Abstract
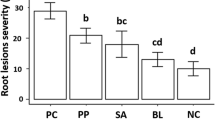
The native plant microbiome is composed of diverse microbial communities that influence overall plant health, with some species known to promote plant growth and pathogen resistance. Here, we show the antibacterial and growth promoting activities of autoclaved culture metabolites (ACM) from native endophytic bacteria (NEB). These NEB were isolated from a papaya cultivar (var. Cariflora) that is tolerant to bacterial crown rot (BCR) caused by Erwinia mallotivora. In this cultivar, bacterial colonization in tissues recovering from the disease was observed before onset of tissue regeneration or ‘regrowth’. We further isolated and characterized these bacteria and were able to identify two culturable stem NEB related to plant endophytic genera Kosakonia sp. (ex. Enterobacter sp., isolate EBW), and to Sphingomonas sp. (isolate EBY). We also identified root NEB under genus Bacillus (isolates BN, BS, and BT). Inhibition assays indicated that ACM from these NEB promptly (within 18-30 h) and efficiently inhibited (60–65% reduction) E. mallotivora proliferation in vitro. When surface-sterilized papaya seeds were soaked in ACM from isolates EBY and EBW, germination was variably retarded (20–60% reduction) depending on plant genotype, but plant biomass accumulation was significantly stimulated, at around two-fold increase. Moreover, greenhouse experiments show that ACM from all isolates, especially isolate EBW, significantly reduced BCR incidence and severity in a susceptible genotype (var. Solo), at around two-fold. In general, our observations of pathogen antagonism and plant growth promotion leading to disease reduction, suggested the influence of native endophytic bacteria to increased fitness in plants, and tolerance against the re-emerging crown rot disease of papaya.




Similar content being viewed by others
References
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., & Lipman, D. J. (1990). Basic local alignment search tool. Journal of Molecular Biology, 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2.
Annapurna, K., Govindasamy, V., Sharma, M., Ghosh, A., and Chikara, S. K. (2018). Whole genome shotgun sequence of Bacillus paralicheniformis strain KMS 80, a rhizobacterial endophyte isolated from rice (Oryza sativa L.). 3 biotech 8, 223. doi:https://doi.org/10.1007/s13205-018-1242-y.
Azri, M. H., Ismail, S., & Abdullah, R. (2018). An endophytic Bacillus strain promotes growth of oil palm seedling by fine root biofilm formation. Rhizosphere, 5, 1–7. https://doi.org/10.1016/j.rhisph.2017.10.003.
Borkar, S. G. (2017). Laboratory techniques in plant bacteriology. Boca Raton : Taylor & Francis, 2017.: CRC press doi:https://doi.org/10.1201/9781315206882.
Brady, C., Cleenwerck, I., Venter, S., Coutinho, T., & De Vos, P. (2013). Taxonomic evaluation of the genus Enterobacter based on multilocus sequence analysis (MLSA): Proposal to reclassify E. nimipressuralis and E. amnigenus into Lelliottia gen. Nov. as Lelliottia nimipressuralis comb. nov. and Lelliottia amnigena comb. nov. Systematic and Applied Microbiology, 36, 309–319. https://doi.org/10.1016/j.syapm.2013.03.005.
Chen, W., & Kuo, T. (1993). A simple and rapid method for the preparation of gram-negative bacterial genomic DNA. Nucleic Acids Research, 21, 2260–2260. https://doi.org/10.1093/nar/21.9.2260.
Cipollini, D., Purrington, C. B., & Bergelson, J. (2003). Costs of induced responses in plants. Basic and Applied Ecology, 4, 79–89. https://doi.org/10.1078/1439-1791-00134.
Coombs, J. T., & Franco, C. M. M. (2003). Isolation and identification of Actinobacteria from surface-sterilized wheat roots. Applied and Environmental Microbiology, 69, 5603–5608. https://doi.org/10.1128/AEM.69.9.5603-5608.2003.
dela Cueva, F., Waje, A., Magdalita, P., Justo, V., Pathania, N., & Vawdrey, L. (2017). Evaluation of inoculation techniques to screen for bacterial crown rot resistance in different breeding lines of Carica papaya. J. Plant Pathol., 99. https://doi.org/10.4454/jpp.v99i2.3887.
