Abstract
Banana (Musa spp.) production worldwide is seriously threatened by Fusarium wilt caused by Fusarium oxysporum f. sp. cubense (Foc). The best way to control this disease is to grow resistant cultivars. However, it requires large-scale field evaluations and labor- and time-consuming to obtain disease-resistant germplasm. Development of early, reliable, and reproducible selection strategies are considered as the efficient approach which could speed up the selection procedure. In this study, two pairs of sequence-related amplified polymorphism (SRAP) primers related to banana Foc resistance/susceptibility were screened from 100 pairs of random primers. Correspondingly, two pairs of sequence characterized amplified region (SCAR) markers (namely SC4-F/SC4-R and SC14-F/SC14-R, respectively) were successfully generated from these two SRAP markers using 30 cultivars either resistant or susceptible to Foc. Both SCAR markers were located in mitochondrion genome and showed discriminatory power of 96.67% and 100%, respectively. Mitochondrial proteins possibly play a very important role in banana resistance to Foc. In additional, these two SCAR markers were employed simultaneously to screen potential resistant germplasm from 53 accessions with unknown resistance to Foc, and the results revealed a consistency of 83.0% with each other, further indicating their high reliability and reproducibility. These results suggest that both SCAR markers could be used in molecular marker-assisted selection for banana germplasm resistant to Fusarium.
Similar content being viewed by others
Introduction
Banana (Musa spp.) is one of the most important fruit and food crops in the world, which is cultivated in more than 130 countries worldwide, with annual production of more than 100 Million Metric Tons . China is the second largest producer with annual production of more than 10 Million Metric Tons in recent years (FAOSTAT 2023). In Africa, banana is a staple food for millions of people (Dotto et al. 2018). Banana is rich in nutrients (such as starch, vitamins and minerals/K) and plays a very important role in food security (Dotto et al. 2018), food industry (Kumar et al. 2019; Martins et al. 2019; Lau et al. 2020) and some other aspects, such medicine (Vilhena et al. 2018) and industrial wastewater treatment (Mouiya et al. 2019).
However, Fusarium wilt, a soil-borne disease caused by Fusarium oxysporum f. sp. cubense (Foc), is threatening banana industry and the plantations of the producing regions worldwide (Ploetz 2015; Koberls et al. 2017; Mostert et al. 2017; Staver et al. 2020). Foc is classified into several physiological races. Among them, race 1 (termed as Foc1) affects ‘Gros Michel’ (Musa spp. AAA), ‘Silk’ (Musa spp. AAB) and ‘Pisang Awak’ (Musa spp. ABB), race 2 (Foc2) attacks ‘Bluggoe’ (Musa spp. ABB), while race 4 (Foc4) is virulent not only on Cavendish (Musa spp. AAA) but also on Foc1- and Foc2-susceptible varieties (Ploetz 2015). This pathogen infects banana roots through the wound and causes wilt symptom (yellowing of old leaves, withering and even death of plants) through production of gums and gels in the xylem (Ploetz 2015) and causes heavy yield losses among susceptible bananas worldwide (Kumar et al. 2010; Scheerer et al. 2018; Staver et al. 2020; Heck et al. 2021).
Unfortunately, there are no known effective chemical fungicides and biological control agents for this disease. The efficiency of cultural methods is also limited. The best way to control this disease is growing resistant cultivars (Ploetz 2006). However, conventional breeding method has been restricted by several factors, such as the parthenocarpy nature of most cultivars and long growth cycle resulting into long breeding cycle of estimated 10–17 years (Nyine et al. 2018). Selection of disease-resistant germplasm, whatever they are from field screening, or mutation breeding, requires large-scale field evaluations and is labor- and time-consuming (Javed et al. 2004) due to its gigantic size, and long-life cycle.
Molecular markers are very efficient tools in crop improvement due to their functions in linkage mapping, genetic analysis, germplasm characterization and fingerprinting, as well as molecular marker-assisting breeding (Kalia et al. 2017; Kengo 2019; Jian et al. 2021; Maleita et al. 2021; Park et al. 2022; Chen et al. 2023; Habe et al. 2023). These molecular markers include inter-simple sequence repeat (ISSR), random amplified polymorphic DNA (RAPD), sequence characterized amplified regions (SCARs), sequence-related amplified polymorphism (SRAP), single nucleotide polymorphism (SNP), simple sequence repeat (SSR) and so on. Development of early, reliable, and reproducible selection strategies are considered as the efficient approach which could speed up the selection procedure and, eventually, be benefit for banana breeding and the control of banana Fusarium wilt (Javed and Othman 2005). RAPD markers were the first molecular markers successfully used to identify banana resistant germplasm to Foc from susceptible one (Javed et al. 2004; Zambrano et al. 2007; Prasad et al. 2018). But its application is limited due to their drawbacks, such as their dominance nature and low reproducibility (Arinaitwe et al. 2019). SSR and ISSR markers have also been used to distinguish the resistant banana germplasm from the susceptible ones, however they are not successful (Li et al. 2012; Rebouças et al. 2018). No SNP markers involved in the resistance of banana to Foc have been developed successfully yet (Arinaitwe et al. 2019).
