Abstract
Genetic monitoring has become a popular instrument in the conservation of endangered species, allowing to estimate size and genetic structure of wild populations. Long-term monitoring projects are essential to recognize demographic changes and impact of human activities. Since 2011, an extensive monitoring project on the population size and trends, as well as spatial distribution and survival rates, of two grouse species including the western capercaillie, Tetrao urogallus, has been conducted in Tyrol, in the eastern part of the European Alps, where T. urogallus males are huntable under specific regulations. In this case study, we aimed to compile a set of analyses to be employed in evaluating data from dropping and feather samples for conservation studies. Using eleven microsatellite and two sex markers, we genotyped 251 faeces and feathers of T. urogallus collected in East Tyrol in spring 2019. We analysed population structure and mobility patterns, including sex differences in genetic diversity and mobility. The relationship between habitat parameters and genetic diversity was investigated using multiple linear regressions. We showed that the investigated T. urogallus population is well mixed and likely well connected to neighbouring populations. We also found sex-specific mobility patterns that support female-biased dispersal. As the last step, we demonstrated the general feasibility of a modelling approach using habitat parameters. With this pilot study, further analysis of data is possible for the whole monitoring project, giving a better insight in the grouse populations in Tyrol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With their large spatial demands and specific habitat requirements, umbrella species are common targets of conservation measures that should benefit multiple species in the same community (Simberloff 1998; Roberge and Angelstam 2004). The western capercaillie (Tetrao urogallus) is an effective umbrella species for red-listed mountain birds (Suter et al. 2002) and a good candidate for specialist birds in taiga forests (Pakkala et al. 2003). Typically, T. urogallus habitat consists of semi-open, well-structured coniferous forest with trees of varying ages, and particularly old stands with decaying wood, as well as well-developed dwarf-shrub vegetation (Storch 2007; Lentner et al. 2022). The species is closely associated with the structure of the forest, needing open gaps for mating displays and rich ground cover with Vaccinium berries for feeding (Storch 2007). Adults feed mostly on leaves, buds, flowers, and fruits in summer and conifer needles in the winter, while chicks rely on invertebrates (Storch 2007). In Central Europe, the distribution range of T. urogallus is mainly limited to mountain forests (Storch 2007). Because of habitat loss and fragmentation, many of the populations in Central Europe are small and isolated (Storch 2007; Segelbacher et al. 2008; Keller et al. 2020), so that priorities for conservation are preserving habitat, maintaining connectivity, and reducing human disturbance (Storch 2007). The conservation status for Europe was evaluated as of Least Concern with an decreasing population trend (IUCN 2016), while the species is listed as threatened in Western, Central, and South Eastern European countries (Storch 2007; Dvorak et al. 2017) and is included in Annex I of the European Birds Directive (79/409/CEE).
Given the conservation status of T. urogallus and its role as umbrella species (Suter et al. 2002), monitoring is a valuable instrument to assess population trends and responses to management actions. Traditionally, counts of males at leks have been used to estimate population size (Lentner et al. 2018; Abrahams 2019). Starting in late winter, males display collectively in lek sites to attract the females (Storch 2007). This method might underestimate population size (Jacob et al. 2010; Aleix-Mata et al. 2019), since young individuals that are not showing lekking behaviour or using only peripheral territories can be overlooked (Abrahams 2019; Aleix-Mata et al. 2019). Furthermore, certain methodological standards (e.g. simultaneous counts) and the experience of observers (Lentner et al. 2018) may differ among studies, which has impacts on the estimations. Alternatively, genetic analysis of non-invasive samples (e.g. faeces, feathers) has become increasingly popular to study population size and structure of T. urogallus and is commonly used for monitoring (Jacob et al. 2010; Morán-Luis et al. 2014; Rösner et al. 2014; Mollet et al. 2015; Bañuelos et al. 2019).