Dunlap, C. A., Kwon, S.-W., Rooney, A. P., & Kim, S.-J. (2015). Bacillus paralicheniformis sp. nov., isolated from fermented soybean paste. International Journal of Systematic and Evolutionary Microbiology, 65, 3487–3492. https://doi.org/10.1099/ijsem.0.000441.
Evans, E., and Ballen, F. (2012). Overview of global papaya production, trade, and consumption. UF/IFAS Extension. Article. Available at: http://edis.ifas.ufl.edu/fe913.
Fan, B., Blom, J., Klenk, H.-P., and Borriss, R. (2017). Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis form an “operational group B. amyloliquefaciens” within the B. subtilis species complex. Front. Microbiol. 8. doi:https://doi.org/10.3389/fmicb.2017.00022
Fullerton, R. A., Taufa, L., Vanneste, J. L., Yu, J., Cornish, D. A., & Park, D. (2011). First record of bacterial crown rot of papaya ( Carica papaya ) caused by an Erwinia papayae -like bacterium in the Kingdom of Tonga. Plant Disease, 95, 70–70. https://doi.org/10.1094/PDIS-06-10-0455.
Gardan, L., Christen, R., Achouak, W., & Prior, P. (2004). Erwinia papayae sp. nov., a pathogen of papaya (Carica papaya). International Journal of Systematic and Evolutionary Microbiology, 54, 107–113. https://doi.org/10.1099/ijs.0.02718-0.
Goszczynska, T., Serfontein, J. J., and Serfontein, S. (2000). Introduction to practical phytobacteriology : A manual for phytobacteriology. Pretoria : SAFRINET Available at: http://lib.ugent.be/catalog/rug01:000908450.
Hallmann, J., Quadt-Hallmann, A., Mahaffee, W. F., & Kloepper, J. W. (1997). Bacterial endophytes in agricultural crops. Canadian Journal of Microbiology, 43, 895–914. https://doi.org/10.1139/m97-131.
Hanagasaki, T., Yamashiro, M., Gima, K., Takushi, T., & Kawano, S. (2021). Characterization of Erwinia sp. causing black rot of papaya ( Carica papaya ) first recorded in Okinawa Main Island, Japan. Plant Pathology, 70, 932–942. https://doi.org/10.1111/ppa.13334.
Hardoim, P., Nazir, R., Sessitsch, A., Elhottová, D., Korenblum, E., van Overbeek, L., & van Elsas, J. (2013). The new species Enterobacter oryziphilus sp. nov. and Enterobacter oryzendophyticus sp. nov. are key inhabitants of the endosphere of rice. BMC Microbiol, 13, 164. https://doi.org/10.1186/1471-2180-13-164.
Huang, H.-Y., Li, J., Zhao, G.-Z., Zhu, W.-Y., Yang, L.-L., Tang, H.-Y., Xu, L. H., & Li, W. J. (2012). Sphingomonas endophytica sp. nov., isolated from Artemisia annua L. International Journal of Systematic and Evolutionary Microbiology, 62, 1576–1580. https://doi.org/10.1099/ijs.0.031484-0.
Kämpfer, P., McInroy, J. A., Doijad, S., Chakraborty, T., & Glaeser, S. P. (2016). Kosakonia pseudosacchari sp. nov., an endophyte of Zea mays. Systematic and Applied Microbiology, 39, 1–7. https://doi.org/10.1016/j.syapm.2015.09.004.
Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33, 1870–1874. https://doi.org/10.1093/molbev/msw054.
Liu, H., Brettell, L. E., Qiu, Z., & Singh, B. K. (2020). Microbiome-mediated stress resistance in plants. Trends in Plant Science, 25, 733–743. https://doi.org/10.1016/j.tplants.2020.03.014.