SCAR markers are defined as DNA fragments amplified by the PCR using specific 15–30 bp primers, based on the nucleotide sequences generated from cloned RAPD and ISSR fragments related to a trait of interest (Nadeem et al. 2018). When compared to RAPD and ISSR, SCAR markers owe some advantages, such as less sensitive to amplification conditions due to its primer size and hence are more specific and reproducible (Joshi and Chavan 2012). In banana, only three SCAR markers associated with the susceptibility of banana to Foc4 have been developed previously (Wang et al. 2012, 2018). However, there are no SCAR markers related Foc4 have been applied successfully in Foc4-resistant banana germplasm screening.
In the present study, we have developed two SCAR markers with the same set of resistant and susceptible cultivars (7 resistant cultivars/lines, 23 susceptible ones), and the discriminatory power was 96.7% for SC4-F/SC4-R and 100% for SC14-F/SC14-R, suggesting their high reliability. To further confirm their reliability and reproducibility, potential resistant banana germplasm were identified with combined utilization of these two SCAR markers from 53 accessions from cultivars either resistant or susceptible to Foc4, and the results show that the consistency between these two SCAR markers was 83.0%.
Materials & methods
Plant materials used for the development of molecular markers
A total of 30 banana cultivars/mutants belonging to Musa spp. AAA Cavendish were selected as plant materials to generate SCAR markers related banana resistance/susceptibility to Foc4. These 30 cultivars/mutants and their degrees of resistance or susceptibility to Foc4 were shown in Table S1.
DNA preparation
The cetyltrimethylammonium ammonium bromide (CTAB) method (Murray et al. 1980) was employed to extract the DNA from the 30 banana cultivars/mutants. After determination the quality and concentration of DNA with a nucleic acid protein analysis (Eppendorf BioPhotometer Plus, Germany), the concentration was adjusted to 50 ng/μL with ddH2O before use. The size and the integrity of the genomic DNA fragment were tested by gel electrophoresis on 1.0% (w/v) agarose (2 µL DNA + 1 µL 6 × loading buffer + 4 µL ddH2O, 120 V, 15–20 min) followed by visualized on a UV transilluminator and photographed.
Cloning and sequencing of SRAP products
To get SRAP products for further development of SCAR markers associated with banana resistance/susceptibility to Foc4, 100 pairs of random primers (each containing the original 10 bases) were synthesized, and five resistant (‘Nongke 1’, ‘Nantianhuang’, ‘Dongjiao 1’, bxm51, and kangku1) and susceptible cultivars/mutants (g30, ‘Dongguan Zhongba’, ‘Qiwei’, ‘Baxijiao’ and ‘Beida Ai’) were selected as the plant materials. These primers are designed according the rules described by Li et al. (2001) and obtained from Sangon Biotech (Shanghai) Co., LTD (Table S2). PCR amplifications ran in a Bio-Rad PCR (Bio-Rad T100, Bio-Rad, Hercules, CA, USA) according to the manufacturers’ instructions. The total reaction mixture was referred to Table S3 and the conditions for reaction were mentioned in Table S4.
To get the SRAP products, the PCR products (8–10 μL) were fractionated by gel electrophoresis on 1.8% (w/v) agarose gels in 0.5 × Tris–borate-EDTA (TBE) buffer (pH 8.0) after loaded with 4 μL of 6 × loading buffer. Gel was ran at 120 V for approximately 40 min and then visualized using a UV transilluminator and photographed with gel imaging system (Guangzhou Ewell Bio-Technology Co., Ltd., Tanon 4120, CHN) after staining with Goldview. After excised from agarose gels, the PCR products were purified by Tiangel Midi purification kit (Tiangen, CHN) according to the manufacturers’ instructions. To obtain the fragment information of the linked SRAP (the PCR products), they were sent to Sangon Biotech (Shanghai) Co., LTD for sequencing.
Generation of SCAR markers
The SRAP sequences obtained were analyzed by BioXM2.7 software, followed by blasting in NCBI and http://banana-genome-hub.southgreen.fr/blast. Primer3.0 software was employed to design primers for the development SCAR markers from SRAP sequences (Table S5 and Table S6). DNA samples and PCR reaction mixture were the same as that for random markers mentioned above. The reactions were programmed according to Table S7 and Table S8 for SC4-F/SC4-R and SC14-F/SC14-R, respectively. The method for gel electrophoresis of the amplified products was the same as that for SRAP products described above.
Validation of SCAR markers
The SCAR markers obtained were validated firstly with four resistant and susceptible cultivars/mutants, respectively. The additional 22 cultivars/mutants showing different degrees of resistance to Foc4 (Table 1) was employed to validate the reliability of the SCAR markers using the pair of primers with single band after amplification. The PCR reaction system is shown in Table S3 while the conditions for PCR reactions were described above as for the validation of SCAR markers in Table S7 and Table S8.
Molecular marker assistant selection of banana germplasm resistant to Foc4
These two SCAR markers developed in the present study were employed to screen potential resistant banana germplasm from the same group of new accessions (in total 53, of which 21 are natural mutants or mutated from resistant cultivars, 29 from susceptible ones, while the origin of the other three ones was unknown). The leaf from young seedling was used to extract DNA.
