed for most of the traits, albeit staying within the permissible limits recommended by UPOV criteria.
The pooled analysis of the quantitative traits showed a relatively minor difference between PCV and GCV which is a reflection of the low environmental effects on the traits expressions underscoring consistency and stability of the traits (Table 6). Phenotypic and genotypic coefficients of variation were classified into low (< 10%), moderate (10–20%), and high (> 20%) categories. The time of the beginning of flowering (17.81 and 17.72) and the time of appearance of the main umbel (16.52 and 16.40) displayed moderate values of PCV and GCV. Conversely, lower PCV and GCV values were noted for leaf length (4.96 and 2.38). These results suggested a significant impact of genetic components on the expression of most of the traits, while environmental factors had minimal influences. This result revealed high heritability for the time of beginning of flowering (98.99%), time of appearance of the main umbel diameter (98.55%), and thousand seed weight (80.90%), suggesting that both additive and non-additive gene effects have an equal role in their heritability. In contrast, low heritability (23.10%) was observed for leaf length, signifying a prevalence of non-additive gene effects and greater environmental influence on its expression,
thereby limiting its performance stability over time. Overall, the analysis of various genetic parameters for eight quantitative DUS descriptors revealed substantial variability and heritability within the populations. However, leaf length stood out due to its lower heritability, emphasizing the importance of these parameters in accurately characterizing fennel populations.
Principal component analysis (PCA)
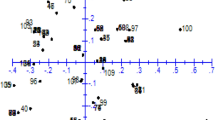
PCA stands out as a highly precise tool for grouping germplasm, effectively subdividing them into subgroups based on the extent of variations in their characteristics. In this study, PCA was implemented on the quantitative morphological traits to group the 11 fennel populations. The first two principal components together represented 86.8% of the phenotypic variation, as indicated by PCA eigenvectors. PC1 accounting for 73.4 percent of the total variation, was directly linked to the time of appearance of the main umbel, time of the beginning of flowering, height at flowering, and, leaf length. Meanwhile, PC2, contributing to 13.4 percent of the overall variance, primarily reflected the traits of main umbel diameter, length of cotyledons, thousand seed weight, and petiole length. Visualization of the first two PCs in a biplot (Fig. 2) effectively demonstrated the population relations. PCA successfully categorized the cultivars based on their quantitative traits, grouping each synthetic cultivar with its respective parental populations in one cluster. Notably, in the biplot, the Chahestan population exhibited distinct separation from all the others populations, even the other late maturity populations, potentially reflecting its unique origins and extensive diversity in morphological traits.
Clustering analysis
The dendrogram constructed based on eight quantitative traits, put breeding populations and parental populations, in three major clusters (Fig. 3) which was similar to the results from the PCA. The first cluster of the dendrogram consisted of Late Maturity Synthetic with its parental populations Hajiabad and Chahestan, the second cluster consisted of Medium Maturity Synthetic with its parental populations Fozve, Moghan, Khash, and Meshkinshahr, and the third cluster consisted of Early Maturity Synthetic with its parental populations Hashtgerd and Fasa. This approach was able to perfectly distinguish the three synthetic cultivars from their relevant parental populations.
Molecular markers characterization
Assessment of polymorphism using SCoT marker
Out of the 20 SCoT primers, 15 amplified clear, sharp, and reproducible bands in the 20 fennel populations. The SCoT primers generated a total of 164 amplified bands, of which 139 (85%) were polymorphic with an average of 10.93 bands for each primer, SCOT-9 with 18 bands had the most bands, whereas SCoT-35 primers with 3 bands had the least. Among the populations, the Late Maturity Synthetic had the most bands (129) while the Sarpolzahab had the least (64). Table 10 shows the findings for the primers that were utilized in this study. The percentages of polymorphisms in the tested primers varied from 54 to 100%. The two primers, SCoT-29 and SCoT-31, showed 100% polymorphism, indicating their significant potential to distinguish different populations from each other.
