Abstract
Central Asia contains rich genetic resources for apricots. With its mountainous geography, Kyrgyzstan is the country where wild and cultivated apricot forms are preserved. The present study revealed genetic diversity and population structures in 91 apricot accessions from different regions of Kyrgyzstan. Fifteen simple sequence repeat (SSR) primers were used, capillary electrophoresis was performed, and the number of alleles per primer ranged from 4 to 11. A significant level of variation was determined among apricots originating from Kyrgyzstan. In the resulting dendrogram, all apricot accessions were divided into five groups, and the materials belonging to the northern and southern regions were grouped separately. At the same time, the population structure of apricots was analyzed. Accordingly, individuals have 0.80 or more membership coefficients; therefore, they are likely pure and non-admixed. There were 34 apricot accessions that have been determined as pure. The remaining 53 individuals had varying membership coefficients and were likely to be admixed by at least two subpopulations. This study is the most comprehensive research on Kyrgyzstan-origin apricots, providing vital information on their identification, preservation, and use in future research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Apricot belongs to the Rosaceae family, Prunus genus, which consists of eight species. Apricots, cultivated worldwide, belong to the Prunus armeniaca L. species. World apricot production is approximately 3.57 million tons. Regarding production, Türkiye ranks first with 800 thousand tons, followed by Uzbekistan and Iran (FAO 2021).
Research revealed that apricot culture began in Asia approximately 5000 years ago. It has been stated that two regions in Asia, the Fergana Valley (border of Uzbekistan, Tajikistan, and Kyrgyzstan) and China, are possible origin regions for domesticated apricots (Vavilov 1951; Decroocq et al. 2016). Further research divided domesticated and wild apricots into two main ecogeographic groups based on fruit size: the first group includes Central Asia, which includes parts of China, Afghanistan, Pakistan, and northern India, and the second is the Dzhungar-Zailij group, which includes Kyrgyzstan, Kazakhstan, and China's Xinjiang province (Kostina 1969). Central Asia, including China, is the center of origin of the apricot species, including a high genetic diversity (Bourguiba et al. 2020). In addition, two more ecogeographic groups, Iranian-Caucasian and European, have been defined for cultivated apricots (Kostina 1969; Decroocq et al. 2016). The Central Asian group is the oldest group with the richest variation. Self-incompatibility is observed in most cultivars in this group; the fruits are small to medium. Likewise, the Iranian-Caucasian group is generally self-incompatible, but compared to the Central Asian group, the fruits are more significant, and the chilling requirements are lower. The European group is younger and less diverse. This group probably originates from some Asian apricots. Also, European cultivars, which are mostly self-compatible, have lower chilling requirements and have only a short maturation period. (Maghuly et al. 2005).
Central Asia has rich resources regarding apricot genetic diversity, including cultivated and wild materials. The apricot genotypes here contain significant variations in high sugar content, fruit shape, fruit color, fruit flesh, late flowering and late ripening, drought, cold, and salt tolerance. International researchers had limited access to evaluate these genetic resources until the collapse of the Soviet Union. Wide-scale use of these resources will allow for broader adaptation, expansion of production regions, and efficient and more sustainable production (Zaurov et al. 2013). On the other hand, it has been reported that Central Asian apricots require higher temperatures for flowering in spring, complete flower development more slowly, and, as a result, have a more extended flowering period. Likewise, it has been emphasized that flower formation stops in Central Asian apricots below 8 °C; on the contrary, the process continues in European apricots under these conditions, and as a result, European apricots are more affected by the cold (Lomakin 1977). Based on these points, it is seen that apricots of Central Asian origin have a more comprehensive adaptation.
Although different marker systems are used to determine plant diversity, DNA-based markers not affected by environmental conditions provide more successful and safe results. Among these, simple sequence repeats (SSR) are considered a preferred marker as they are codominant, have high reproducibility, and have greater genome abundance (Nadeem et al. 2018). This marker system is widely used for genetic diversity and fingerprint research in the apricot species (Hormaza et al. 2002; Zhebentyayeva et al. 2003; Hayashi et al. 2008; Pedryc et al. 2009; Yılmaz et al. 2012; Bakır et al. 2015: Gurcan et al. 2015; Corrado et al. 2021; Ehliz et al. 2021; Sheikh et al. 2021; Wang et al. 2023).