Results
DNA quality evaluation
The quality of the DNA extracted from banana leaves was checked with BioDrop μLite (Holliston, MA, USA). The result showed that the concentration of DNA samples was more than 400 ng/μL, the OD260/OD280 ratio for each sample was in the range of 1.7–2.0, indicating they could be used for further experiment. Fig. S1 showed DNA extracted from some representative samples.
SRAP molecular markers related to Foc4 resistance/susceptibility
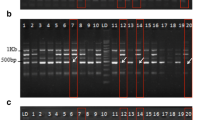
After amplification with a primer set Me1-Em2, a special band of approximately 800 bp appeared in the five susceptible cultivars/mutants (g30, ‘Dongguan Zhongba’, ‘Qiwei’, ‘Baxijiao’ and ‘Beida Ai’) (Fig. 1a). But this was not the case for the five resistant ones (‘Nongke1’, ‘Nantianhuang’, ‘Dongjiao 1’, bxm51 and kangku1). Similarly, a special band of approximately 400 bp was only present in the five susceptible cultivars/mutants after amplified with primer set Me9-Em1 (Fig. 1b). These two pairs of primers were potentially related to the susceptibility of banana to Foc4.
The PCR products amplified with random primer sets in 10 banana (Musa spp. AAA) cultivars/mutants either resistant or susceptible to Foc4. a Primer set Me1-Em2; b Primer set Me9-Em1. M: 2000 bp marker; lanes 1–5: resistant cultivars/mutants; lanes 6–10: susceptible cultivars/mutants; Foc4: Fusarium oxysporum f. sp. cubense race 4; Arrows indicate the specific bands related to disease susceptibility
The PCR products from both Me1-Em2 and Me9-Em1 primer sets mentioned above were cut and purified followed by sequencing. The sequences from ‘Baxijiao’ were selected as the representative (the sequences from different cultivars/mutants were nearly the same) and shown in Fig. 2a (from primer set Me1-Em2) and Fig. 2b (from primer set Me9-Em1), respectively.
Development of SCAR markers from SRAP ones
For the development of a SCAR marker, four pairs of primers (Table S5) were designedfor the PCR product from primer set Me1-Em2 using DNA from eight banana cultivars showing resistance or susceptibility to Foc4 as the template. Another four pairs of primers (Table S6) were designed for the PCR product from primer set Me9-Em1.
A single, clear band of 251 bp or 295 bp was present only in the four susceptible cultivars/mutants (‘Dongguan Zhongba’, ‘Qiwei’, ‘Baxijiao’ and ‘Beida Ai’) but not in the other four resistant ones (‘Nongke 1’, ‘Nantianhuang’, ‘Dongjiao 1’, and bxm51) after amplified with primer set SC4-F/SC4-R (Fig. 3a) or SC14-F/SC14-R (Fig. 3b), respectively. Unfortunately, there were no any assignments with these two fragments when blasted in NCBI, but this was not the case when blasted in Banana Genome Hub database (http://banana-genome-hub.southgreen.fr/blast). With all available Musa genome sequences, we employ the BLAST program to find potential locations of these two markers. Interestingly, both markers seem to locate in the mitochondrion genome. The 251 bp fragment from SC4-F/SC4-R is located in the position from 72 to 322 bp of the original SRAP band and showed a 99.60% similarity to a fragment of 250 bp located in chrUn_random (from 6,569,491 to 6,569,741) of the M. acuminate Pahang (v2) and in the mitochondrion genome (from 2,709,912 to 2,710,162) of the M. acuminate Pahang (v4). Differently, the 295 bp fragment from SC14-F/SC14-R is imperfectly matched to multiple regions of the mitochondrion genome. It was located in the position 15 to 309 bp of the original SRAP band and showed only a 71.53% similarity to a fragment of 211 bp located in Contig78419 of the M. acuminate calcutta4 (v1).
The PCR products amplified from banana cultivars/mutants (Musa spp. AAA) either resistant or susceptible to Foc4 with SCAR primer sets. a SC4-F/SC4-R, 251 bp; b SC14-F/SC14-R, 295 bp; M: 2000 bp marker; lanes 1–4: resistant cultivars/mutants; lanes 5–8: susceptible cultivars/mutants; Foc4: Fusarium oxysporum f. sp. cubense race 4; and SCAR: sequence characterized amplified region
To test the reliability of those two SCAR markers mentioned above, they were used to distinguish the additional 22 cultivars/mutants with known resistance or susceptibility to Foc4. As shown in Fig. 4 and Table 1, both SCAR markers were able to discriminate the resistance from susceptibility of banana to Foc4. An unexpected, weak band only appeared in one (kangku5) out of the 30 cultivars/mutants after amplified with SC4-F/SC4-R. The discriminatory power was 96.67% for SC4-F/SC4-R while it was 100% for SC14-F/SC14-R. These results suggest that the two SCAR markers developed in the present study can be applied in molecular marker-assisted selection for banana germplasm resistant to Fusarium.
