Abstract
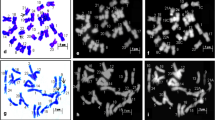
Rice serves as a primary dietary source of nutrients and calories for over half of the global human population. With India being the world's second-largest producer and largest rice exporter, it boasts a vast collection of rice germplasm. Among these, traditional aromatic rice cultivars (TARCs) occupy a prominent position. Despite the challenges posed by the small size of chromosomes and procedural limitations, karyotype analysis of rice has not received adequate attention. However, such evaluation of karyotype diversity may enrich the global gene pool, which is a prerequisite for the success of new breeding programs. The enzymatic maceration and air drying (EMA) method, a fundamental technique in molecular cytogenetics, has rejuvenated plant chromosome research by providing a cytoplasm-free background. The objective of the present study was to document, decode, and conserve the untapped karyotypic diversity in TARCs of India. Karyotype diversity was evaluated in 22 populations of TARCs using the EMA-based chromosome preparation method, followed by staining with non-fluorescent Giemsa and DNA base-specific fluorescent dye DAPI (4’-6-diamidino-2-phenylindole) for the first time. Some unique and foundational highlighting features of this study include the diversity in staining pattern, total chromatin length, morphology of individual and landmark sat-bearing chromosomes in each cultivar, intraspecific sat-bearing chromosome number in several cultivars, the karyotype formula, and the effect of DAPI on sat-bearing chromosomes. The genomic diversity decoded and conserved in Indian rice for its adaptive potential is expected to be highly advantageous for global rice breeders and genome researchers in both conventional and new breeding initiatives.









Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this article.
References
Ainsworth D (2022) UN convention on biological diversity (COP 15) held in Montreal Canada
Ansari HA, Ellison NW, Bassett SA, Hussain SW, Bryan GT, Williams WM (2016) Fluorescence chromosome banding and FISH mapping in perennial ryegrass. Lolium Perenne l BMC Genom 17(1):1–9
Bhuvaneswari S, Gopala Krishnan S, Bollinedi H, Saha S, Ellur RK, Vinod KK, Singh IM, Prakash N, Bhowmick PK, Nagarajan M, Singh NK (2020) Genetic architecture and anthocyanin profiling of aromatic rice from Manipur reveals divergence of Chakhao landraces. Front Genet 11:570731
Chatterjee SD, Adhikari B, Ghosh A, Ahmed J, Bal Neogi S, Pandey N (2007) Rice diversity of West Bengal. Published by West Bengal biodiversity board, Dept. of environment, Govt. of West Bengal
Cheng Z, Buell CR, Wing RA, Gu M, Jiang J (2001) Toward a cytological characterization of the rice genome. Genome Res 11(12):2133–2141
Cheng CY, Motohashi R, Tsuchimoto S, Fukuta Y, Ohtsubo H, Ohtsubo E (2003) Polyphyletic origin of cultivated rice: based on the interspersion pattern of SINEs. Mol Biol Evol 20:67–75
Choi JY, Platts AE, Fuller DQ, Wing RA, Purugganan MD (2017) The rice paradox: multiple origins but single domestication in Asian rice. Mol Biol Evol 34(4):969–979
Civáň P, Brown TA (2017) Origin of rice (Oryza sativa L.) domestication genes. Genet Resour Crop Evol 6:1125–1132
Civáň P, Craig H, Cox CJ, Brown TA (2015) Three geographically separate domestications of Asian rice. Nat Plants 1(11):1–5
Civáň P, Ali S, Batista-Navarro R, Drosou K, Ihejieto C, Chakraborty D, Ray A, Gladieux P, Brown TA (2019) Origin of the aromatic group of cultivated rice (Oryza sativa L.) traced to the Indian subcontinent. Genome Biol Evol 11(3):832–843
Das S, Khanda CM (2020) Origin and evolution of aus type fragrant rice (Oryza sativa L.): a review. Oryza 57(3):169–180
Deakin JE, Potter S, O’Neill R, Ruiz-Herrera A, Cioffi MB, Eldridge MD, Fukui K, Marshall Graves JA, Griffin D, Grutzner F, Kratochvíl L (2019) Chromosomics: bridging the gap between genomes and chromosomes. Genes 10(8):627
Deb D (2005) Seeds of tradition, seeds of future. Research foundation for science, technology & ecology (RFSTE/ Vrihi). New Delhi