Magdalita, M., dela Cueva, F., Waje, A., Justo, V., & Vawdrey, L. (2016). Regrowth and genotype differences in papaya infected with bacterial crown rot. J. Trop. Plant Pathol., 52, 1–22.
Magdalita, P., dela Cueva, F., Justo, V., & Vawdrey, L. (2015). Performance of bacterial crown rot- tolerant regrowth selections and hybridization with PRSV-tolerant lines in papaya. J. Trop. Plant Pathol., 51, 47–63.
Maktar, N. H., Kamis, S., Mohd Yusof, F. Z., & Hussain, N. H. (2008). Erwinia papayae causing papaya dieback in Malaysia. Plant Pathology, 57, 774–774. https://doi.org/10.1111/j.1365-3059.2008.01877.x.
Maughan, H., & Van der Auwera, G. (2011). Bacillus taxonomy in the genomic era finds phenotypes to be essential though often misleading. Infection, Genetics and Evolution, 11, 789–797. https://doi.org/10.1016/j.meegid.2011.02.001.
Mohd Taha, M. D., Mohd Jaini, M. F., Saidi, N. B., Abdul Rahim, R., Md Shah, U. K., & Mohd Hashim, A. (2019). Biological control of Erwinia mallotivora, the causal agent of papaya dieback disease by indigenous seed-borne endophytic lactic acid bacteria consortium. PLoS One, 14, e0224431. https://doi.org/10.1371/journal.pone.0224431.
Obrero, F. (1980). Bacterial crown rot of papaya in the Philippines: Etiology, epidemiology and screening for disease resistance (PhD Thesis). Available at: University of the Philippines, Los Banos, Laguna, Philippines.
Ortiz-Castro, R., & López-Bucio, J. (2019). Review: Phytostimulation and root architectural responses to quorum-sensing signals and related molecules from rhizobacteria. Plant Science, 284, 135–142. https://doi.org/10.1016/j.plantsci.2019.04.010.
Ploetz, R. (2004). Tropical fruit crops and diseases that affect their production. In: Tropical biology and conservation management – Vol. III. UNESCO-Encyclopedia of Life Support Systems (EOLSS).
Podolich, O., Ardanov, P., Zaets, I., Pirttilä, A. M., & Kozyrovska, N. (2015). Reviving of the endophytic bacterial community as a putative mechanism of plant resistance. Plant and Soil, 388, 367–377. https://doi.org/10.1007/s11104-014-2235-1.
Pordesimo, A., & Quimio, A. (1965). Bacterial soft rot of papaya in the Philippines. J. Trop. Plant Pathol., 1, 19–20.
Rivas, R., Abril, A., Trujillo, M. E., & Velázquez, E. (2004). Sphingomonas phyllosphaerae sp. nov., from the phyllosphere of Acacia caven in Argentina. International Journal of Systematic and Evolutionary Microbiology, 54, 2147–2150. https://doi.org/10.1099/ijs.0.63102-0.
Ruiz-García, C., Béjar, V., Martínez-Checa, F., Llamas, I., & Quesada, E. (2005). Bacillus velezensis sp. nov., a surfactant-producing bacterium isolated from the river Vélez in Málaga, southern Spain. International Journal of Systematic and Evolutionary Microbiology, 55, 191–195. https://doi.org/10.1099/ijs.0.63310-0.
Saitou, N., & Nei, M. (1987). The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution. https://doi.org/10.1093/oxfordjournals.molbev.a040454.
Singh, B. K., Trivedi, P., Singh, S., Macdonald, C. A., & Verma, J. P. (2018). Emerging microbiome technologies for sustainable increase in farm productivity and environmental security. Microbiol. Aust., 39, 17–23. https://doi.org/10.1071/MA18006.
Song, G. C., Choi, H. K., Kim, Y. S., Choi, J. S., & Ryu, C.-M. (2017). Seed defense biopriming with bacterial cyclodipeptides triggers immunity in cucumber and pepper. Scientific Reports, 7, 14209. https://doi.org/10.1038/s41598-017-14155-9.