Validation of SCAR markers in 23 banana cultivars (Musa spp. AAA) either resistant or susceptible to Foc4. a SC4-F/SC4-R, 251 bp; b SC14-F/SC14-R, 295 bp; M: 2000 bp marker; lanes 1–4 and 23: resistant cultivars/mutants; lanes 5–22: susceptible cultivars/mutants; Foc4: Fusarium oxysporum f. sp. cubense race 4; SCAR: sequence characterized amplified region. * ‘Nongke 1’ is the cultivar also among the eight ones in Fig. 3
Identification of potential resistant germplasm with two SCAR markers
In the present study, two SCAR markers were employed to screen potential Foc resistant banana germplasm from 53 accessions (50 from resistant or susceptible cultivars, 3 from resistance-unknown accessions). SC4-F/SC4-R revealed that 17 accessions were identified as resistant germplasm while the other 36 as susceptible ones, and it was 16 resistant accessions and 37 susceptible ones for SC14-F/SC14-R (Fig. 5, Table 2). Forty-four out of the 53 tested accessions revealed the same resistance to Foc between these two SCAR markers and the consistence was 83.0% (it was 88.0% when the 30 cultivars/mutants used for the development of SCAR markers were included in). These results further confirmed the reliability and reproducibility of the two SCAR markers developed in the present study.
Screening of potential resistant gerplasm from 53 banana accessions (Musa spp. AAA Cavendish) from cultivars either resistant or susceptible to Foc4. Resistant-origin: 5 (nk4), 6 (rk5), 7 (reke1), 14–17, 21 (B40-Yueyoukang 1), 22 (B20-Yueke), 23 (B40-Yueke), 24–25 (B20-Reyan1), 26 (B30-Reyan 1), 27–28 (B20-Nantianhuang), 29 (B30-Nantianhuang), 30 (k1j1), 38 [l1n1(1)], 40 (l3-3), 44 (l3), 49 (r1-2–1); Susceptible-orgin: 1 (hongyan 3 ny3), 2 (‘Hongyan 2’), 3 (‘Hongyan 3’), 4(‘Hongyan 5’), 8 (‘Reke2’), 9–11 (2–21), 12–13 (1–10), 18–20 (B40-BX), 31 (bx1-1 nai), 32 (b23-1), 33 [g20k71(1)], 34,36 (g20k72), 37 (g20-11–71), 39 (g20k71), 41 (g20-1 k); 42 (rk2), 43 (rk3), 45 [g20k71(2)], 46 (g20-1–1), 50 (wghd3), 51 (wghd 51), 52 (wghd 52), 53 (wghd); Unknown origin: 35 (wgh), 47 (hb1), 48 (hb3)
In the present study, most tested accessions kept the resistance or susceptibility from their mother plants, but four susceptible-origin accessions (bx1-1 nai, hongyan 3 hy3, ‘Hongyan 5’, and ‘Reke 2’) were identified as resistant germplasm with both SCAR markers. On the contrary, nine resistant-origin accessions [l3, l3-3, r1-2-1, nk4, B40-Yueyoukang 1 (1–5)] were identified as susceptible ones. In total, two newly obtained cultivars, four mutants and 6 offsprings generated directly from mutated cultivars were identified as resistant germplasm by both SCAR markers developed in the present study (Fig. 5, Table 2).
Discussion
It is estimated that by 2040, only Foc tropical race 4 (Foc TR4) might spread to 17% of the world’s banana production regions and the yield is estimated as 36 million tons which is worth over US$10 billion (Staver et al. 2020). A banana breeding program aiming to control this devastating disease is extremely important and necessary. Nevertheless, traditional hybrid breeding strategy is not suitable for Musa AAA cultivars due to parthenocarpy. Some other strategies, such as somaclonal variants (Hwang and Ko 2004), field tested in more tropical regions (Molina et al. 2016), and mutation breeding (Kishor et al. 2018) have been employed commercially. However, they are time-consuming and costly due to the necessity of verifying the results by field evaluation over an extended period (Javed et al. 2004). Furthermore, there is a risk in spread of the pathogen when field disease resistance test is carried out. Thus, the development of rapid and reliable methods to screen resistant germplasm at earlier developmental stages would be of great interest for the banana breeding against Foc (Collard and Mackill 2008). Molecular markers associated with the resistance or susceptibility fit this requirement very well due to their several advantages, such as accuracy, stability and effectiveness of operation (Park et al. 2022). Most importantly, molecular-assisted breeding only needs a sample of less than 0.5 g from a new germplasm, instead of a large quantity of offsprings from the new germplasm as used in conventional disease resistance test. This will greatly shorten the breeding cycle.