Kyrgyzstan, located within the homeland region of apricots, has a significant richness of cultivated and wild apricots. This study was conducted to determine the genetic diversity and population structure among apricot genotypes containing wild and cultivated forms collected from different regions of Kyrgyzstan.
Materials and methods
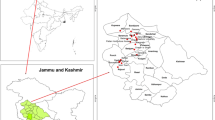
Ninety-one apricot accessions collected from the Chuy, Issyk-Kul, Osh, and Batken regions of Kyrgyzstan were used as material. Chuy and Issyk-Kul are located in northern Kyrgyzstan, while Batken and Osh are located in southern Kyrgyzstan. As described in Table 1, the materials consist of wild apricots, landraces, and cultivars and represent the apricot diversity of that region. The cultivars whose names are mentioned in Table 1 are widely grown in the Central Asian region (Zaurov et al. 2013).There is an elevated level of observable variation among the fruit morphologies of the apricots used in the study (Fig. 1).
Fruit differences in some apricot accessions used in the study (Accession number of apricots described in Table 1; 16: Cultivar; 25: Wild type; 36: Wild type; 91: Landrace)
DNA isolation from young leaves of apricot accessions was performed using the CTAB method, according to Doyle and Doyle (1990). DNA concentrations were determined with the help of a semi-automatic microplate reader (PowerWave HT, BIO-TEK Instruments, Inc., Winooski, VT, USA), and DNA concentrations of 10 ng/µl were prepared for PCR research.
SSR analysis
SSR primers specified by Gürcan et al. (2015) were used for SSR analyses. All accessions were studied with 15 SSR primers, which successfully resulted in the tests performed with these primers (Table 2). PCR components and PCR cycles were made as reported by the same researchers and described below.
The polymerase chain reaction (PCR) was performed according to the method described by Schuelke (2000). PCR reactions were performed including fluorescently labeled M13 primer with 6-FAM, NED, PET, or VIC. The PCR reaction mix consisted of 2 µL of 10X PCR buffer, 0.6 µL of 50 mM MgCl2, 2 µL of 10 mM dNTP, 0.15 µL of 10 µM of a sequence-specific forward primer with M13(-21) tail at its 5´ end, 0.15 µL of 10 µM of a sequence-specific reverse primer, and 0.20 µL of 10 µM the universal fluorescent-labeled M13(-21) primer, 0.2 µL 5U/µL Taq DNA polymerase, and 3 µL of 25 ng/µL sample DNA. The final reaction volume was brought up to 20 µL with distilled water. The thermal cycler was programmed for denaturation at 94 °C for 5 min followed by 35 cycles of 94 °C for 40 s for denaturation, 60 °C for 30 s for primer annealing, 72 °C for 40 s for extension, and a final 5-min extension step at 72 °C. Four multiplex groups were assigned to primer pairs.
Allele lengths were determined by running PCR results on the ABI 3500 capillary electrophoresis system (Applied Biosystems, Foster City, CA, USA) at Erciyes University Genome and Stem Cell Center.
PowerMarker V3.025 software was run to determine the expected allele number (n) (He) and observed heterozygosity (Ho) and polymorphism information content (PIC). The neighbor-joining (NJ) method made the dendrogram using a genetic similarity matrix in MEGA6 (Tamura et al. 2007) and PowerMarker programs.
Results and discussion
Genetic diversity
Ninety-one apricot genotypes used were studied with 15 SSR primers. The allele numbers expected (He) and observed heterozygosity (Ho) and PIC (Polymorphism Information Content) values of these primers are given in Table 2. Accordingly, 107 alleles were obtained in all genotypes with 15 SSR primers. The highest allele was found in the SSR 3 primer (11), and the lowest was found in the SSR 1 and SSR4 primers (4). Expected heterozygote values in the primers are between 0.70 (SSR 4) and 0.88 (SSR 6). The observed heterozygosity values ranged between 0.58 (SSR 14) and 0.95 (SSR 3). On the other hand, PIC values were found between 0.65 (SSR 4) and 0.86 (SSR 6). Estimates of He was higher than that of Ho for ten loci (Table 2) showing that null alleles are high for the loci. The high frequency of null alleles indicates amplification failure for one allele in heterozygotes, which results in a loss of data and a decrease in the apparent heterozygosity (Dakin and Avise 2004). In some previous research on apricots, the number of alleles per marker was generally higher. Bourguiba et al. (2012), in their study of apricots from the regions around the Mediterranean basin, found the number of alleles per marker to be between 5 and 18. Gürcan et al. (2015) determined the number of alleles between 5 and 20 in apricot cultivars from five countries. Decroocq et al. (2016), in their study with 230 wild and 142 cultivated apricot materials, determined the number of alleles per marker as 6–22 and the observed heterozygosity values as 0.57–0.81. The relatively higher number of alleles obtained in these investigations, probably was because the cultivars used belong to different regions of the world.