The polymorphism bands ranged from 2 (SCoT-2) to 17 (SCoT-9) with an average of 9.13. The amount of polymorphism of a marker is represented by the polymorphic information content (PIC), which can range from zero to half, and it is calculated using the frequency and number of amplified fragments (Roldàn-Ruiz et al. 2000). Markers with greater PIC values have a stronger ability to distinguish populations. The PIC values for fifteen primers ranged from 0.10 (SCoT-31) to 0.40 (SCoT-11), with an average of 0.25. Moreover, the resolving power (RP), a key indicator of a primer’s ability to distinguish between various populations (Powell et al. 1996), ranged from 1.20 (SCoT-35) to 7.50 (SCoT-9) in our study, with an average of 4.03. Due to their high RP, SCoT-9 and SCoT-11 were able to differentiate most of the populations. Marker Index (MI), as an indicator for the total effectiveness of a primer, in the study ranged from 0.38 (SCoT-2) to 4.80 (SCoT-11) with the average of 2.50. Considering the PIC, RP, and MI measures, revealed that primer SCoT-11 was the most efficient in examining the variation among the populations (PIC 0.40, RP 7.20, and MI 4.80). Other primers such as SCoT-9 and SCoT-30 were also performed well.
Analysis of genetic relationships
The 20 fennel populations were discriminated by the polymorphic SCoT markers employed in this investigation. The Jaccard coefficient was used to calculate the genetic similarity matrix for the fennel populations (Fig. 4). The genetic similarity of the fennel populations varied from 0.08 to 0.81; this indicated that there is a sufficient variation among the fennel populations. The similarity matrix showed that among all the 20 populations, Sardasht and Nayriz were the most similar, whereas Early Maturity Synthetic and Rafsanjan were the least similar populations.
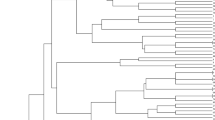
Cluster analysis based on marker data, a useful way to evaluate genetic relationships among individuals (Randi and Lucchini 2002). Cluster analysis was performed using Jaccard similarity coefficients and the UPGMA technique, which effectively distinguished all the 20 populations. The fennel populations were classified into seven distinct groups, highlighting a wide range of genetic diversity (Fig. 5). Groups V and VI each comprised a single fennel population, namely Early Maturity Synthetic and Fozveh, respectively. Within Group IV, which comprised multiple populations, including Medium Maturity Synthetic, Late Maturity Synthetic, Meshkinshahr, Fasa, and Hashtgerd, were clustered together. This classification successfully differentiated Early and Late Maturity Synthetic cultivars from their respective parent populations and discriminated among three synthetic fennel cultivars and their parental populations.
Discussion
Accurate identification and evaluation of plant materials is necessary for variety creation, the registration process, and breeders’ rights protection (Hale et al. 2006). In this study, the distinctness, uniformity and stability of three breeding populations of fennel and their elite parental populations was evaluated using morphological traits and SCoT markers.