Since 2011, an extensive grouse-monitoring project has been taking place in Tyrol, in the eastern part of the European Alps (Masoner 2012; Lentner et al. 2018; Vallant et al. 2018), investigating two grouse species (T. urogallus and the black grouse, Lyrurus tetrix). Four reference areas covering 1100 km2 were established to investigate 5–10% of the grouse populations in Tyrol in a 5-year cycle (Lentner et al. 2018). Each year, one of the reference areas is systematically searched for faeces and feathers using a transect mapping approach. Additionally, synchronous counts at known lek sites are performed. This is done in intensive sampling sites that cover yearly a total area of 1112–1418 ha for T. urogallus. Genetic analysis of the collected samples allows to determine individuals and to estimate population sizes through a mark-recapture method (Pennell et al. 2013; Lentner et al. 2018). The currently estimated population size of T. urogallus in Tyrol is 1700–2300 cocks (Lentner et al. 2022). Via a standardised sampling protocol and repetition every 5 years (1 year break after a completed round), this monitoring aims at observing changes of grouse population over a longer time period. Furthermore, a main aim of the project is to calibrate and validate the conventional lek counts of the 5-year regional grouse monitoring of the Tyrolean Hunters’ Association by comparison with the population estimates based on genetic data from selected reference areas.
In this pilot study, we analysed non-invasive T. urogallus samples collected in 2019 in a reference area in East Tyrol as part of the Tyrolean grouse monitoring using multi-locus genotyping. We aimed to compile a set of analyses that could be a starting point for the analysis of the complete monitoring data from 2011 to 2019 for both T. urogallus and Lyrurus tetrix. We analysed population structure using Bayesian cluster analysis and calculating population differentiation. Also, mobility patterns and differences between males and females were investigated comparing pairwise relatedness between individuals, genetic diversity, and inbreeding. Furthermore, multiple linear regressions were used with a set of explanatory variables (habitat parameters) and response variables (genetic parameters) to find which habitat parameters explain variation in genetic diversity.
Materials and methods
Sampling
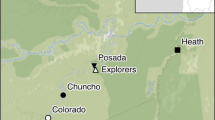
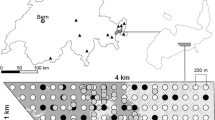
Fieldwork was conducted in East Tyrol (Austria) from end of March to end of April 2019. As part of the grouse monitoring in Tyrol, a reference area of 38,286 ha was defined in Defereggen valley and the front part of Virgen valley. In this area, 11 sampling sites (Fig. 1; mean area: 100 ha, standard deviation: 49 ha) were systematically searched for faeces and feathers of T. urogallus along transects about 100 m in altitude apart from each other (and 100 m of horizontal distance on flat terrain) covering the complete sampling site. These two different transect distances were chosen to be able to map both steep and flat terrain, with steep terrain representing the vast majority of all terrain sampled here. The surveyors followed isohypses but adjusted their paths to include relevant habitat structures (stands of old and dead coniferous trees and lek sites). For each sample, GPS coordinates were noted using Garmin GPSMAP 64sx and comparable devices of the same product series. To reduce researcher and sampling bias, each sampling site was visited a second time after 16–19 days by a different surveyor using a transect route different from that used in the first survey.
Locations of Tetrao urogallus samples in East Tyrol. Circles represent faeces and feathers (n = 251) collected along transects in eleven monitoring sampling sites. Every site was visited twice by different surveyors, and transect routes covered the whole sampling sites. Inset: distribution of T. urogallus from BirdLife International and Handbook of the Birds of the World (2019) Bird species distribution maps of the world. Version 2019.1. Available at http://datazone.birdlife.org/species/requestdis
In total, 651 faeces and feathers were collected. For the molecular-genetic analyses, the number of samples for each sampling site was chosen proportionally to its area among all samples collected from that site, to achieve a total sample size of 251 [n (faeces) = 245; n (feathers) = 6], which corresponds to the sample size of the grouse monitoring. Sampling points were defined as circular areas of 50 m diameter, and for any sampling point, only one sample was analysed unless definitively belonging to different individuals. This was the case for samples that could be assigned to male or female individuals based on their size (faeces) or colour (feathers), as noted on field protocols. Priority was given to fresh samples collected on snow. Samples were equally distributed between sampling rounds.