On the other hand, Corrado et al. (2021), in their study of a narrower apricot population in Italy, determined the number of alleles to be between 2 and 10, similar to the results in our study.
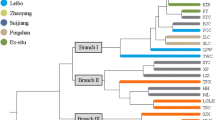
According to the dendrogram obtained with the 15 SSR primers used in this study, it is seen that there are significant variations among genotypes (Fig. 2). Almost all of the genotypes are genetically separated from each other. On the other hand, apricot genotypes taken from northern (Chuy and Issyk-Kul, red color in the dendrogram) and southern Kyrgyzstan (Batken and Osh, blue color in the dendrogram) regions were grouped geographically. Genotypes are divided into five basic groups in the dendrogram. With some exceptions, genotypes taken from the southern regions of Kyrgyzstan were included in groups 1, 4, and 5, and genotypes taken from the northern regions were included in groups 2 and 3. In general, genotypes in groups 1, 4, and 5 consist of dried apricots, and those in groups 2 and 3 consist of table apricots. Decroocq et al. (2016), in their study with SSR markers, reported that wild apricots taken from the Issyk-Kul region of Kyrgyzstan are in the same group as apricots taken from the Kazakhstan region and that they belong to the Dzhungar-Zailig ecogeographic group. Researchers emphasized that the apricots taken from Sarı Chelek, located in the south of Kyrgyzstan and close to the Ferghana region, are in a completely different group from the others and belong to the Central Asian ecogeographic group.
There was a total of seven genotypes in group 1 in the dendrogram. In this group, only genotype 1 belongs to the northern region (Chuy), while the remaining six belong to the southern Batken (73, 75, 77) and Osh (91, 92, 94) regions. There are 18 genotypes in group number two, all belonging to the northern regions. Genotype 56 was taken from the Chuy region, and the remaining 17 were taken from the Issyk-Kul region.
Group number three is the most populous group in the dendrogram and has 33 genotypes. Again, all of the genotypes in this group belonging to Issyk-Kul region, except genotype 57 (Chuy). In this group, genotypes 13 and 15 and 21 and 22 were the most similar dual genotypes. This large group was divided into two subgroups, with 23 and 10 apricot genotypes in the subgroups, respectively. All apricot genotypes in the fourth group in the dendrogram belong to the Batken and Osh regions south of Kyrgyzstan. The genotypes within the group were separated, and there were 16 genotypes. The two 'Supkhani' cultivars (79, 81) included in the study are located very close to each other in this group.
The last group, the fifth group, consists of 12 genotypes, of which genotypes 5, 6, and 7 belong to the northern Kyrgyzstan (Issyk-Kul) region. These three genotypes are grouped in a small subgroup, separate from the others. The remaining nine genotypes were taken from the southern regions of Kyrgyzstan. ‘Kandak’ (83 and 87) and ‘Mirsandjali’ cultivars (85, 86) in this group were reported to have the same AFLP profiles in a previous study (Zhebentyayeva et al. 2008).
Population structure
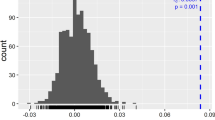
The population structure of apricot accessions was determined using the STRUCTURE 3.2 program defined by Evanno et al. (2005). LnP(D) value reached its maximum at K = 3, and this method's most appropriate population number was evaluated as 3 (Fig. 3). Based on SSR markers, the substructuring Bayesian analysis identified three subpopulations in our data set (Fig. 4). Accessions with membership coefficiency of 0.80 (Q value) or more to a subpopulation are considered pure (Fukunaga et al. 2005). In contrast, individuals with less membership coefficients are considered admixed by at least two distinct genotypes. In this study, 38 individuals have 0.80 or more membership coefficients; therefore, they are likely pure and non-admixed. The remaining 53 individuals had varying membership coefficients and were likely to be admixed by at least two subpopulations.