Doležel J, Lucretti S, Molnár I, Cápal P, Giorgi D (2021) Chromosome analysis and sorting. Cytometry A 99(4):328–342
Fukui K (1996) Plant chromosomes at mitosis. In: Fukui K, Nakayama S (eds) Plant chromosomes: laboratory methods. CRC Press Inc., Boca Raton, pp 1–17
Garris AJ, Tai TH, Coburn J, Kresovich S, McCouch S (2005) Genetic structure and diversity in Oryza sativa L. Genetics 169(3):1631–1638
Glaszmann JC (1987) Isozymes and classification of Asian rice varieties. Theor Appl Genet 74(1):21–30
Goff SA (1999) Rice as a model for cereal genomics. Curr Opin Plant Biol 2(2):86–89
Goff SA, Ricke D, Lan TH, Presting G, Wang R, Dunn M, Glazebrook J, Sessions A, Oeller P, Varma H, Hadley D (2002) A draft sequence of the rice genome (Oryza sativa L. ssp japonica). Science 296(5565):92–100
Griglione A, Erica L, Chiara C, Davide B, Cecilla C, Patrizia R, Carlo B, Barbara S (2015) High-quality Italian rice cultivars: chemical indices of ageing and aroma quality. Food Chem 172:305–313
Guerra M (2000) Patterns of heterochromatin distribution in plant chromosomes. Genet Mol Biol 23:1029–1041
Hizume M (2015) Fluorescent banding pattern in chromosomes of Tsuga forrestii and T. sieboldii Pinaceae. Chromosom Bot 10(3):95–100
Jha TB (2021) Karyotype diversity in cultivated and wild Indian rice through EMA-based chromosome analysis. J Genet 100(2):1–5
Jha TB, Bhowmick BK (2021) Unravelling the genetic diversity and phylogenetic relationships of Indian Capsicum through fluorescent banding. Genet Resour Crop Evol 68(1):205–225
Jha TB, Halder M (2016) Searching chromosomal landmarks in Indian lentils through EMA-based Giemsa staining method. Protoplasma 253(5):1223–1231
Jha TB, Saha PS (2017) Characterization of some Indian Himalayan capsicums through floral morphology and EMA-based chromosome analysis. Protoplasma 254(2):921–933
Jha TB, Saha PS (2022) Evaluation of morphological traits, fluorescent banding and rDNA ITS sequences in cultivated and wild Indian lentils (Lens spp.). Genet Resour Crop Evol 69(1):349–362
Jha TB, Saha PS, Nath S, Das A, Jha S (2017) Morphological and cytogenetical characterization of “Dalle Khursani”: a polyploid cultivated capsicum of India. Sci Hortic 215:80–90
Kato S (1928) On the affinity of rice varieties as shown by fertility of hybrid plants. Bull Sci Fac Agric Kyushu Univ 3:132–147
Khush GS (2000) Taxonomy and origin of rice. In: Singh RK, Singh US, and Khush GS (eds) Aromatic rices. New Delhi: Oxford and IBH Publishing Co. Pvt. Ltd, pp 5–13
Konak MA, Hasancebi S, Beser N (2021) Morphological and molecular evaluation of Turkish rice (Oryza sativa L.) landraces. Genet Resour Crop Evol 68(6):2367–2377
Kondo T, Hizume M (1982) Banding for the chromosomes of Cryptomeria japonica D. Don J Jpn for Soc 64:356–358
Kou Y, Liao Y, Toivainen T, Lv Y, Tian X, Emerson JJ, Gaut BS, Zhou Y (2020) Evolutionary genomics of structural variation in Asian rice (Oryza sativa) domestication. Mol Biol Evol 37(12):3507–3524
Kurata N, Omura T (1978) Karyotype analysis in rice a new method for identifying all chromosome pairs. Japan J Genet 53(4):251–255
Kuwada Y (1910) A Cytological study of Oryza sativa L. with plate VIII. Shokubutsugaku Zasshi 24(287):en267-81
Li JQ, Shahid MQ, Feng JH, Liu XD, Zhao XJ, Lu YG (2012) Identification of neutral alleles at pollen sterility gene loci of cultivated rice (Oryza sativa L.) from wild rice (O. rufipogon Griff). Plant Syst Evo 298(1):33–42
Mao AA, Hynniewta TM, Sanjappa M (2009) Plant wealth of Northeast India with reference to ethnobotany. Indian J Tradi Know 8:96–103
Matsuo T (1997) Origin and differentiation of cultivated rice. In: Matsuo T, Futsuhara Y, Kikuchi F, Yamaguchi H (eds) Science of the rice plant, vol 3. Food and agriculture policy research center, Tokyo, pp 69–88
Mehta S, Singh B, Dhakate P, Rahman M, Islam MA (2019) Rice, marker-assisted breeding, and disease resistance. In: Wani SH (ed) Disease resistance in crop plants. Springer, Cham, pp 83–111
Mishra R, Joshi RK, Zhao K (2018) Genome editing in rice: recent advances, challenges, and future implications. Front Plant Sci 9:1361