DNA extraction and genotyping
DNA from faeces samples was extracted using the QIAamp Fast DNA Stool Mini kit (Qiagen, Hilden, Germany) following the protocol of the manufacturer. For feather samples, the QIAamp DNA Mini Kit (Qiagen) was used with the modifications described by Vallant et al. (2018). Eluted DNA was stored at − 20 °C.
Samples were genotyped three times using eleven microsatellite markers (Table 1) and two additional sex markers, Tt_Wa and Tt_Za (Haider et al. 2023), targeting the CHD gene. Markers were organized into three multiplex PCR sets (Table 1).
PCRs were done using the multiple tube approach (Taberlet et al. 1996) on a UnoCycler (VWR, USA) in 10 µl reaction volume with 1 µl template DNA, 1× Qiagen Multiplex PCR Master Mix, 0.6 µl non-acetylated BSA (2.5 µg/µl; Thermo-Fisher), 3 mM MgCl2, 0.16 µM of each primer, and RNase-free water (Qiagen). Cycling conditions were as follows: initial denaturation at 95 °C for 15 min, 37 cycles of denaturation at 94 °C for 30 s, annealing at 56 °C for 120 s, elongation at 72 °C for 30 s, and final extension at 72 °C for 45 min. Negative and positive controls were included as well as allelic ladders (Andesner et al. 2021).
Fragments were sized by capillary electrophoresis at University of Chicago Comprehensive Cancer Center DNA Sequencing & Genotyping Facility (Chicago, USA). Allele calling was done using GeneMarker v2.7 (SoftGenetics, State College, PA, USA). If for a particular marker a sample failed completely, did not reach the minimum signal intensity, and/or did not show clear peaks, the marker was classified as missing for this sample.
Species and genotype determination
Species identification was done based on distinct allele-sizes ranges for loci BG15, BG19, and sTuT2 (Vallant et al. 2018). Unique genotypes were identified using the R package allelematch v2.5.1 (Galpern et al. 2012) allowing for three mismatches. Allowed number of mismatches was determined using the implanted function amUniqueProfile. To check the individualization, probability of identity (Pi) and probability of identity for siblings (Pisib) were calculated with GeneCap v1.4 (Wilberg and Dreher 2004).
Population genetics
Loci were tested for linkage disequilibrium and deviation from Hardy–Weinberg equilibrium (HWE) with probability tests implemented in Genepop (Rousset 2008). Bonferroni–Holm correction was applied for multiple comparisons (Holm 1979). For the following analyses, only one sampling site was randomly selected for genotypes found at several sampling sites. All calculation using R packages were done in R v4.2.3 (R Core Team 2023).
Inbreeding coefficient (FIS) and pairwise fixation index (FST) were calculated using the package hierfstat v0.5-11 (Goudet 2005). Individual inbreeding was estimated as multi-locus heterozygosity (MLH) and mean squared distance between alleles (d2) (Coulson et al. 1998). The packages adegenet v 2.1.10 (Jombart 2008) was used to calculate observed heterozygosity (HO). Allelic richness (AR) was calculated using PopGenReport v3.0.4 (Adamack and Gruber 2014). Calculations were done also for males and females separately. Differences between sexes were tested by Mann–Whitney U test using the package rstatix v0.7.2 (Kassambara 2023).
Isolation by distance (IBD) in males and females was tested using Mantel tests (9999 permutations) in ade4 v1.7-15 (Chessel et al. 2004). Geographic and Edward’s genetic distances (Edwards 1971) between the individuals were calculated using geosphere v1.5-10 (Hijmans et al. 2019) and poppr v2.8.6 (Kamvar et al. 2014), respectively.