When the bar plot obtained by STRUCTURE analysis is examined, it is seen that there have been mixtures and gene transfers between apricot genotypes over many years in the Central Asian region in general (Fig. 4). The STRUCTURE analysis results were compatible with the groupings obtained in the NJ dendrogram in Fig. 2. Consistent with the results in the dendrogram, a grouping appropriate to the geographical distribution of genotypes was obtained in the STRUCTURE analysis. Bourguiba et al. (2017) found that geographical origin and propagation mode are essential for structuring genetic diversity in North African apricot species. Also, Bourguiba et al. (2020) identified strong relationships between memberships of accessions within the clusters and their geographical regions for global apricot accessions. Similarly, Sheikh et al. (2021) determined a relationship between the genetic structure and the geographical origins of the materials in their study on Himalayan apricots.
There were 21 genotypes in the red group in the bar plot, and the Q value was above 0.80 in 8 of them (Fig. 4). Genotypes 13, 15, 21, and 22 in this group showed an almost homogeneous structure. In addition, genotypes such as 19, 42, 46, 47, 16, 29, and 24 are included in the red group, but they also contain infrastructure in the other two groups. On the other hand, numbers 30, 32, and 43 showed almost half the red and blue group resolution. The red group in the bar plot consists of genotypes in group 3 in the NJ dendrogram (Fig. 2). This group generally consists of genotypes belonging to the northern Kyrgyzstan region.
In the bar plot, the green-colored group consists of 29 genotypes, and the Q value of 12 (91, 57, 88, 83, 85, 56, 67, 64, 62, 61, 86, 60) is above 0.80. All genotypes in this group belong to the Batken and Osh regions in southern Kyrgyzstan. The feature of these genotypes is that they are mostly drying. This reveals genetic differences between northern (Issyk-Kul) and southern (Batken, Osh) Kyrgyzstan apricot populations. Similarly, Bourguiba et al. (2020) noted the genetic structure within Central Asia's wild Prunus armeniaca populations, with a highly differentiated Kyrgyz population in the South.
The accessions in this group are distributed in groups 4 and 5 in the dendrogram. In the dendrogram, 2 of the three samples of the 'Mirsandjali' cultivar (85, 86) were placed in group 5, and one (65) was placed in group 4. However, in the bar plot, three samples (60, 80, and 81) were in the green group. Similarly, two “Kandak” genotypes (83, 87) in group 5 in the dendrogram are in the green group (78, 82) in the bar plot. It is seen that the Q values of such genotypes in group 5 in the dendrogram belong to the blue and red groups at a rate of approximately 20–40% in the green group on the barplot. In group 4 in the dendrogram, the Q value of the blue and red groups is much lower than that of the green group. This shows that the genetic background is less complex in these genotypes.
In the bar plot, the blue-colored group contained the most accessions. There are 40 genotypes in this group, and 18 of them have a Q value greater than 0.80. The genotypes in this group are generally distributed in groups 1, 2, 3, and 5 in the NJ dendrogram. The entire group numbered two, and a subgroup of groups 1, 3, and 5 formed the green-colored group in the bar plot. These genotypes mainly include genotypes belonging to the Issyk-Kul region. However, genotypes 64, 73, 74, 75, 76, 78, and 82 belong to the Batken region in southern Kyrgyzstan. On the other hand, it is seen that 20–50% of the 15 genotypes in this group have a green and red group background. This shows that there are interactions between genotypes in the process and that seed propagation occurs occasionally.
Conclusion
The results of our study show that Kyrgyzstan has a significant level of apricot diversity. This study is the first comprehensive genetic diversity study on apricots in this country. The presence of wild, local, and standard cultivars has enabled the formation of a more complex genetic structure with the interactions between them over time. However, the unique genetic diversity in the Central Asian region is threatened with extinction due to unauthorized human economic activity, overgrazing, and the almost complete absence of natural seed regeneration caused by the lack of protective and forest restoration measures (Shalpykov and Dolotbakov 2011). In order to preserve this diversity, it is necessary to establish ex situ core collections and in situ conservation strategies. The results of our study also provide information for establishing such collections. In addition, these results could be used for parent selection and other events for new research.