The findings indicate that certain morphological DUS descriptors effectively can distinguish some of the populations. Therefore, additional markers or descriptors may be necessary to complement the morphological DUS descriptors for distinctiveness. Morphological traits are commonly employed to identify plant materials, however, this approache is debatable since environmental factors can have a significant impact on morphological traits. Furthermore, due to the time and expense needed, this method is inefficient (Singh et al. 2004). Moreover, morphological traits on theirown are incapable of detecting discrimination between certain cultivars that are morphologically similar but exhibit diverse agronomical characteristics. DNA analysis can help to overcome all of these constraints because it is quicker, more cost-effective, and more precise (Korir et al. 2013). In recent years, various novel marker approaches have emerged, and among them, Start Codon Targeted (SCoT) markers are becoming increasingly popular owing to their high polymorphism and resolvability (Guo et al. 2012; Bhattacharyya et al. 2013). The SCoT marker system is characterized by its simplicity, cost-effectiveness, polymorphic nature, repeatability, and reliability (Gorji et al. 2011; Zhang et al. 2015). To use molecular approaches for cultivar identification, first must understand the level of polymorphism shown by the different methods in the species under investigation. In their study, Farshadfar et al. (2017) found that employing ten SCoT primers to amplify DNAs from 16 fennel accessions resulted in 55 bands, of which 51 were polymorphic, indicating a polymorphism rate of 93%. Similarly, Bahmani et al. (2013) reported obtaining 1043 polymorphic DNA fragments in their research involving 25 fennel accessions from various regions in Iran, utilizing 10 RAPD markers to assess genetic diversity. Furthermore, Bahmani et al. (2012a, b) and Poudineh et al. (2018) examined genetic diversity among fennel cultivars using ISSR markers. Both investigations disclosed a notable level of polymorphism. Our results demonstrated a substantial percentage of polymorphic primers (85 percent), indicating that the diversity of SCoT markers utilized in this study serves as an effective tool for distinguishing and identifying cultivars. Out of the 20 SCoT markerse used in our study, 15 of threm were able to discriminateall the populations, showcasing their potential utility in DUS testing as well. Numerous research recorded the utilization of molecular markers in crop registration testing. In rice (Singh et al. 2004), cucumber (Bernet et al. 2003), and rapeseed (Tommasini et al. 2003), fingerprinting has extensively employed RAPD, AFLP, and SSR markers. When comparing molecular and morphological features, various scientists have revealed that molecular markers provide for more exact discrimination than morphological features, as demonstrated in maize (Dillmann et al. 1997). According to previous findings molecular markers could be employed for pre-screening or classification of existing and emerging cultivars. Furthermore, the outcomes of clustering using SCoT markers and morphological features in the present investigation did not seem comparable, revealing differences in exploring genetic diversity. This could be due to the fact that different molecular markers represent genetic variety in different sections of the genome. The current study highlighted the efficacy of SCoT markers in demonstrating the distinctness of the fennel populations for DUS test.
Conclusion
