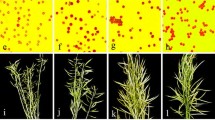
Abstract
Broadening the genetic base of the C genome of Brassica napus canola by use of B. oleracea is important. In this study, the prospect of developing B. napus canola lines from B. napus × B. oleracea var. alboglabra, botrytis, italica and capitata crosses and the effect of backcrossing the F1’s to B. napus were investigated. The efficiency of the production of the F1’s varied depending on the B. oleracea variant used in the cross. Fertility of the F1 plants was low—produced, on average, about 0.7 F2 seeds per self-pollination and similar number of BC1 seeds on backcrossing to B. napus. The F3 population showed greater fertility than the BC1F2; however, this difference diminished with the advancement of generation. The advanced generation populations, whether derived from F2 or BC1, showed similar fertility and produced similar size silique with similar number of seeds per silique. Progeny of all F1’s and BC1’s stabilized into B. napus, although B. oleracea plant was expected, especially in the progeny of F1 (ACC) owing to elimination of the A chromosomes during meiosis. Segregation distortion for erucic acid alleles occurred in both F2 and BC1 resulting significantly fewer zero-erucic plants than expected; however, plants with ≤ 15% erucic acid frequently yielded zero-erucic progeny. No consistent correlation between parent and progeny generation was found for seed glucosinolate content; however, selection for this trait was effective and B. napus canola lines were obtained from all crosses. Silique length showed positive correlation with seed set; the advanced generation populations, whether derived from F2 or BC1, were similar for these traits. SSR marker analysis showed that genetically diverse canola lines can be developed by using different variants of B. oleracea in B. napus × B. oleracea interspecific crosses.







Similar content being viewed by others
References
Abel S, Möllers C, Hecker HC (2005) Development of synthetic Brassica napus lines for the analysis of ‘‘fixed heterosis’’ in allopolyploid plants. Euphytica 146:157–163
Attri R, Rahman H (2018) Introgression of allelic diversity from genetically distinct variants of Brassica rapa into Brassica napus canola and inheritance of the B. rapa alleles. Crop Pasture Sci 69:94–106
Bahrani J, McVetty PBE (2008) Relationship of seed quality traits for greenhouse-grown versus field-grown high erucic acid rapeseed: Is seed quality trait selection for greenhouse-grown seed worthwhile? Can J Plant Sci 88:419–423
Bennett RA, Thiagarajah MR, King JR, Rahman MH (2008) Interspecific cross of Brassica oleracea var. alboglabra and B. napus: effects of growth condition and silique age on the efficiency of hybrid production, and inheritance of erucic acid in the self-pollinated backcross generation. Euphytica 164:593–601
Bennett RA, Séguin-Swartz G, Rahman H (2012) Broadening genetic diversity in canola (Brassica napus L.) using the C-genome species B. oleracea. Crop Sci 52:2030–2039
Bus A, Korber N, Snowdon RJ, Stich B (2011) Patterns of molecular variation in a species-wide germplasm set of Brassica napus. Theor Appl Genet 123:1413–1423
Butruille DV, Guries RP, Osborn TC (1999) Increasing yield of spring oilseed rape hybrids through introgression of winter germplasm. Crop Sci 39:1491–1496
Chen BY, Heneen WK (1989) Fatty acid composition of resynthesized Brassica napus L., B. campestris L. and B. alboglabra Bailey with special reference to the inheritance of erucic acid content. Heredity 63:309–314