References
Bakır M, Dumanoğlu H, Erdoğan V, Ernim C, Macit T (2015) Characterization of wild apricot (Prunus armeniaca L.) genotypes selected from cappadocia region (Nevşehir-Türkiye) by SSR markers. J Agricult Sci 25:498–507. https://doi.org/10.15832/ankutbd.457850
Bourguiba H, Audergon JM, Krichen L, Trifi-Farah N, Mamouni A, Trabelsi S et al (2012) Loss of genetic diversity as a signature of apricot domestication and diffusion into the Mediterranean Basin. BMC Plant Biol 12:49. https://doi.org/10.1186/1471-2229-12-49
Bourguiba H, Batnini MA, Krichen L, Trifi-Farah N, Audergoni JM (2017) Population structure and core collection construction of apricot (Prunus armeniaca L.) in north Africa based on microsatellite markers. Plant Genetic Res 15(1):21–28. https://doi.org/10.1017/S1479262115000313
Bourguiba H, Scotti I, Sauvage C, Zhebentyayeva T, Ledbetter C, Krška B, Remay A, D’Onofrio C, Iketani H, Christen D, Krichen L, Trifi-Farah N, Liu W, Roch G, Audergon JM (2020) Genetic structure of a worldwide germplasm collection of Prunus armeniaca L. reveals three major diffusion routes for varieties coming from the species’ center of origin. Front Plant Sci 11:638. https://doi.org/10.3389/fpls.2020.00638
Corrado G, Forlani M, Rao R, Basile B (2021) Diversity and relationships among neglected apricot (Prunus armeniaca L.) landraces using morphological traits and SSR markers: implications for agro-biodiversity conservation. Plants 10:1341. https://doi.org/10.3390/plants10071341
Dakin EE, Avise JC (2004) Microsatellite null alleles in parentage analysis. Heredity 93:504–509. https://doi.org/10.1038/sj.hdy.6800545
Decroocq S, Cornille A, Tricon D, Babayeva S, Chague A, Eyquard JP (2016) New insights into the history of domesticated and wild apricots and its contribution to Plum pox virus resistance. Mol Ecol 25:4712–4729. https://doi.org/10.1111/mec.13772
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ehliz F, Karakurt Y, Çelik C (2021) Molecular Characterization of Apricot (Prunus armeniaca L) Cultivars with SSR Markers. J Faculty Agricult 16(1):79–85. https://doi.org/10.15832/ankutbd.457850
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 8:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
FAO (2021) https://www.fao.org/faostat/en/#data/QCL
Fukunaga K, Hill J, Vigouroux Y, Matsuoka Y, Sanchez GJ et al (2005) Genetic diversity and population structure of Teosinte. Genetics 169:2241–2254. https://doi.org/10.1534/genetics.104.031393
Gürcan K, Ocal N, Yilmaz KU, Ullah S, Erdogan A, Zengin Y (2015) Evaluation of Turkish apricot germplasm using SSR markers: genetic diversity assessment and search for Plum pox virus resistance alleles. Sci Hort 193:155–164. https://doi.org/10.1016/j.scienta.2015.07.012
Hayashi K, Shimazu K, Yaegaki H, Yamaguchi M, Iketani H, Yamamoto T (2008) Genetic diversity in fruiting and flower ornamental Japanese apricot (Prunus mume) germplasms assessed by SSR markers. Breeding Sci 58:401–410. https://doi.org/10.1270/jsbbs.58.401
Hormaza JI (2002) Molecular characterization and similarity relationships among apricot (Prunus armeniaca L.) genotypes using simple sequence repeats. Theor Appl Genet 104:321–328. https://doi.org/10.1007/s001220100684
Kostina KF (1969) The use of varietal resources of apricots for breeding. Proceed Trudi Nikita Botan Garden 40:45–63
Lomakin EN (1977) Genepool of apricots, goals and breeding work in Central Asia, p. 13–22. In: Nauchno-metodicheskoe soveshanie pokulturye abrikosa v Srednei Azii [Scientific-methodological conference on apricot culture], MSKh Uz.S.S.R., Tashkent, Uzbekistan [in Russian]