Pairwise relatedness between individuals was estimated by the coefficients implemented in the R package related v1.0 (Pew et al. 2015). Relatedness within sites and between sites for males and females separately were compared using Mann-Whitney U test using the R package rstatix v 0.7.2 (Kassambara 2023).
A Bayesian cluster analysis was performed in STRUCTURE v2.3.4 (Pritchard et al. 2000) using an admixture model with correlated allele frequencies, 1,000,000 generations burn-in, and 4,000,000 MCMC generations with 10 replicates. The most likely number of clusters was determined by ΔK (rate of change of probability between the different numbers of populations used) (Evanno et al. 2005) using the R package pophelper v2.0.3 (Francis 2017).
Habitat and genetic diversity
Two habitat suitability models were developed for the grouse monitoring Tyrol (Masoner 2012; Lehne 2014; Lentner 2017). The first is an ecological niche model generated using MaxEnt (Phillips et al. 2004), expressing the potential habitat suitability for T. urogallus as a probability. The second represents habitat suitability as a score resulting from expert assessment of landscape parameters (Exp) (Masoner 2012; Lehne 2014; Lentner 2017). Data were provided by Michael Haupolter, Regional Government of Tyrol, Department of Environmental Protection – tiris (geographical information system of Regional Government of Tyrol). In order to consider the total number of genotypes per site, all detected sampling sites for a genotype were used for the modelling.
Linear regressions were used to determine the effect of habitat parameters on genetic diversity. Explanatory variables were: area, perimeter, shape (perimeter/area), two habitat suitability models (MaxEnt and Exp), isolation measures Hanski (MaxEnt) and Hanski (Exp) (Hanski et al. 1994), and mean distance to other sites.
For the two isolation measures of sampling sites, Hanski (MaxEnt) and Hanski (Exp), Hanski‘s I4 (Hanski et al. 1994) was modified to include habitat suitability (Hanski et al. 1994; Rabasa et al. 2007): \({I4}_{i} = {\sum }_{j\ne i}{e}^{-\alpha *{d}_{ij}}\text{*}{A}_{j}\text{*}{S}_{j}\), where \({d}_{ij}\) is the distance between sites \(i\) and \(j\) (km), \({A}_{j}\) the area (ha) of site \(i\), \(\alpha\) the reciprocal of the average dispersal distance (5 km), and \({S}_{j}\) the habitat suitability of site \(j\). The measure was calculated for both habitat suitability models.
Response variables were d2, MLH, FIS, AR, HO, and the number of genotypes (N.genotypes). Since the number of genotypes depends largely on the number of samples analysed, which is proportional to the area of the sites, number of genotypes divided by area was added to the response variables.
First, simple linear regressions were calculated for every combination explanatory/response variable. Then, multiple linear regressions were used for number of genotypes divided by area. To identify the subset of habitat parameters that best explained these variables, stepwise model selection was performed. The best model was chosen based on Akaike Information Criterion (AIC).
Results
Species and genotype determination
In total, 251 samples were analysed; five were determined as Lyrurus tetrix, two as Tetrastes bonasia, 239 as Tetrao urogallus, and the remaining five could not be genotyped for a sufficient number of species-specific loci to allow determination. Unique genotypes were identified using 236 T. urogallus samples that were genotyped successfully at ≥ 8 loci; we found 55 males and 28 females of T. urogallus (Supplementary Table 1). Each genotype was found 2.9 times on average (males: 3.5, females: 1.6), 36 were found only once, 45 multiple times (males: max = 17, mean = 5.0; females: max = 6, mean = 2.6). The maximum distance between recaptured genotypes was 5.5 km for males and 2.5 km for females. Pid and Psib were 2.93 × 10−10 and 3.69 × 10−4, respectively.