In summary, this DUS test using both SCoT markers and morphological traits was able successfully show the distinctiveness, uniformity, and stability of our newly developed fennel cultivars. According to the results, it is easier to properly identify and classify cultivars when molecular markers and morphological characteristic analysis are combined. These analyses revealed that the newly developed populations of native fennel are generally distinct from the parent populations and can be utilized for seed purity registration and testing. Beside our previous open field testing operations to assess yiled performance of the three fennel synthetic cultivars, the DUS test here in this current study also showed that synthetic cultivar can be a great breeding approach to develop high yielding cultivars in crops such as fennel.
References
Afifi SM, El-Mahis A, Heiss AG, Farag MA (2021) Gas chromatography–mass spectrometry-based classification of 12 fennel (Foeniculum vulgare Miller) varieties based on their aroma profiles and estragole levels as analyzed using chemometric tools. ACS Omega 6:5775–5785
Akbari A, Izadi Darbandi A, Bahmani K, Ramshini HA (2015) Evaluation of drought tolerance in synthetic varieties and superior ecotypes of fennel (Foeniculum vulgare Mill.). Iran J Field Crop Sci 17(3):193–204
Akbari A, Izadi-Darbandi A, Bahmani K, Farhadpour M, Ebrahimi M, Ramshini H, Esmaeili Z (2023) Assessment of phenolic profile and antioxidant activity in developed breeding populations of fennel (Foeniculum vulgare Mill). Biocatal Agric Biotechnol 48:102639
Akbari A, Bahmani K, Kafkas NE, Hamijo T, Darbandi AI (2024) Evaluation of seed yield, essential oil compositions, and fatty acid profiles in advanced fennel (Foeniculum vulgare Mill) breeding populations. Biocatal Agric Biotechnol 16:103118
Bahmani K, Izadi Darbandi A, Sadat Noori SA, Jafari AA (2012a) Determination of interrelationships among phenotypic traits of Iranian fennel (Foeniculum vulgare Mill.) using correlation, stepwise regression, and path analyses. J Essent Oil Bear Plants 15(3):424–444
Bahmani K, Izadi-Darbandi A, Jafari AA, Sadat Noori SA, Farajpour M (2012b) Assessment of genetic diversity in Iranian fennels using ISSR markers. J Agric Sci 4:79–84
Bahmani K, Izadi Darbandi A, Sadat Noori SA, Jafari AA (2013) Assessment of the genetic diversity in Iranian fennels by RAPD markers. J Herbs Spices Med Plants 19:275–285. https://doi.org/10.1080/10496475.2013.791909
Bahmani K, Izadi Darbandi A, Ramshini HA, Moradi N, Akbari A (2015) Agro-morphological and phytochemical diversity of various Iranian fennel ecotypes. Ind Crops Prod 77:282–294
Bahmani K, Izadi Darbandi A, Faleh Alfekaiki D, Sticklen M (2016) Phytochemical diversity of fennel ecotypes from various growth types and origin. Agrobiol Rec 14(5):1530–1547
Bahmani K, Akbari A, Izadi Darbandi A, Warner RM (2023) Development of high-yielding fennel synthetic cultivars based on polycross progeny performance. Agric Res 12:357–363
Bahmani K, Akbari A, Izadi Darbandi A, Ghamari T (2024) Phenological traits, seed yield, and essential oil yield of fifty populations of bitter fennel (Foeniculum vulgare). Int J Hortic Sci 11:391–410
Barros L, Carvalho AM, Ferreira ICFR (2010) The nutritional composition of fennel (Foeniculum vulgare): shoots, leaves, stems, and inflorescences. LWT—Food Sci Technol 43:814–818. https://doi.org/10.1016/J.LWT.2010.01.010
Bernet GP, Bramardi S, Calvache D, Carbonell EA, Asins MJ (2003) Applicability of molecular markers in the context of protection of new varieties of cucumber. Plant Breed 122:146–152
Bhattacharyya P, Kumaria S, Kumar S, Tandon P (2013) Start codon targeted (SCoT) marker reveals genetic diversity of Dendrobium nobile Lindl., an endangered medicinal orchid species. Gene 529:21–26. https://doi.org/10.1016/j.gene.2013.07.096
Collard BCY, Mackill DJ (2009) Start codon targeted (SCoT) polymorphism: a simple, novel DNA marker technique for generating gene-targeted markers in plants. Plant Mol Biol Rep 27:86–93. https://doi.org/10.1007/S11105-008-0060-5
Cooke R (1995) Varietal identification of crop plants. In: Skerritt JH, Appels R (eds) New Diagnostics in Crop Sci. CAB International, Wallingford, pp 33–63
De Riek J, Calsyn E, Everaert I, Van Bockstaele E, De Loose M (2001) AFLP based alternatives for the assessment of distinctness, uniformity, and stability of sugar beet varieties. Theor Appl Genet 103:1254–1265. https://doi.org/10.1007/S001220100710
