Abstract
Background
Gastric cancer (GC) seriously threatens people’s health and life quality worldwide.
Aim
The current study sought to explore prognostic immune genes and their regulatory network in GC.
Methods
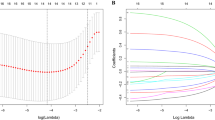
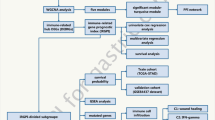
First, expression data in GC and normal samples were analyzed based on bioinformatics analysis. Immune-related genes were identified and confirmed with univariate/multivariate Cox analysis and receiver-operating characteristic curve. The upstream transcription factors of immune genes were subsequently predicted, and their regulatory network was constructed. GC and adjacent normal tissues were obtained from 76 patients with GC to determine the expression patterns of immune genes and their correlation with overall prognosis. CD8+ T-cell infiltration of patients with high or low risk was detected by means of immunohistochemistry.
Results
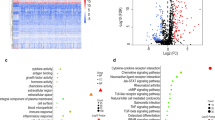
Bioinformatics analysis highlighted 3689 differentially expressed genes in GC, including 87 immune genes, 8 of which were significantly associated with patient survival. CGB5 and INHBA were high-risk genes, while TRAJ19 was identified as a low-risk gene, all of which were found to be regulated by 11 different transcription factors. Furthermore, CGB5 and INHBA exhibited negative correlation with the prognosis of GC patients; however, TRAJ19 was positively correlated with GC patient prognosis. The incidence of lymph node metastasis was higher, the pathological stage was advanced and the infiltrated CD8+ T cells were fewer in the high-risk GC group.
Conclusions
Overall, our findings identified the key roles of CGB5, INHBA and TRAJ19 in prognosis GC patients, serving as an important gene set for prognostic prediction.










Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- GC:
-
Gastric cancer
- GTEx:
-
Genotype-Tissue Expressionx
- DEGs:
-
Differentially expressed genes
- FACS:
-
Fluorescence activated cell sorting
- RT-qPCR:
-
Reverse transcription quantitative polymerase chain reaction
References
Eusebi LH, Telese A, Marasco G et al. Gastric cancer prevention strategies: a global perspective. J Gastroenterol Hepatol. 2020;35:1495–1502.
Machlowska J, Baj J, Sitarz M et al. Gastric cancer: epidemiology, risk factors, classification, genomic characteristics and treatment strategies. Int J Mol Sci 2020;21:4012.
Zhang Y, Yu J. The role of MRI in the diagnosis and treatment of gastric cancer. Diagn Interv Radiol 2020;26:176–182.
Sitarz R, Skierucha M, Mielko J et al. Gastric cancer: epidemiology, prevention, classification, and treatment. Cancer Manag Res. 2018;10:239–248.
Refolo MG, Lotesoriere C, Messa C et al. Integrated immune gene expression signature and molecular classification in gastric cancer: new insights. J Leukoc Biol. 2020;108:633–646.
Foukakis T, Lovrot J, Matikas A et al. Immune gene expression and response to chemotherapy in advanced breast cancer. Br J Cancer. 2018;118:480–488.
Tsakonas G, Lewensohn R, Botling J et al. An immune gene expression signature distinguishes central nervous system metastases from primary tumours in non-small-cell lung cancer. Eur J Cancer. 2020;132:24–34.
Huang R, Chen Z, Li W et al. Immune systemassociated genes increase malignant progression and can be used to predict clinical outcome in patients with hepatocellular carcinoma. Int J Oncol. 2020;56:1199–1211.
Li L, Wang XL, Lei Q et al. Comprehensive immunogenomic landscape analysis of prognosis-related genes in head and neck cancer. Sci Rep. 2020;10:6395.
Nakamura K, Hatakeyama K, Furukawa K et al. Prediction of S-1 adjuvant chemotherapy benefit in Stage II/III gastric cancer treatment based on comprehensive gene expression analysis. Gastric Cancer. 2020;23:648–658.
Newman AM, Liu CL, Green MR et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods. 2015;12:453–457.
Ali HR, Chlon L, Pharoah PD et al. Patterns of immune infiltration in breast cancer and their clinical implications: a gene-expression-based retrospective study. PLoS Med. 2016;13:e1002194.