Till today, several types of molecular markers, e.g. RAPD, SSR and ISSR, have been employed in studing the resistance/susceptibility of bananas to Fusarium wilt. For example, the genetic diversity of wild Musa acuminata ssp. Malaccensis in Malaysia showing varying degrees of resistance to Foc TR4, was assessed with RAPD markers by Javed et al. (2004). The authors found that a band specific to either resistant or susceptible seedlings could be obtained from three primers (namely primer OPA-03, 24 and 21), respectively. A RAPD marker named OPK-03 was identified as capable of distinguishing the susceptible cultivar from the resistant ones, because it could amplify a band of 485 bp only in the susceptible banana genotype (Zambrano et al. 2007).Recently, Kishor et al. (2018) have also identified two RAPD primers, namely OPN-06 and OPR-07, to screen potential mutants resistant to Foc1. However, the application of RAPD markers in molecular-assisted breeding has been restricted due to their disadvantages. Firstly, the PCR products from RAPD primers often have more than one band after amplification of multiple gene loci. Secondly, they have relatively lower reproducibility when compared to some other molecular markers. The amplification reactions for RAPD markers require highly standardized experimental protocols because they are very sensitive to the reaction conditions (Jarocki et al. 2016). Moreover, RAPD markers generally require DNA in relatively higher quality and at higher molecular weight (Athe et al. 2018). SSR markers owe some advantages over other molecular markers, such as RAPD and SCAR. These include the multi-allelic and co-dominant natures of the genome providing wide coverage as well as high reproducibility these features enable them to distinguish homozygotes from heterozygotes (Athe et al. 2018). However, till today, the SSR markers have been able to detect genetic diversity but could not differentiate between banana genotypes susceptible to Foc4 from those that are resistant (Rebouças et al. 2018). Similar results were observed with ISSR markers in relation to Fusarium wilt of banana (Li et al. 2012), though SSR and ISSR markers have been utilized successfully in studying Fusarium wilt resistance in some other crops (Anjani et al. 2018; Mahmoud and Abd El-Fatah 2020).
SCAR markers, in contrast to RAPDs, are dominant and could detect a single locus. In banana, Cunha et al. (2015) have successfully generated a SCAR marker associated with the susceptibility of banana to Foc1 from a RAPD primer OPP-12. The PCR products from this marker were present only in the 20 tested susceptible banana cultivars but not in the resistant ones. Wang et al. (2012) have successfully converted two RAPD primers into SCAR markers (ScaU1001 and ScaS0901), which could distinguish bananas resistant and susceptible to Foc4. However, the cultivar number for the development of these SCAR markers was only five, which might result in low reliability and reproducibility. Moreover, one of these two markers was identified as not efficient (Silva et al. 2016). More cultivars showing different resistance to the pathogen used in developing molecular markers will be benefit for their reliability and reproducibility. Wang et al. (2018) have also successfully developed one SCAR marker linked to Foc4 susceptibility but without application of this marker in resistant banana germplasm screening. In the present study, two SCAR markers associated with banana susceptibility to Foc4 were successfully generated from SRAP markers using 30 cultivars/mutants that were either resistant or susceptible to Foc4. We used both SCAR markers to screen 53 banana germplasm accessions from resistant and susceptible cultivars. The results were highly consistent, confirming the marker’s reliability and reproducibility. The combined use of these two SCAR markers will accelerate banana breeding against Foc4.
Two SCAR markers obtained in the present study were predicted to be located in mitochondrion genome. Similar result was observed by Wang et al. (2018). Interestingly, we have analyzed available sequences amplified from banana Foc1 or Foc4 resistant linked markers in previous studies (Wang et al. 2012; Cunha et al. 2015) and found that two out of three were also present in mitochondrion genome. These findings indicated that mitochondrion DNA plays a crucial role in in banana resistance to Foc. Molecular markers linked to several aspects, including disease resistance, have been successfully generated from mitochondrial DNA (mtDNA) of some other plants (Ding et al. 2021; Murali et al. 2021) and many evidences have already demonstrated that mitochondrial proteins contributed to plant resistance/tolerance to biotic and abiotic stresses (Robles et al. 2018; Kim et al. 2021; Qiu et al. 2021; Meng et al. 2022).
In the present study, the two SCAR markers developed do not exhibit any annotated functional genes in adjacent regions (upstream 100 kb and downstream 100 kb). This is, at least partially, due to the fact that no complete banana mitochondrial chromosome data is currently available. More transcriptome data might be helpful in verifying the direct association of this marker with coding genes. Another alternative explanation is that the locus of marker SC4-F/SC4-R functions in cis. For better understanding the functions of mitochondrial proteins, further work should be focused on assembly and analysis of the complete mitochondrial genome sequences of banana cultivars either resistant or susceptible to Foc4. We believe that Pacbio hifi-sequencing will play a crucial role in resolving this problem in the near future.
Conclusions
In the present study, two reliable SCAR markers (SC4-F/SC4-R and SC14-F/SC14-R) were developed from SRAP markers successfully using 30 cultivars/mutants showing varying resistance to Foc. In addition, we have combined the use of them to screen potential resistant banana germplasm and have got highly consistent results. This method could screen large amount of mutants or any other newly obtained germplasm rapidly and at low cost. We are sure the combination application of these two SCAR markers could further improve the reliability and reproducibility of identification, and finally greatly speed up banana breeding against Foc4.