Population genetics
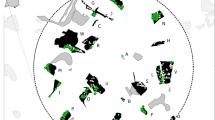
Loci were tested for deviation from HWE and for linkage disequilibrium. Based on significant deviation from HWE, locus mTuT1 was excluded from further analyses. Pairwise FST values between sites were small with only a few high values (FST > 0.05) between sites where only one individual was found (Tu_27, highest value 0.15) and non-significant, without clear geographic patterns of differentiation (Fig. 2).
Multi-locus heterozygosity and FST were significantly different between females and males (Fig. 3d, f; Mann-Whitney U test: W = 613, p = 0.001; W = 1068, p = 0.003). We found no significant difference in AR (W = 32, p = 0.19), d2 (W = 694, p = 0.47), FIS (W = 53, p = 0.85), nor HO (W = 63, p = 0.34) between females and males (Fig. 3).
Comparisons of genetic diversity and inbreeding between females (F) and males (M) of Tetrao urogallus. None of the calculated values were significantly different between sexes except multi-locus heterozygosity (MLH; f). AR allelic richness, a d2 mean squared distance between alleles, b FIS inbreeding coefficient, c FST fixation index, d HO observed heterozygosity, e Dots represent mean values, error bars standard deviation. Significance level of Mann–Whitney U test: **p < 0.01
IBD was statistically significant for males but not for females (Mantel test males: r = 0.068, p = 0.03; females: r = − 0.056, p = 0.88, Fig. 4).
Isolation by distance (IBD) for a males and b females of Tetrao urogallus with regression line. Warmer colours indicate higher density of points. a There is a significant increase of genetic distance over geographic distance in males, whereas b in females the result is not significant. Significance level of Mantel test: ns p > 0.05, *p ≤ 0.05
Pairwise relatedness for males within sampling sites was significantly higher than for males from different sites for three out of the seven relatedness coefficients calculated (Fig. 5, Supplementary Fig. 1). No difference was found for females (Fig. 5, Supplementary Fig. 1).
Comparison of relatedness coefficients using different methods [Lynch–Ritland (Lynch and Ritland 1999) and Queller–Goodnight (Queller and Goodnight 1989)] for males (a, c) and females (b, d) of Tetrao urogallus. a, b There was no differences between inter and intra relatedness using Lynch–Ritland. c, d In males, intra relatedness was significantly higher than inter relatedness. In females, no difference was detected. Inter: between sampling sites, intra: within sampling sites. Dots represent mean values, error bars standard deviation. Significance levels of Mann–Whitney U test: *p < 0.05
ΔK statistics indicated a best value of K = 4 for Bayesian clustering (Supplementary Fig. 2). The population showed complete admixture (Fig. 6). The same was observed for other values of K (Fig. 6).
Habitat and genetic diversity
Simple linear regressions were used to find the most appropriate response variables among genetic parameters. Regressions were significant for N.genotypes (Table 2).
Since this parameter is strongly affected by the area of the sampling site because of sampling design, density of genotypes (N.genotypes/area) was used for the multiple linear regressions. The best model (stepwise selection) contained Hanski isolation measure calculated using the expert model and shape of the sampling site (perimeter/area) (Table 3).
Discussion
In this pilot study, we tested various data-analysis methods for non-invasive samples of Tetrao urogallus from East Tyrol to investigate population structure and mobility using ten polymorphic microsatellite markers. The relationship between habitat parameters and genetic diversity was also analysed using linear regressions. The markers discriminated well among individuals with probability of identity (Pi) and probability of identity for siblings (Pisib) lower than the recommended values (10−2 − 10−4) (Waits et al. 2001). We also genotyped each sample three times to minimize the error rates (Taberlet et al. 1996; Beja-Pereira et al. 2009) and saw only minor differences (mainly a minimized number of failed alleles), indicating that the marker set used is very reliable (Supplementary Table 2).
Locus mTuT1 deviated significantly from Hardy-Weinberg equilibrium (HWE) and was subsequently excluded from population genetic calculations. A deviation from HWE across multiple loci could be explained by population stratification, assortative mating, and/or poor DNA quality causing allelic dropout. Since only one locus was affected, it is more likely that null alleles are responsible, as found in other studies using this marker (Jacob et al. 2010; Vallant et al. 2018).