Schleif RF. Modulation of DNA binding by gene-specific transcription factors. Biochemistry. 2013;52:6755–6765.
Mierke CT. The matrix environmental and cell mechanical properties regulate cell migration and contribute to the invasive phenotype of cancer cells. Rep Prog Phys. 2019;82:064602.
Kang BW, Kim JG, Lee IH et al. Clinical significance of tumor-infiltrating lymphocytes for gastric cancer in the era of immunology. World J Gastrointest Oncol. 2017;9:293–299.
Farhood B, Najafi M, Mortezaee K. CD8(+) cytotoxic T lymphocytes in cancer immunotherapy: a review. J Cell Physiol. 2019;234:8509–8521.
Nie K, Shi L, Wen Y et al. Identification of hub genes correlated with the pathogenesis and prognosis of gastric cancer via bioinformatics methods. Minerva Med. 2019;111:213–225.
Yang Y, Shi Y, Hou Y et al. CGB5 expression is independently associated with poor overall survival and recurrence-free survival in patients with advanced gastric cancer. Cancer Med. 2018;7:716–725.
Chen Y, Kibriya MG, Jasmine F et al. Do placental genes affect maternal breast cancer? Association between offspring’s CGB5 and CSH1 gene variants and maternal breast cancer risk. Cancer Res. 2008;68:9729–9734.
Li X, Yu W, Liang C et al. INHBA is a prognostic predictor for patients with colon adenocarcinoma. BMC Cancer. 2020;20:305.
Liu X, Liu X, Qiao T et al. Identification of crucial genes and pathways associated with colorectal cancer by bioinformatics analysis. Oncol Lett. 2020;19:1881–1889.
Wang W, He Y, Zhao Q et al. Identification of potential key genes in gastric cancer using bioinformatics analysis. Biomed Rep. 2020;12:178–192.
Biskup K, Blanchard V, Castillo-Binder P et al. N- and O-glycosylation patterns and functional testing of CGB7 versus CGB3/5/8 variants of the human chorionic gonadotropin (hCG) beta subunit. Glycoconj J. 2020;37:599–610.
Sun YL, Zhang Y, Guo YC et al. A prognostic model based on the immune-related genes in colon adenocarcinoma. Int J Med Sci. 2020;17:1879–1896.
Wan H, Liu Z, Tan X et al. Application of immune cell infiltration in the diagnosis and prognosis of non-small cell lung cancer. Sheng Wu Gong Cheng Xue Bao 2020;36:740–749.
Wu M, Ding J, Song J et al. Genomic analysis and clinical implications of immune cell infiltration in gastric cancer. Biosci Rep. 2020;40:BSR20193308.
De Rossi A, Huaman SD, Leon J et al. Fibroblast growth factor receptor 2 expression in apical periodontitis in mice. Int Endod J. 2020;53:1111–1119.
Ramakrishna B, Yewale R, Vijayakumar K et al. Gastric IgG4-related disease presenting as a mass lesion and masquerading as a gastrointestinal stromal tumor. J Pathol Transl Med. 2020;54:258.
Lou JS, Wang JF, Fei MM et al. Targeting LAG-3 reverses T lymphocytes dysfunction and improves survival in murine polymicrobial sepsis. J Infect Dis. 2020;222:1051–1061.
Yuan X, Tang Y, Zhao Z et al. Identification and verification of EOMEs regulated network in Alopecia areata. Int Immunopharmacol. 2020;84:106544.
Taveirne S, Wahlen S, Van Loocke W et al. The transcription factor ETS1 is an important regulator of human NK cell development and terminal differentiation. Blood 2020;136:288–298.
Acknowledgments
We acknowledge and appreciate our colleagues for their valuable contributions to this work.
Funding
Project ZR2019PH102 supported by Shandong Provincial Natural Science Foundation.
Author information
Authors and Affiliations
Contributions
BJ and WZ designed the project and prepared the main manuscript. LLQ prepared the manuscript and figures. All authors agreed to the publication. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

Rights and permissions
About this article
Cite this article
Ji, B., Qiao, L. & Zhai, W. CGB5, INHBA and TRAJ19 Hold Prognostic Potential as Immune Genes for Patients with Gastric Cancer. Dig Dis Sci 68, 791–802 (2023). https://doi.org/10.1007/s10620-022-07513-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-022-07513-9