References
Anjani K, Ponukumatla B, Mishra D, Ravulapalli DP (2018) Identification of simple-sequence-repeat markers linked to Fusarium wilt (Fusarium oxysporum f. sp carthami) resistance and marker-assisted selection for wilt resistance in safflower (Carthamus tinctorius L.) interspecific offsprings. Plant Breed 137(6):895–902. https://doi.org/10.1111/pbr.12636
Arinaitwe IK, Teo CH, Kayat F, Tumuhimbise R, Uwimana B, Kubiriba J, Harikrishna JA, Othman RY (2019) Molecular markers and their application in Fusarium Wilt studies in Musa spp. Sains Malays 48(9):1841–1853. https://doi.org/10.17576/jsm-2019-4809-05
Athe R, Naha B, Neerasa G, Parthasarathi PCB, Nukala R, Devara D (2018) Molecular markers-characteristics and applications in animal breeding. Int J Livest Res 8(1):1–7. https://doi.org/10.5455/ijlr.20170424050432
Chen A, Sun J, Viljoen A, Mostert D, Xie Y, Mangila L, Bothma S, Lyons R, Hřibová E, Christelová P, Uwimana B et al (2023) Gene identification and marker validation for host plant resistance to the race 4 of Fusarium oxysporum f. sp. cubense using Musa acuminata ssp. malaccensis. Pathogens 12:820. https://doi.org/10.3390/pathogens12060820
Collard BCY, Mackill DJ (2008) Marker-assisted selection: an approach for precision plant breeding in the twenty-first century. Phil Trans R Soc 363:557–572. https://doi.org/10.1098/rstb.2007.2170
Cunha CMS, Hinz RH, Pereira A, Tcacenco FA, Paulino EC, Stadnik MJ (2015) A SCAR marker for identifying susceptibility to Fusarium oxysporum f. sp. cubense in banana. Sci Hortic 191:108–112. https://doi.org/10.1016/j.scienta.2015.04.038
Ding CQ, Chen CT, Su N, Lyu WH, Yang JH, Hu ZY, Zhang MF (2020) Identification and characterization of a natural SNP variant in ALTERNATIVE OXIDASE gene associated with cold stress tolerance in watermelon. Plant Sci 304:110735. https://doi.org/10.1016/j.plantsci.2020.110735
Dotto J, Matemu AO, Ndakidemi PA (2018) Potential of cooking bananas in addressing food security in East Africa. Int J Biosci 13(4):278–294. https://doi.org/10.12692/ijb/13.4.278-294
FAOSTAT (2023) Available online: http://www.fao.org/faostat/en/#data/QC/visualize Accessed on 20 May 2023
Habe I, Sakamoto Y, Matsumoto K (2023) The development and efficient utilization of molecular markers for the major quantitative trait locus of bacterial wilt resistance in potato. Euphytica 219:68. https://doi.org/10.1007/s10681-023-03187-0
Heck DW, Dita M, Ponte EMD, Mizubuti ESG (2021) Incidence, spatial pattern and temporal progress of Fusarium wilt of bananas. J Fungi 7(8):646. https://doi.org/10.3390/jof7080646
Hwang SC, Ko WH (2004) Cavendish banana cultivars resistant to Fusarium wilt acquired through somaclonal variation in Taiwan. Plant Dis 88(6):580–588. https://doi.org/10.1094/PDIS.2004.88.6.580
Jarocki P, Podleśny M, Komoń-Janczara E, Kucharska J, Glibowska A, Targoński Z (2016) Comparison of various molecular methods for rapid differentiation of intestinal bifidobacteria at the species, subspecies and strain level. BMC Microbiol 16(1):159. https://doi.org/10.1186/s12866-016-0779-3
Javed MA, Othman RY (2005) Characterization of Fusarium wilt-resistant and Fusarium wilt-susceptible somaclones of banana cultivar Rastali (Musa AAB) by random amplified polymorphic DNA and retrotransposon markers. Plant Mol Biol Rep 23(3):241–249. https://doi.org/10.1007/BF02772754
Javed MA, Chai M, Othman RY (2004) Study of resistance of Musa acuminata to Fusarium oxysporum using RAPD markers. Biol Plantarum 48(4):93–99. https://doi.org/10.1023/B:BIOP.0000024281.85427.6d
Jian Y, Yan W, Xu J, Duan S, Li G, Jin L (2021) Genome-wide simple sequence repeat markers in potato: abundance, distribution, composition, and polymorphism. DNA Res 28(6):1–9. https://doi.org/10.1093/dnares/dsab020
Joshi K, Chavan P (2012) Development of sequence characterized amplified region from random amplified polymorphic DNA amplicons. Methods Mol Biol 862:123–134. https://doi.org/10.1007/978-1-61779-609-8_10
Kalia P, Saha P, Ray S (2017) Development of RAPD and ISSR derived SCAR markers linked to Xca1Bo gene conferring resistance to black rot disease in cauliflower (Brassica oleracea var. botrytis L.). Euphytica. https://doi.org/10.1007/s10681-017-2025-y
Kengo O (2019) The Rx gene derived USDA 41956 and Rx1 gene derived CPC 1673 confer equal resistance to the migration of Potato virus X from potato leaves to tubers. Euphytica 215:90. https://doi.org/10.1007/s10681-019-2413-6