Moscone EA, Lambrou M, Ehrendorfer F (1996) Fluorescent chromosome banding in the cultivated species of Capsicum (Solanaceae). Plant Syst Evol 202(1):37–63
Muthuramalingam P, Jeyasri R, Rakkammal K, Satish L, Shamili S, Karthikeyan A, Valliammai A, Priya A, Selvaraj A, Gowri P, Wu QS (2022) Multi-Omics and integrative approach towards understanding salinity tolerance in rice: a review. Biology 11(7):1022
Pachauri V, Singh MK, Singh AK, Singh S, Shakeel NA, Singh VP, Singh NK (2010) Origin and genetic diversity of aromatic rice varieties, molecular breeding and chemical and genetic basis of rice aroma. J Plant Biochem Biotechnol 19(2):127–143
Qaisar U, Tayyeb A, Bhat TA (2017) Techniques of chromosomal studies. In: Bhat TA, Wani AA (eds) Chromosome structure and aberrations. Springer, New Delhi, pp 307–330
Second G (1982) Origin of the genic diversity of cultivated rice (Oryza spp.): Study of the polymorphism scored at 40 isozyme loci. Jpn J Genet 57(1):25–57
Sharma GR, Manda D (1980) Excavations at mahagara 1977–1978. A neolithic settlement in belan valley. Archeology of the Vindhyas and Ganga Valley, 6. Dept. of Ancient History, Culture and Archeology, Univ. of Allahabad, India
Sharma A, Sandhya AS, Singh S, Mishra S, Mohan S, Chhavi AS, Singh AK, Jaiswal HK (2021) Aromatic rice of India: It’s types and breeding strategies. In: Huang M (ed) Integrative advances in rice researc. Intech Open. https://doi.org/10.5772/intechopen.99232
Shetty PK, Hegde MR, Mahadevappa M (2013) Innovations in rice production (NIAS Books and Special Publications No. SP1–2013). NIAS, Bangalore. ISBN 978-81-87663706
Sokal RR, Rohlf FJ (1995) Biometry: the principles and practice of statistics in biological research, 3rd edn. Freeman & Co., New York
Sweeney M, McCouch S (2007) The complex history of the domestication of rice. Ann Bot 100(5):951–957
Xie X, Du H, Tang H, Tang J, Tan X, Liu W, Li T, Lin Z, Liang C, Liu YG (2021) A chromosome-level genome assembly of the wild rice Oryza rufipogon facilitates tracing the origins of Asian cultivated rice. Sci China Life Sci 64(2):282–293
Xiong W, Reynolds M, Xu Y (2022) Climate change challenges plant breeding. Curr Opin Plant Biol 70:102308
Yadav S, Kumar V (2018) Feeding the world while caring for the planet. Direct Seeded Rice Consor 1(2):1–18
Yamamoto M, Terakami S, Yamamoto T (2015) Enzymatic maceration/air-drying method for chromosome observations in the young leaf of pear (Pyrus spp.). Chromosom Sci 18(1–2):29–32
Yamamoto M, Takeuchi M, Nashima K, Yamamoto T (2019) Enzyme maceration, fluorescent staining, and FISH of rDNA of pineapple (Ananas comosus (L.) Merr.) chromosomes. Hortic J 88(4):455–461
Yu J, Hu S, Wang J, Wong GK, Li S, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, Cao M (2002) A draft sequence of the rice genome (Oryza sativa L. ssp. indica). Science 296(5565):79–92
Yu S, Ali J, Zhang C, Li Z, Zhang Q (2020) Genomic breeding of green super rice varieties and their deployment in Asia and Africa. Theor Appl Genet 133(5):1427–1442
Zaghum MJ, Ali K, Teng S (2022) Integrated genetic and omics approaches for the regulation of nutritional activities in rice (Oryza sativa L.). Agric 12(11):1757
Zhou X, Huang X (2019) Genome-wide association studies in rice: how to solve the low power problems? Mol Plant 12(1):10–12
Acknowledgements
TBJ acknowledges Usha Gram Trust and Prof. S. Bhattacharyya BCKV, Nadia, West Bengal, Mr. R. Kole of Hooghly West Bengal and Dr. P. K. Das for providing Rice cultivars. Principal Dr. S. Dutta, and Dr. D. Mukhopadhyay, Head, Dept. of Botany Maulana Azad College is duly acknowledged for providing all basic facilities.
Funding
The authors declare that no funds, grants, were received.
Author information
Authors and Affiliations
Contributions
TBJ and MH have contributed equally. Both the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jha, T.B., Halder, M. Evaluation of karyotype diversity in Indian traditional aromatic rice cultivars through EMA-based non-fluorescent Giemsa and fluorescent DAPI staining. Genet Resour Crop Evol 71, 1379–1400 (2024). https://doi.org/10.1007/s10722-023-01696-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01696-4