Chen S, Nelson MN, Ghamkhar K, Fu T, Cowling WA (2008) Divergent patterns of allelic diversity from similar origins: the case of oilseed rape (Brassica napus L.) in China and Australia. Genome 51:1–10
Cheng X, Xu J, Xia S, Gu J, Yang Y, Fu J, Qian X, Zhang S, Wu J, Liu K (2009) Development and genetic mapping of microsatellite markers from genome survey sequences in Brassica napus. Theor Appl Genet 118:1121–1131
Chiang BY, Grant WF, Chiang MS (1978) Transfer of resistance to race 2 of Plasmodiophora brassicae from Brassica napus to cabbage (B. oleracea var. capitata). II. Meiosis in the interspecific hybrids between B. napus and 2x and 4x cabbage. Euphytica 27:81–93
Cowling WA (2007) Genetic diversity in Australian canola and implications for crop breeding for changing future environments for crop breeding for changing future environments. Field Crop Res 104:103–111
Cui C, Ge X, Gautam M, Kang L, Li Z (2012) Cytoplasmic and genomic effects on meiotic pairing in Brassica hybrids and allotetraploids from pair crosses of three cultivated diploids. Genetics 191:725–738
Diers BW, Osborn TC (1994) Genetic diversity of oilseed Brassica napus germplasm based on restriction fragment length polymorphisms. Theor Appl Genet 88:662–668
dos Santos JB, Nienhuis J, Skroch P, Tivang J, Slocum MK (1994) Comparison of RAPD and RFLP genetic markers in determining genetic similarity among Brassica oleracea L. genotypes. Theor Appl Genet 87:909–915
Falk DE (2010) Generating and maintaining diversity at the elite level in crop breeding. Genome 53:982–991
Friedt W, Snowdon R (2010) Oilseed rape. In: Vollmann J, Rajcan I (eds) Oil crops. Handbook of plant breeding 4. Springer, Dordrecht Heidelberg London New York, pp 91–126
Fu YB, Gugel RK (2010) Genetic diversity of Canadian elite summer rape (Brassica napus L.) cultivars from the pre- to post-canola quality era. Can J Plant Sci 90:23–33
Gyawali S, Hegedus DD, Parkin IAP, Poon J, Higgins E, Horner K, Bekkaoui D, Coutu C, Buchwaldt L (2013) Genetic diversity and population structure in a world collection of Brassica napus accessions with emphasis on South Korea, Japan, and Pakistan. Crop Sci 53:1537–1545
Hasan M, Seyis F, Badani AG, Pons-Kuhnemann J, Friedt W, Lühs W, Snowdon RJ (2006) Analysis of genetic diversity in the Brassica napus L. gene pool using SSR markers. Genet Res Crop Evol 53:793–802
Hasan M, Friedt W, Kühnemann JP, Freitag NM, Link K, Snowdon RJ (2008) Association of gene-linked SSR markers to seed glucosinolate content in oilseed rape (Brassica napus ssp. napus). Theor Appl Genet 116:1035–1049
Hill J, Lethenborg P, Li PW, Rahman MH, Sørensen H, Sørensen JC (2003) Inheritance of progoitrin and total aliphatic glucosinolates in oilseed rape (Brassica napus L). Euphytica 134:179–187
Holm SN, Rahman MH, Stølen O, Sørensen H (1985) Studies on pollination requirement in rapeseed (Brassica campestris). In: Sørensen H (ed) Advances in the production and utilization of cruciferous crops. Martinus Nijhoff Publ, Dordrecht, pp 245–253
Howell PM, Sharpe AG, Lydiate DJ (2003) Homoeologous loci control the accumulation of seed glucosinolates in oilseed rape (Brassica napus). Genome 46:454–460
Izzah NK, Lee J, Perumal S, Park JY, Ahn K, Fu D, Kim G-B, Nam Y-W, Yang T-J (2013) Microsatellite-based analysis of genetic diversity in 91 commercial Brassica oleracea L. cultivars belonging to six varietal groups. Genet Resour Crop Evol 60:1967–1986
Kebede B, Thiagarajah MR, Zimmerli C, Rahman MH (2010) Improvement of open-pollinated spring rapeseed (Brassica napus L.) through introgression of genetic diversity from winter rapeseed. Crop Sci 50:1236–1243