Kim M, Schulz V, Brings L, Schoeller T, Kühn K, Vierling E (2021) mTERF18 and ATAD3 are required for mitochondrial nucleoid structure and their disruption confers heat tolerance in Arabidopsis thaliana. New Phytol 232(5):2026–2042. https://doi.org/10.1111/nph.17717
Kishor H, Prabhuling G, Abhijith YC, Manjunatha N, Prasad YP (2018) Early screening of Fusarium wilt and molecular analysis of banana variants. Int J Curr Microbiol Appl Sci 7(03):2313–2321. https://doi.org/10.20546/ijcmas.2018.703.271
Koberls M, Dita M, Martinuz A, Staver C, Berg G (2017) Members of Gammaproteobacteria as indicator species of healthy banana plants on Fusarium wilt infested fields in Central America. Sci Rep 7:45318. https://doi.org/10.1038/srep45318
Kumar B, Shankar U, Nayaka C, Kini SR (2010) Biochemical characterization of Fusarium oxysporum f.sp. cubense isolates from India. Afr J Biotechnol 9(4):523–530. https://doi.org/10.5897/AJB09.1069
Kumar PS, Saravanan A, Sheeba N, Uma S (2019) Structural, functional characterization and physicochemical properties of green banana flour from dessert and plantain bananas (Musa spp.). LWT-Food Sci Technol 116:108524. https://doi.org/10.1016/j.lwt.2019.108524
Lau BF, Kong KW, Leong KH, Sun J, He X, Wang Z, Mustafa MR, Ling TC, Ismail A (2020) Banana inflorescence: its bio-prospects as an ingredient for functional foods. Trends Food Sci Technol 97:14–28. https://doi.org/10.1016/j.tifs.2019.12.023
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103(2–3):455–461. https://doi.org/10.1007/s001220100570
Li X, Peng T, Xu F, Xie J, Chen YTH (2012) Genetic diversity analysis for different wilt disease resistant bananas by ISSR. Guangdong Agri Sci 39(9):6–8. https://doi.org/10.16768/j.issn.1004-874x.2012.09.056
Mahmoud AF, Abd El-Fatah BES (2020) Genetic diversity studies and identification of molecular and biochemical markers associated with Fusarium wilt resistance in cultivated Faba bean (Vicia faba). Plant Pathology J 36(1):11–28. https://doi.org/10.5423/PPJ.OA.04.2019.0119
Maleita C, Cardoso JMS, Rusinque L, Esteves I, Abrantes I (2021) Species-specific molecular detection of the root knot nematode Meloidogyne luci. Biology 10:775. https://doi.org/10.3390/biology10080775
Martins ANA, Pasquali MAD, Schnorr CE, Martins JJA, de Araujo GT, Rocha APT (2019) Development and characterization of blends formulated with banana peel and banana pulp for the production of blends powders rich in antioxidant properties. J Food Sci Tecchnol 56(12):5289–5297. https://doi.org/10.1007/s13197-019-03999-w
Meng XX, Li L, Pascual J, Rahikainen M, Yi CY, Jost R, He CM, Fournier-Level A, Borevitz J, Kangäsjarvi S, Whelan J, Berkowitz O (2022) GWAS on multiple traits identifies mitochondrial ACONITASE3 as important for acclimation to submergence stress. Plant Physiol 188(4):2039–2058. https://doi.org/10.1093/plphys/kiac011
Molina AB, Sinohin VO, Fabregar EG, Ramillete EB, Loayan MM, Chao CP (2016) Field resistance of Cavendish somaclonal variants and local banana cultivars to tropical race 4 of Fusarium wilt in the Philippines. Acta Hortic 1114:227–230. https://doi.org/10.17660/actahortic.2016.1114.31
Mostert D, Molina AB, Daniells J, Fourie G, Hermanto C, Chao CP, Viljoen A (2017) The distribution and host range of the banana Fusarium wilt fungus, Fusarium oxysporum f. sp cubense. Asia. Plos ONE 12(7):e0181630. https://doi.org/10.1371/journal.pone.0181630
Mouiya M, Bouazizi A, Abourriche A, Benhammou A, El Hafiane Y, Ouammou M, Abouliatim Y, Younssi SA, Smith A, Hannache H (2019) Fabrication and characterization of a ceramic membrane from clay and banana peel powder: application to industrial wastewater treatment. Mater Chem Phys 227:291–301. https://doi.org/10.1016/j.matchemphys.2019.02.011
Murali P, Hilda K, Ramakrishnan M, Ganesh A, Bhuvaragavan S, Janarthanan S (2021) Molecular genotypic diversity of populations of brinjal shoot and fruit borer, Leucinodes orbonalis and development of SCAR marker for pesticide resistance. Mol Biol Rep 48(12):7787–7800. https://doi.org/10.1007/s11033-021-06791-2
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8(19):4321–4325. https://doi.org/10.1093/nar/8.19.4321
Nadeem MA, Nawaz MA, Shahid MQ, Doğan Y, Comertpay G, Yıldız M, Ahmad F, Alsaleh A, Labhane N, Özkan H, Chung G, Baloch FS (2018) DNA molecular markers in plant breeding: current status and recent advancements in genomic selection and genome editing. Biotechnol Biotechnol Equip 32(2):261–285. https://doi.org/10.1080/13102818.2017.1400401
Nyine M, Uwimana B, Blavet N, Hřibová E, Vanrespaille H, Batte M, Doležel J (2018) Genomic prediction in a multiploid crop: genotype by environment interaction and allele dosage effects on predictive ability in banana. Plant Genome 11:2. https://doi.org/10.3835/plantgenome2017.10.0090