Dillmann C, Bar-Hen A, Guerin D, Charcosset A, Murigneux A (1997) Comparison of RFLP and morphological distances between maize Zea mays L. inbred lines. Theor Appl Genet 95:92–102. https://doi.org/10.1007/BF00299755
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Edoardo M, Curcuruto G, Ruberto G (2010) Screening the essential oil composition of wild Sicilian fennel. Biochem Syst Ecol 38:213–223
Ehsanipour A, Razmjoo J, Zeinali H (2012) Effect of nitrogen rates on yield and quality of fennel (Foeniculum vulgare Mill.) accessions. Ind Crops Prod 35:121–125
Elagayyar M, Draughon FA, Golden DA (2001) Antimicrobial activity of essential oil from plants against selected pathogenic and saprophytic microorganisms. J Food Prot 64:1019–1024
FAO (2021) FAOstat. Available online: http://www.fao.org/home/en. Accessed 22 Dec 2021
Farshadfar M, Moradzade N, Farshadfar E, Shirvani H (2017) Genetic diversity among fennel (Fueniculum vulgare Mill.) accessions using morphological and SCoT markers. Iran J Rageland Forest Plan Breed. Genet Res 25(2):212–231
Gebhardt C, Ritter E, Debener T, Schachtschabel U, Walkemeier B, Uhrig H, Salamini F (1989) RFLP analysis and linkage mapping in Solanum tuberosum. Theor Appl Genet 78:65–75. https://doi.org/10.1007/BF00299755
Gilbert JE, Lewis RV, Wilkinson MJ, Caligari PDS (1999) Developing an appropriate strategy to assess genetic variability in plant germplasm collections. Theor Appl Genet 98:1125–1131. https://doi.org/10.1007/S001220051176
Gorji AM, Poczai P, Polgar Z, Taller J (2011) Efficiency of arbitrarily amplified dominant markers (SCoT, ISSR and RAPD) for diagnostic fingerprinting in tetraploid potato. Am J Potato Res 88(3):226–237. https://doi.org/10.1007/s12230-011-9187-2
Guo DL, Zhang JY, Liu CH (2012) Genetic diversity in some grape varieties revealed by SCoT analyses. Mol Biol Rep 39:5307–5313
Gupta AJ, Anandhan S, Manjunathagowda DC, Benke AP, Mahajan V, Kad SK, Singh M (2022) Complement test for distinctiveness, uniformity and stability testing of kharif onion (Allium cepa L.) varieties. Genet Resour Crop Evol 69:2217–2229. https://doi.org/10.1007/S10722-022-01372-Z
Hale AL, Farnham MW, Menz MA (2006) Effectiveness of PCR-based markers for differentiating elite broccoli inbreds. J Am Soc Hort Sci 131:418–423
Hamdy Roby MH, Sarhan MA, Selim KA, Khalel KI (2013) Antioxidant and antimicrobial activities of essential oil and extracts of fennel (Foeniculum vulgare) and chamomile (Matricaria chamomilla). Ind Crop Prod 44:437–445
Izadi-Darbandi A, Akbari A, Bahmani K, Warner R, Ebrahimi M, Ramshini H (2023) Fatty acid profiling and oil content variation among iranian fennel (Foeniculum vulgare Mill. Var. vulgare) Landraces. Int J Hortic Sci Technol 10(2):193–202
Jaccard P (1908) Nouvelles recherches sur la distribution fl orale. Bull Soc Vaud Sci Nat 44:223–270
Kafkas SO, Özkan H, Ak BE, Acar I, Atlı HS, Koyuncu S (2006) Detecting DNA polymorphism and genetic diversity in a wide pistachio germplasm: comparison of AFLP, ISSR and RAPD markers. J Am Soc Hort Sci 131(4):522–529
Khorshidi J, Mirahmadi SF, Tabatabaei MF (2010) Oil content and yield of Foeniculum vulgare Mill. cv. Soroksary seeds as affected by different plant cultivation densities. J Am Sci 6(11):1098–1100
Kooti W, Moradi M, Ali Akbari S, Sharafi N, Asadi M, Ashtary Larky D (2015) Therapeutic and pharmacological potential of Foeniculum vulgare Mill: a review. J Herbmed Pharmacol 4(1):1–9
Korir NK, Han J, Shanqquan L, Wang C, Kayesh E, Zhang J, Fang J (2013) Plant variety and cultivar identification: advances and prospects. Crit Rev Biotechnol 33:111–125. https://doi.org/10.3109/07388551.2012.675314
Lietava J (1992) Medicinal plants in a middle paleolithic grave Shanidar IV. J Ethnopharmacol 35(2):263–266