Kondra ZP, Stefansson BR (1970) Inheritance of the major glucosinolates of rapeseed (Brassica napus) meal. Can J Plant Sci 50:643–647
Kubik TJ, Hawkins GP, Stringam GR (1999) Cytological stability of doubled haploid lines derived from interspecific crosses between B. napus L. and B. rapa L. In: Proceedings of the 10th International Rapeseed Congr, Australia
Lázaro A, Aguinagalde I (1998) Genetic diversity in Brassica oleracea L. (Cruciferae) and wild relatives (2n = 18) using RAPD markers. Ann Bot 82:829–833
Li H, Chen X, Yang Y, Xu J, Gu J, Fu J, Qian X, Zhang S, Wu J, Liu K (2011) Development and genetic mapping of microsatellite markers from whole genome shotgun sequences in Brassica oleracea. Mol Breed 28:585–596
Li Q, Mei J, Zhang Y, Li J, Ge X, Li Z, Qian W (2013) A large-scale introgression of genomic components of Brassica rapa into B. napus by the bridge of hexaploid derived from hybridization between B. napus and B. oleracea. Theor Appl Genet 126:2073–2080
Li Q, Zhou Q, Mei J, Zhang Y, Li J, Li Z, Ge X, Xiong Z, Huang Y, Qian W (2014) Improvement of Brassica napus via interspecific hybridization between B. napus and B. oleracea. Mol Breed 34:1955–1963
Liu H (1985) Rapeseed genetics and breeding. Shanghai Science and Technology Press, Shanghai, pp 556–559
Mailer RJ, Pratley JE (1990) Field studies of moisture availability effects on glucosinolate and oil concentration in the seed of rape, (Brassica napus L.) and turnip rape (Brassica rapa L. var. silvestris (Lam.) Briggs). Can J Plant Sci 70:399–407
Mason AS, Huteau V, Eber F, Coriton O, Yan G, Nelson MN, Cowling WA, Chevre AM (2010) Genome structure affects the rate of autosyndesis and allosyndesis in AABC, BBAC and CCAB Brassica interspecific hybrids. Chromosome Res 18:655–666
Mei J, Fu Y, Qian L, Xu X, Li J, Qian W (2011) Effectively widening the gene pool of oilseed rape (Brassica napus L.) by using Chinese B. rapa in a ‘virtual allopolyploid’ approach. Plant Breed 130:333–337
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci 76:5269–5273
Nicolas SD, Monod H, Eber F, Chèvre AM, Jenczewski E (2012) Non-random distribution of extensive chromosome rearrangements in Brassica napus depends on genome organization. Plant J 70:691–703
Pelé A, Trotoux G, Eber F, Lodé M, Gilet M, Deniot G, Falentin C, Nègre S, Morice J, Rousseau-Gueutin M, Chèvre A-M (2017) The poor lonesome A subgenome of Brassica napus var. Darmor (AACC) may not survive without its mate. New Phytol 213:1886–1897
Prakash S, Wu XM, Bhat SR (2012) History, evolution, and domestication of Brassica crops. Plant Breed Rev 35:19–84
Qian W, Chen X, Fu D, Zou J, Meng J (2005) Inter sub-genomic heterosis in seed yield potential observed in a new type of Brassica napus introgressed with partial Brassica rapa genome. Theor Appl Genet 110:1187–1194
Qian W, Meng J, Li M, Frauen M, Sass O, Noack J, Jung C (2006) Introgression of genomic components from Chinese Brassica rapa contributes to widening the genetic diversity in rapeseed (B. napus L.), with emphasis on the evolution of Chinese rapeseed. Theor Appl Genet 113:49–54
Qian W, Sass O, Meng J, Li M, Frauen M, Jung C (2007) Heterotic patterns in rapeseed (Brassica napus L.): I. Crosses between spring and Chinese semi-winter lines. Theor Appl Genet 115:27–34
Qian W, Li Q, Noack J, Sass O, Meng J, Frauen M, Jung C (2009) Heterotic patterns in rapeseed (Brassica napus L.): II. Crosses between European winter and Chinese semi-winter lines. Plant Breed 128:466–470