Population structure
Our results suggest that the individuals analysed belong to a single well-mixed population. Low values of the fixation index FST indicate a high level of gene flow among the sampling sites without clear geographical pattern (Fig. 2). However, there is an exception with high pairwise FST values observed in the Tu_27 and Tu_32 sampling sites, which can be explained by the number of individuals found there. Although even FST values smaller than 0.05 could indicate some differentiation (Balloux and Lugon-Moulin 2002), the STRUCTURE clustering showed complete admixture at all K values analysed (Fig. 6). However, one has to be aware that gene flow as parameter for differentiation cannot provide insight in e.g. the demographic history (Marko and Hart 2011). In our case, the sampling sites are likely well connected, and the area investigated is relatively small, with a maximum distance between sites of 23 km. Furthermore, the sites do not represent distinct habitat fragments. On the one hand, (Rösner et al. 2014) found that movement of single capercaillies of up to 30 km can maintain gene flow across larger distances (up to 85 km) in the Bohemian Forest. On the other hand, significant differentiation has been observed for a fragmented population in the Black Forest (Germany) even at fine spatial scale (Segelbacher et al. 2008). It is possible that the complex topography does not allow free dispersal in our study area with high mountain ranges does not allow free dispersal, in contrast to the Bohemian and the Black Forest (Segelbacher and Storch 2002). Nevertheless, this does not seem to significantly limit gene flow at the scale we studied.
Mobility patterns
The low FST values and the results of the STRUCTURE clustering reflect high connectivity and ongoing gene-flow among sampling sites. This is also supported by the recapture of single individuals at multiple sites, with a maximum air-line distance of 5.5 km between recaptures. In previous years of the monitoring, the longest distance recorded was 10.2 km (Lentner 2017). Since sampling was conducted over three weeks in the mating season, this result could reflect short-term movement rather than dispersal patterns. Storch (2007) found that juveniles and subadult capercaillies visited several leks and had larger home ranges, while adult males occupied smaller, overlapping territories closer to lek centres. Females also visited multiple leks (Storch 1997).
Our results suggest sex-specific differences in movement patterns. While most comparisons of inbreeding and genetic diversity were not statistically significant, MLH and FST were significantly different between the sexes with higher MLH and lower FST in females than males (Fig. 3). This could be explained by immigration of females from a population that has higher heterozygosity. Another measure of individual inbreeding, d2, did not differ between males and females and showed high variability within groups (Fig. 3). Few highly heterozygous loci might control individual values of d2, whereas the value of MLH is influenced by a larger number of loci (Hansson 2010), possibly leading to different results. Females within a site and from different sites were similarly related. In contrast, relatedness was higher for males from the same site for most of the calculated coefficients, suggesting some degree of differentiation from limited movement of males (Fig. 5, Supplementary Fig. 1). Similarly, our analysis of IBD supports female-biased dispersal and male philopatry, with genetic distance increasing significantly with geographic distance in males and no trend in females (Fig. 4). Overrepresentation of closely related males visiting lek sites in the mating season (Segelbacher et al. 2007; Cayuela et al. 2021) was likely avoided through standardized transect sampling.