Liu J, Li MJ, Zhang Q, Wei X, Huang XH (2020) Exploring the molecular basis of heterosis for plant breeding. J Integr Plant Biol 62:287–298. https://doi.org/10.1111/jipb.12804
Lu X, Liu L, Gong Y, Zhao L, Song X (2009) Cultivar identification and genetic diversity analysis of broccoli and its related species with RAPD and ISSR markers. Sci Hortic 4:645–648
Mailer R, Scarth R (1994) Discrimination among cultivars of rapeseed (Brassica napus L.) using DNA polymorphisms amplified from arbitrary primers. Theor Appl Genet 87:697–704. https://doi.org/10.1007/BF00222895
Maydell D, Brandes J, Lehnert H, Junghanns W, Marthe F (2021) Breeding synthetic varieties in annual caraway: observations on the outcrossing rate in a polycross using a high-throughput genotyping system. Euphytica 217:1–15
Mohamed MAH, Abdu M (2004) Growth and oil production of fennel (Foeniculum vulgare Mill): effect of irrigation and organic fertilization. Biol Agric Hort 22:31–39
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F, Alsaleh A, Labhane N, Özkan H (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32(2):261–285
Oliveira F, Alves M, Damasceno L, Oliveira M, Dias J, Crotti M, Tavares C (2015) Cytotoxicity screening of essential oils in cancer cell lines. Rev Bras Farmacogn 25:183–188
Ouariachi E, Lahhit N, Bouyanzer A, Hammouti B, Paolini J, Majidi L, Desjobert M, Costa J (2014) Chemical composition and antioxidant activity of essential oils and solvent extracts of Foeniculum vulgare Mill. from Morocco. J Chem Pharm Res 6(4):743–748
Poehlman JM, Sleper DA (1995) Breeding Field Crops, 4th edn. Iowa State University Press, Ames, p 494
Poudineh Z, Fakheri BA, Sirosmehr AR, Shojaei S (2018) Genetic and morphological diversity of fennel by using ISSR marker and biplot analysis. Indian J Plant Physiol 23(3):564–572. https://doi.org/10.1007/s40502-018-0390-8
Powell W, Morgante M, Andre C, Hanafey M, Vogel J, Tingey S, Rafalski A (1996) The comparison of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238. https://doi.org/10.1007/BF00564200
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112. https://doi.org/10.1007/S001220051046
Ramanujam S, Joshi BS, Saxena MBL (1964) Extent and randomness of cross-pollination in some umbelliferous spices of India. 62–67
Randi E, Lucchini V (2002) Detecting rare introgression of domestic dog genes into wild wolf (Canis lupus) populations by Bayesian admixture analyses of microsatellite variation. Conserv Genet 3:31–45. https://doi.org/10.1023/A:1014229610646
Rather MA, Dar BA, Sofi SN, Bhat BA, Qurishi MA (2016) Foeniculum vulgare: a comprehensive review of its traditional use, phytochemistry, pharmacology, and safety. Arabian J Chem 9:S1574–S1583
Roldán-Ruiz I, Dendauw J, Van Bockstaele E, Depicker A, De Loose M (2000) AFLP markers reveal high polymorphic rates in ryegrasses (Lolium spp.). Springer 6:125–134. https://doi.org/10.1023/A:1009680614564
Salami M, Rahimmalek M, Ehtemam MH, Sabzalian MR (2016) Variation in bioactive compounds and anatomical characteristics of different fennel (Foeniculum vulgare Mill.) populations as affected by self-pollination. J Appl Bot Food Qual 89:38–48
Sawargaonkar SL, Singh AK, Sahu S (2018) Stability analysis for seed yield and yield attributing traits in fennel (Foeniculum vulguare Mill.). J Spices Aromat Crops 74–80
Sheidai M, Kalhor-Home N, Poorneydanei A (2007) Cytogenetic study of some populations of Foeniculum vulgare (Umbelliferae) in Iran. Caryologia 60(3):257–261
Shojaiefar S, Mirlohi A, Sabzalian R, Yaghini H (2015) Seed yield and essential oil content of fennel influenced by genetic variation and genotype×year interaction. Ind Crop Prod 71:97–105
Shukla S, Joshi D, Singh RS (2011) Suitability of RAPD and ISSR to complement agro-morphological DUS descriptors for establishing distinctiveness in indigenous local strains of Kalanamak rice. Indian J Agric Sci 81(11):994–1000