Supian, S., Saidi, N. B., Wee, C. Y., & Abdullah, M. P. (2017). Antioxidant-mediated response of a susceptible papaya cultivar to a compatible strain of Erwinia mallotivora. Physiological and Molecular Plant Pathology, 98, 37–45. https://doi.org/10.1016/j.pmpp.2017.02.005.
Tamura, K., & Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution. https://doi.org/10.1093/oxfordjournals.molbev.a040023.
Thomas, P., & Kumari, S. (2010). Inconspicuous endophytic bacteria mimicking latex exudates in shoot-tip cultures of papaya. Sci. Hortic. (Amsterdam)., 124, 469–474. https://doi.org/10.1016/j.scienta.2010.02.013.
Thomas, P., Kumari, S., Swarna, G. K., & Gowda, T. K. S. (2007a). Papaya shoot tip associated endophytic bacteria isolated from in vitro cultures and host–endophyte interaction in vitro and in vivo. Canadian Journal of Microbiology, 53, 380–390. https://doi.org/10.1139/W06-141.
Thomas, P., Kumari, S., Swarna, G. K., Prakash, D. P., & Dinesh, M. R. (2007b). Ubiquitous presence of fastidious endophytic bacteria in field shoots and index-negative apparently clean shoot-tip cultures of papaya. Plant Cell Reports, 26, 1491–1499. https://doi.org/10.1007/s00299-007-0363-2.
Thompson, J. D., Higgins, D. G., & Gibson, T. J. (1994). CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Research, 22, 4673–4680.
Trivedi, P., Leach, J. E., Tringe, S. G., Sa, T., & Singh, B. K. (2020). Plant–microbiome interactions: From community assembly to plant health. Nature Reviews. Microbiology, 18, 607–621. https://doi.org/10.1038/s41579-020-0412-1.
Trujillo, E. E., & Schroth, M. N. (1982). Two bacterial diseases of papaya trees caused by Erwinia species in the Northern Mariana Islands. Plant Disease, 66, 116. https://doi.org/10.1094/PD-66-116.
Ventura, J., Costa, H., and Fabra, A. (2004). Papaya diseases and integrated control. In: NAQVI SAMH (ed.). Diseases of fruits and vegetables. Kluwer Academic Publishers. .
von Rant, A. (1931). U¨ ber eine Bakterienkrankheit bei dem Melonenbaume (Carica papaya L.) auf Java (in German). Zentralblatt für Bakteriol. Parasitenkd. Infekt. und Hyg. 84, 481–487.
Wang, Y., Liu, H., Liu, K., Wang, C., Ma, H., Li, Y., Hou, Q., Liu, F., Zhang, T., Wang, H., Wang, B., Ma, J., Ge, R., Xu, B., Yao, G., Xu, W., Fan, L., Ding, Y., & du, B. (2017). Complete genome sequence of Bacillus paralicheniformis MDJK30, a plant growth-promoting Rhizobacterium with antifungal activity. Genome Announcements, 5. https://doi.org/10.1128/genomeA.00577-17.
Webb, R. (1984). Epidemiology and control of bacterial canker of papaya caused by an Erwinia sp. on St. Croix, U.S. Virgin Islands. Plant Dis., 69, 305. https://doi.org/10.1094/PD-69-305.
Weisburg, W. G., Barns, S. M., Pelletier, D. A., & Lane, D. J. (1991). 16S ribosomal DNA amplification for phylogenetic study. Journal of Bacteriology, 173, 697–703. https://doi.org/10.1128/jb.173.2.697-703.1991.
Whitman, W., DeVos, P., Dedysh, S., Hedlund, B., Kämpfer, P., Rainey, F., et al. (2015). Bergey’s manual of systematics of archaea and Bacteria. Wiley. https://doi.org/10.1002/9781118960608.
Wicaksono, W. A., Jones, E. E., Casonato, S., Monk, J., & Ridgway, H. J. (2018). Biological control of Pseudomonas syringae pv. Actinidiae (Psa), the causal agent of bacterial canker of kiwifruit, using endophytic bacteria recovered from a medicinal plant. Biol Control, 116, 103–112. https://doi.org/10.1016/j.biocontrol.2017.03.003.