Sex-biased dispersal occurs when one sex is more philopatric than the other and tends to breed in its natal site or social group (Greenwood 1980). Female-biased dispersal is common among most bird families, while male-biased dispersal is predominant in mammals (Greenwood 1980; Pusey 1987; Mabry et al. 2013). Using genetic data, dispersal bias can be detected by comparing markers with different inheritance (e.g. mitochondrial and nuclear) or sex-specific patterns of genetic distance for bi-parentally inherited markers (Goudet et al. 2002; Prugnolle and de Meeus 2002). Corrales and Höglund (2012) showed that dispersing females maintain gene flow in a panmictic L. tetrix population in northern Sweden. Evidence of female-biased dispersal in T. urogallus was found in the Vosges Mountains (eastern France), in the Bohemian Forest (Germany and Czech Republic), and in the Black Forest (Germany) (Segelbacher et al. 2008; Rösner et al. 2014; Cayuela et al. 2021). In contrast, Mäki-Petäys et al. (2007) found roughly equal dispersal in T. urogallus in northern Finland by looking at IBD patterns of pairwise FST among lek sites over distances up to 350 km. Factors such as dispersal rate, bias intensity, and sampling scheme affect the power of inference of sex-biased dispersal based on genetic data (Goudet et al. 2002). Additionally, different spatial scales of sampling can influence the outcome of such tests (Yannic et al. 2012; Vangestel et al. 2013).
Habitat and genetic diversity
We found no statistically significant relation between habitat parameters and genetic diversity, except for the number of genotypes per sampling site. Given the close relationship between number of genotypes and number of samples analysed, density of genotypes was used in the multiple linear regression to account for this. Several of the explanatory variables were strongly autocorrelated (Supplementary Fig. 3), partly because the sampling sites were relatively similar in these variables. For example, all sampling sites had approximately the same shape, leading to correlation of area and perimeter. Also, mean habitat suitability was similar across sites. Shape of the sampling site (perimeter/area) and the isolation measure calculated using the expert model explained differences in density of genotypes among sampling sites (Table 3). In this study, sampling sites are not isolated patches in an unsuitable matrix; instead, they are adjacent to areas with similar habitat suitability. Narrower sampling sites have larger values of the perimeter/area metric. The positive relationship between density of genotypes and this metric of shape could be explained by the larger area of contact to neighbouring potential habitat. Habitat suitability models were not included directly, but the expert model was included as part of the isolation measure. The density of genotypes was negatively affected by isolation of the sites. This result seems relevant for conservation in principle, but we note that further work is still required: firstly, in-depth analyses of the basically suitable (but unsampled) habitat connecting sampling sites to check if the number and thus relevance of possible barriers correlates with the linear distance between sites, and secondly, corroboration by additional data from the other reference areas of the monitoring project in Tyrol. Should the result be confirmed, this would be of relevance for nature conservation in the future and should be considered in habitat management planning. However, in the context of this case study, the specific result is less relevant than demonstrating the feasibility of the statistical approach, which would also be given if no significant result at all had emerged from the limited data used here.
In this study, we showed that the investigated T. urogallus population is well mixed and likely well connected to populations from neighbouring valleys. We also found sex-specific mobility patterns, supporting female-biased dispersal. We demonstrated the general feasibility of the modelling approach using habitat parameters, which will be used for the analysis of the complete monitoring dataset including three more sites and one additional species (Lyrurus tetrix). We refrain from interpreting the results presented here in detail as this is only one out of four areas of the ongoing grouse-monitoring project in Tyrol. For the analysis of data from multiple areas and years, more complex statistical instruments will be available, including random effects, which may increase the sensitivity of the approach and allow to identify more general patterns.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