Park JS, Kang MY, Shim EJ, Oh J, Seo KI, Kim KS, Sim SC, Chung SM, Park Y, Lee GP, Lee WS et al (2022) Genome-wide core sets of SNP markers and Fluidigm assays for rapid and effective genotypic identification of Korean cultivars of lettuce (Lactuca sativa L.). Hortic Res 9:uhac119. https://doi.org/10.1093/hr/uhac119
Ploetz RC (2006) Fusarium wilt of banana is caused by several pathogens referred to as Fusarium oxysporum f. sp. cubense. Phytopathology 96(6):653–656. https://doi.org/10.1094/PHYTO-96-0653
Ploetz RC (2015) Management of Fusarium wilt of banana: a review with special reference to tropical race 4. Crop Protect 73:7–15. https://doi.org/10.1016/j.cropro.2015.01.007
Prasad YP, Manjunatha N, Prabhuling G, Kishor H, Abhijith YC (2018) Early screening of Fusarium wilt and molecular analysis of banana variants. Int J Curr Microbiol Appl Sci. 7(3):2313–2321. https://doi.org/10.20546/ijcmas.2018.703.271
Qiu TC, Zhao XS, Feng HJ, Qi LL, Yang J, Peng YL, Zhao WS (2021) OsNBL3, a mitochondrion-localized pentatricopeptide repeat protein, is involved in splicing nad5 intron 4 and its disruption causes lesion mimic phenotype with enhanced resistance to biotic and abiotic stresses. Plant Biotechnol J 19(11):2277–2290. https://doi.org/10.1111/pbi.13659
Robles P, Navarro-Cartagena S, Ferrández-Ayela A, Núñez-Delegido E, Quesada V (2018) The characterization of Arabidopsis mterf6 mutants reveals a new role for mTERF6 in tolerance to abiotic stress. Int J Mol Sci 19(8):2388. https://doi.org/10.3390/ijms19082388
Scheerer L, Pemsl DE, Dita M, Perez VL, Staver C (2018) A quantified approach to projecting losses caused by Fusarium wilt tropical race 4. Acta Hortic 1196:211–218. https://doi.org/10.17660/ActaHortic.2018.1196.26
Silva PRO, de Jesus ON, Bragança CAD, Haddad F, Amorim EP, Ferreira CF (2016) Development of a thematic collection of Musa spp. accessions using SCAR markers for preventive breeding against Fusarium oxysporum f. sp. cubense tropical race 4. Genet Mol Res 15(1):15017765. https://doi.org/10.4238/gmr.15017765
Staver C, Pemsl DE, Scheerer L, Vicente LP, Dita M (2020) Ex ante assessment of returns on research investments to address the impact of Fusarium wilt tropical race 4 on global banana production. Front Plant Sci 11:844. https://doi.org/10.3389/fpls.2020.00844
Vilhena RD, Fachi MM, Marson BM, Dias BL, Pontes FLD, Tonin FS, Pontarolo R (2018) Antidiabetic potential of Musa spp. inflorescence: a systematic review. J Pharm Pharmacol 70(12):1583–1595. https://doi.org/10.1111/jphp.13020
Wang W, Hu Y, Sun D, Staehelin C, Xin D, Xie J (2012) Identification and evaluation of two diagnostic markers linked to Fusarium wilt resistance (race 4) in banana (Musa spp.). Mol Biol Rep 39(1):451–459. https://doi.org/10.1007/s11033-011-0758-6
Wang F, Xia L, Lv S, Xu C, Niu Y, Liu W, Hu B (2018) Development of a mitochondrial SCAR marker related to susceptibility of banana (Musa AAA Cavendish) to Fusarium oxysporum f. sp. cubense race 4. Not Bot Horti Agrobo. 46(2):509–516. https://doi.org/10.15835/nbha46211053
Zambrano AY, Martínez G, Gutiérrez Z, Manzanilla E, Ruíz L, Díaz AL, Marquez A, Demey JR (2007) Identification of RAPD marker linked to resistance of Musa to Fusarium oxysporum. Acta Hortic 32(11):775–779. https://doi.org/10.17660/ActaHortic.2007.738.69
Funding
This work was supported by the Key Research and Development Program of Guangdong Province (2021B0707010004), the National Key Research and Development Program of China (2018YFD1000300), the Earmarked Fund for Modern Agro-industry Technology Research System (CARS-31-04), the Guangdong Province Special Fund for Modern Agriculture Industry Technology Innovation Teams (2023KJ109), the Rural Revitalization Strategic Special Funds of Dongguan-Guangdong Province (20211800400052).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. M.Q., G.F., S.C., C.C., L.D., H.L. implemented the experiment and analyzed the data; C.X., Z.S., M.Q. and G.F. drafted the original draft; C.X., M.Q. and S.C. reviewed the original draft; C.X., H.C., X.L and F.W. provided funds; Z.S., F.W., S.L. and B.H. provided plant materials. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qv, M., Feng, G., Chen, S. et al. The development and utilization of two SCAR markers linked to the resistance of banana (Musa spp. AAA) to Fusarium oxysporum f. sp. cubense race 4. Euphytica 220, 69 (2024). https://doi.org/10.1007/s10681-024-03323-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-024-03323-4