Worrall, D., Holroyd, G. H., Moore, J. P., Glowacz, M., Croft, P., Taylor, J. E., Paul, N. D., & Roberts, M. R. (2012). Treating seeds with activators of plant defence generates long-lasting priming of resistance to pests and pathogens. The New Phytologist, 193, 770–778. https://doi.org/10.1111/j.1469-8137.2011.03987.x.
Yang, S., Zhang, X., Cao, Z., Zhao, K., Wang, S., Chen, M., & Hu, X. (2014). Growth-promoting S phingomonas paucimobilis ZJSH1 associated with D endrobium officinale through phytohormone production and nitrogen fixation. Microbial Biotechnology, 7, 611–620. https://doi.org/10.1111/1751-7915.12148.
Yoon, S.-H., Ha, S.-M., Kwon, S., Lim, J., Kim, Y., Seo, H., & Chun, J. (2017). Introducing EzBioCloud: A taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. International Journal of Systematic and Evolutionary Microbiology, 67, 1613–1617. https://doi.org/10.1099/ijsem.0.001755.
Zhu, B., Zhou, Q., Lin, L., Hu, C., Shen, P., Yang, L., An, Q., Xie, G., & Li, Y. (2013). Enterobacter sacchari sp. nov., a nitrogen-fixing bacterium associated with sugar cane (Saccharum officinarum L.). International Journal of Systematic and Evolutionary Microbiology, 63, 2577–2582. https://doi.org/10.1099/ijs.0.045500-0.
Acknowledgements
The authors would like to thank Valeriana Justo, Lorele Trinidad, Ireneo Pangga, Teresita Dalisay, Fe dela Cueva, Demetrio Alvarez, Ramon Cortez, Gil Cueto, Besseluz Abayon and Marcelino Gregorio of the University of the Philippines Los Baños in Laguna, Philippines, and Eka Mirnia of the Indonesian Agency of Agricultural Research and Development in West Sumatra, Indonesia, for their technical support during the conduct of the experiments. The authors would also like to thank Karla Manigbas, and the staff of Diamed Enterprises in Laguna, Philippines for their DNA sequencing logistics services.
Availability of data and Supplementary materials
Sequences of the 16S rRNA gene for each bacterial isolate are deposited in NCBI GenBank, with accession numbers MW405488, MW405489, MW405490, MW405491 and MW405492. All supplementary materials from this study can be downloaded through this link: https://doi.org/10.6084/m9.figshare.13499409.v1
Funding
The authors are very grateful for the support received from two funding institutions: the University of the Philippines Los Baños (UPLB) Basic Research Program (Project number 9116004), and the Australian Centre for International Agricultural Research (ACIAR, Project code HORT/2012/113).
Author information
Authors and Affiliations
Contributions
MPSR and PMM conceptualized the study, and acquired funding. PMM supervised all experiments. Materials preparation, experimental setup and data collection were performed by MPSR, with substantial assistance from EPP and SFMD. MPSR did the analysis of raw data, and wrote the first draft of the manuscript, with all the tables and figures. All authors contributed revisions and proofreading, and approved the final manuscript.
Corresponding author
Ethics declarations
Informed consent
All authors have approved the manuscript and agreed on its submission to the European Journal of Plant Pathology. The authors confirm that no other person involved in this research have equivalent contributions to what is listed above, that may merit authorship. Support from other involved individuals are acknowledged accordingly.
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This research article is not submitted elsewhere for publication and complies with the Ethical Rules applicable for this journal.
Human and animal rights
This article does not contain any studies with human participants or vertebrate animals.
Rights and permissions
About this article
Cite this article
Rivarez, M.P.S., Parac, E.P., Dimasingkil, S.F.M. et al. Influence of native endophytic bacteria on the growth and bacterial crown rot tolerance of papaya (Carica papaya). Eur J Plant Pathol 161, 593–606 (2021). https://doi.org/10.1007/s10658-021-02345-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10658-021-02345-1