Singh RK, Sharma RK, Singh AK, Singh VP, Singh NK, Tiwari SP, Mohapatra T (2004) Suitability of mapped sequence tagged microsatellite site markers for establishing distinctness, uniformity and stability in aromatic rice. Euphytica 135:135–143. https://doi.org/10.1023/B:EUPH.0000014905.10397.08
Sousa TV, Caixeta ET, Alkimim ER, de Oliveira ACB, Pereira AA, Zambolim L, Sakiyama NS (2017) Molecular markers useful to discriminate Coffea arabica cultivars with high genetic similarity. Euphytica. https://doi.org/10.1007/S10681-017-1865-9
Tiwari J, Ali N, Devi S, Kumar V (2018) Development of microsatellite markers set for identification of Indian potato varieties. Sci Hortic 231:22–30
Tommasini L, Batley J, Arnold GM (2003) The development of multiplex simple sequence repeat (SSR) markers to complement distinctness, uniformity and stability testing of rape (Brassica napus L.) varieties. Theor Appl Genet 106:1091–1101
UPOV (2002) General introduction to the examination of distinctness, uniformity and stability and the development of harmonized descriptions of new varieties of plants. International Union for the Protection of New Varieties of Plants, Document TG/I/3. UPOV, Geneva, Switzerland
UPOV (2008a) ‘Examining distinctness’, in UPOV TGP/9/1, pp 1–36, website: http://www.upov.int/edocs/tgpdocs/en/tgp_9_1.pdf (accessed 17 June 2013)
UPOV (2008b) ‘Examining uniformity’, in UPOV TGP/10/1, pp 1–14, website: http://www.upov.int/edocs/tgpdocs/en/tgp_10_1.pdf (accessed 17 June 2013)
UPOV (2018) Guidelines for the conduct of tests for distinctness, homogeneity and stability- Fennel (Foeniculum vulgare Mill.). UPOV TG/183/4 (proj.1)
Van Gastel A (1996) Variety evaluation. In: Seed science and technology, International Center for Agricultural Research in the Dry Areas (ICARDA), pp 141–150
Wang L, Jun Q, Chang L, Liu L, Huang XH (2015) Assessment of wheat variety distinctness using SSR markers. Euphytica 195:435–452
Watson S (2004) Using the DUS software. WORKSHOP ON DATA HANDLING organized by the International Union for the Protection of New Varieties of Plants (UPOV) in cooperation with the State Forestry Administration of China, the Ministry of Agriculture of China and the State Intellectual Property Office of China with the financial assistance of the Ministry of Agriculture, Forestry and Fisheries of Japan. Beijing, June 9 to 11, 2004. UPOV/DATA/BEI/04/12: 2-19
Yadav V, Sciences IS-A (2010) Comparative evaluation of maize inbred lines (Zea mays L.) according to DUS testing using morphological, physiological and molecular markers. Agric Sci 1:131–142
Yan H, Zhang Y, Zeng B, Yin G, Zhang X, Ji Y, Huang L, Jiang X, Liu X, Peng Y, Ma X (2016) Genetic diversity and association of EST-SSR and SCoT markers with rust traits in orchardgrass (Dactylis glomerata L.). Molecules 21(1):66
Zhang J, Xie W, Wang Y, Zhao X (2015) Potential of start codon targeted (SCoT) markers to estimate genetic diversity and relationships among Chinese Elymus sibiricus accessions. Molecules 20(4):5987–6001. https://doi.org/10.3390/molecules20045987
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK). The authors have not disclosed any funding.
Author information
Authors and Affiliations
Contributions
A.A conducted the experiments and processed the experimental data, contributing significantly to the original draft preparation. A.I.D and S.K provided supervision, ensuring the quality and coherence of the research. K.B and E.K were instrumental in reviewing and editing the manuscript for refinement. O.B contributed to the data analysis, while K.B, M.K, J.R, and O.B were involved in the investigation process. All authors have thoroughly reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have not disclosed any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Akbari, A., Bahmani, K., Kazan, M. et al. Analysis of Fennel Breeding Populations Based on Distinctness, Uniformity and stability (DUS) testing via Morphological Descriptors and DNA Molecular Markers. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01934-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01934-3