Maghuly F, Borroto Fernandez E, Ruthner S, Pedryc A, Laimer M (2005) Microsatellite variability in apricots (Prunus armeniaca L.) reflects their geographic origin and breeding history. Tree Genet Genomes 1:151–165. https://doi.org/10.1007/s11295-005-0018-9
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Hatipoğlu R, Ahmad F et al (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32:261–285. https://doi.org/10.1080/13102818.2017.1400401
Pedryc A, Ruthner S, Herman R, Krska B, Hegedus A, Halasz J (2009) Genetic diversity of apricot revealed by a set of SSR markers from linkage group G1. Sci Hort 121:19–26. https://doi.org/10.1016/j.scienta.2009.01.014
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol 18:233–234. https://doi.org/10.1038/72708
Shalpykov KT, Dolotbakov AK (2011) Current status of wild crop relatives in Kyrgyzstan. In: Conservation and sustainable use of biodiversity of fruit crops and wild fruit species (Ed: MK Turdieva, AK Kayimov, KI Baymetov, FU Mustafina, EA Butkov). Bioversity International, ISBN 978-92-9043-914-1
Sheikh ZN, Sharma V, Shah RA, Raina S, Aljabri M, Mir JI, Alkenani N, Hakeem KR (2021) Elucidating genetic diversity in apricot (Prunus armeniaca L.) cultivated in the north-western himalayan provinces of india using SSR markers. Plants 10(12):2668. https://doi.org/10.3390/plants10122668
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol 24:1596–1599
Vavilov NI (1951) Phytogeographical basis of plant breeding. In: The Origin, Variation, Immunity and Breeding of Cultivated Plants, Vol. 13, Chester KS (trans), pp. 13–54. Chronica Botanica, Ronald Press Company, New York
Wang X, Wang L, Sun Y, Chen J, Liu Q, Dong S (2023) Genetic diversity and conservation of Siberian apricot (Prunus sibirica L.) based on microsatellite markers. Sci Rep 13:11245. https://doi.org/10.1038/s41598-023-37993-2
Yılmaz KU, Paydas-Kargi S, Dogan Y, Kafkas S (2012) Genetic diversity analysis based on ISSR, RAPD and SSR among Turkish Apricot Germplasms in Iran Caucasian eco-geographical group. Sci Hort 138:138–143. https://doi.org/10.1016/j.scienta.2012.02.017
Zaurov DE, Molnar TJ, Eisenman SW, Ford TM, Mavlyanova RF, Capik JM, Funk CR, Goffreda JC (2013) Genetic resources of apricots (Prunus armeniaca L.) in Central Asia. Hortscience 48(6):681–691. https://doi.org/10.21273/HORTSCI.48.6.681
Zhebentyayeva TN, Reighard GL, Gorina VM, Abbott AGH (2003) Simple sequence repeat (SSR) analysis for assessment of genetic variability in apricot germplasm. Theor Appl Genet 106:435–444. https://doi.org/10.1007/s00122-002-1069-z
Zhebentyayeva TN, Reighard GL, Lalli DA, Gorina VM, Krška B, Abbott AG (2008) Origin of resistance to plum pox virus in Apricot: what new AFLP and targeted SSR data analyses tell. Tree Genet Genomes 4:403–417. https://doi.org/10.1007/s11295-007-0119-8
Acknowledgements
The authors thank the Scientific Research Projects Unit of Erciyes University for funding and supporting the project (No. FOA-2018-7877).
Funding
Open access funding provided by the Scientific and Technological Research Council of Türkiye (TÜBİTAK).
Author information
Authors and Affiliations
Contributions
AU, HP Methodology, Writing. KG Methodology, Editing. KT Methodology. EY Validation. MI Data curation. SD Validation. All authors commented on previous versions and read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Uzun, A., Pinar, H., Gürcan, K. et al. Genetic diversity and population structure of wild and cultivated apricots collected from Kyrgyzstan. Genet Resour Crop Evol (2024). https://doi.org/10.1007/s10722-024-01894-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10722-024-01894-8