Rahman MH (2001) Production of yellow-seeded Brassica napus through interspecific crosses. Plant Breed 120:463–472
Rahman H (2013) Review: breeding spring canola (Brassica napus L.) by the use of exotic germplasm. Can J Plant Sci 10:24–29
Rahman MH, Bennett RA, Yang RC, Kebede B, Thiagarajah MR (2011) Exploitation of the late flowering species Brassica oleracea L. for the improvement of earliness in B. napus L.: an untraditional approach. Euphytica 177:365–374
Rahman H, Peng G, Yu F, Falk KC, Kulkarni M, Selvaraj G (2014a) Genetics and breeding for clubroot resistance in Canadian spring canola (Brassica napus L.). Can J Plant Path 36(S1):122–134
Rahman H, Kebede B, Zimmerli C, Yang RC (2014b) Genetic study and QTL mapping of seed glucosinolate content Brassica rapa L. Crop Sci 54:537–543
Rahman H, Bennett RA, Séguin-Swartz G (2015) Broadening genetic diversity in Brassica napus canola: development of canola-quality spring B. napus from B. napus × B. oleracea var. alboglabra interspecific crosses. Can J Plant Sci 95:29–41
Rohlf FJ (2000) NTSYS-pc numerical taxonomy and multivariate analysis system. Exeter Software, New York
Rücker B, Röbbelen G (1994) Inheritance of total and individual glucosinolate contents in seeds of winter oilseed rape (Brassica napus L.). Plant Breed 113:206–216
SAS Institute (2011) SAS statistical analysis software version 9.3. SAS Inst, Cary
Simonsen V, Heneen WK (1995) Genetic variation within and among different cultivars and landraces of Brassica campestris L. and B. oleracea L. based on isozymes. Theor Appl Genet 91:346–352
Song KM, Osborn TC, Williams PH (1988) Brassica taxonomy based on nuclear restriction fragment length polymorphisms (RFLPs). 1. Genome evolution of diploid and amphidiploid species. Theor Appl Genet 75:784–794
Thormann CE, Ferreira ME, Camargo LEA, Tivang JG, Osborn TC (1994) Comparison of RFLP and RAPD markers to estimating genetic relationships within and among cruciferous species. Theor Appl Genet 88:973–980
Toroser D, Thormann CE, Osborn TC, Mithen R (1995) RFLP mapping of quantitative trait loci controlling seed aliphatic glucosinolate content in oilseed rape (Brassica napus L.). Theor Appl Genet 91:802–808
Udall JA, Quijada PA, Polewicz H, Vogelzang R, Osborn TC (2004) Phenotypic effects of introgressing Chinese winter and resynthesized Brassica napus L. germplasm into hybrid spring canola. Crop Sci 44:1990–1996
Xiong Z, Pires CJ (2011) Karyotype and identification of all homoeologous chromosomes of allopolyploid Brassica napus and its diploid progenitors. Genetics 187:37–49
Zaman MW (1988) Limitations for introgression of yellow seed coat colour in Brassica napus. J Swedish Seed Assoc 98:157–161
Zhou J, Tan C, Cui C, Ge X, Li Z (2016) Distinct subgenome stabilities in synthesized Brassica allohexaploids. Theor Appl Genet 129(7):1257–1271
Acknowledgements
H.R. gratefully acknowledges the Natural Sciences and Engineering Research Council of Canada (NSERC) (Grand No. CRDPJ 419391-11) and Crop Production Services (CPS) (Grand No. CRDPJ 419391-11) for financial support to this project. Authors also acknowledge technical staff of the Canola Program of the University of Alberta for assistance in different routine operations, and Dr. Berisso Kebede for guidance in molecular marker analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Iftikhar, R., Wang, X. & Rahman, H. Broadening the genetic base of Brassica napus canola by interspecific crosses with different variants of B. oleracea. Euphytica 214, 133 (2018). https://doi.org/10.1007/s10681-018-2213-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2213-4