References
Abrahams C (2019) Comparison between lek counts and bioacoustic recording for monitoring western Capercaillie (Tetrao urogallus L). J Ornithol 160:685–697. https://doi.org/10.1007/s10336-019-01649-8
Adamack AT, Gruber B (2014) PopGenReport: simplifying basic population genetic analyses in R. Methods Ecol Evol 5:384–387. https://doi.org/10.1111/2041-210X.12158
Aleix-Mata G, Adrados B, Boos M et al (2019) Comparing methods for estimating the abundance of western Capercaillie Tetrao urogallus males in Pyrenean leks: singing counts versus genetic analysis of non-invasive samples. Bird Study 66:565–569. https://doi.org/10.1080/00063657.2020.1720594
Andesner P, Vallant S, Seeber T et al (2021) A reference allelic ladder for western Capercaillie (Tetrao urogallus) and black grouse (Tetrao tetrix) enables linking grouse genetic data across studies. Conserv Genet Resour 13:97–105. https://doi.org/10.1007/s12686-020-01180-6
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165. https://doi.org/10.1046/j.0962-1083.2001.01436.x
Bañuelos M-J, Blanco-Fontao B, Fameli A et al (2019) Population dynamics of an endangered forest bird using mark–recapture models based on DNA-tagging. Conserv Genet 20:1251–1263. https://doi.org/10.1007/s10592-019-01208-x
Beja-Pereira A, Oliveira R, Alves PC et al (2009) Advancing ecological understandings through technological transformations in noninvasive genetics. Mol Ecol Resour 9:1279–1301. https://doi.org/10.1111/j.1755-0998.2009.02699.x
Caizergues A, Dubois S, Loiseau A et al (2001) Isolation and characterization of microsatellite loci in black grouse (Tetrao tetrix). Mol Ecol Notes 1:36–38. https://doi.org/10.1046/j.1471-8278.2000.00015.x
Cayuela H, Prunier JG, Laporte M et al (2021) Demography, genetics, and decline of a spatially structured population of lekking bird. Oecologia. https://doi.org/10.1007/s00442-020-04808-4
Chessel D, Dufour AB, Thioulouse J (2004) The ade4 package—I: one-table methods. R News 4:6
Corrales C, Höglund J (2012) Maintenance of gene flow by female-biased dispersal of black grouse Tetrao tetrix in northern Sweden. J Ornithol 153:1127–1139. https://doi.org/10.1007/s10336-012-0844-0
Coulson TN, Pemberton JM, Albon SD et al (1998) Microsatellites reveal heterosis in red deer. Proc R Soc B Biol Sci 265:489–495
Dvorak M, Landmann A, Teufelbauer N et al (2017) The conservation status of the breeding birds of Austria: Red List (5th version) and birds of conservation concern (1st version). 38
Edwards AWF (1971) Distances between populations on the basis of gene frequencies. Biometrics 27:873–881. https://doi.org/10.2307/2528824
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Francis RM (2017) Pophelper: an R package and web app to analyse and visualize population structure. Mol Ecol Resour 17:27–32. https://doi.org/10.1111/1755-0998.12509
Galpern P, Manseau M, Hettinga P et al (2012) Allelematch: an R package for identifying unique multilocus genotypes where genotyping error and missing data may be present. Mol Ecol Resour 12:771–778. https://doi.org/10.1111/j.1755-0998.2012.03137.x
Goudet J (2005) Hierfstat, a package for R to compute and test hierarchical F-statistics. Mol Ecol Notes 5:184–186
Goudet J, Perrin N, Waser P (2002) Tests for sex-biased dispersal using bi-parentally inherited genetic markers. Mol Ecol 11:1103–1114. https://doi.org/10.1046/j.1365-294X.2002.01496.x
Greenwood PJ (1980) Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28:1140–1162. https://doi.org/10.1016/S0003-3472(80)80103-5
Haider M, Steixnr R, Zeni T, Vallant S, Lentner R, Schlick-Steiner BC, Steiner FM (2023) The influence of sample size on two approaches to estimate black grouse (Lyrurus tetrix) population size using non-invasive sampling (In preparation)
Hanski I, Kuussaari M, Nieminen M (1994) Metapopulation structure and migration in the butterfly Melitaea cinxia. Ecology 75:747–762. https://doi.org/10.2307/1941732
Hansson B (2010) The use (or misuse) of microsatellite allelic distances in the context of inbreeding and conservation genetics. Mol Ecol 19:1082–1090. https://doi.org/10.1111/j.1365-294X.2010.04556.x
Hijmans RJ, Karney (GeographicLib) C, Williams E, Vennes C (2019) geosphere: spherical trigonometry. Version 1.5-10. https://CRAN.R-project.org/package=geosphere
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70. https://www.jstor.org/stable/4615733
IUCN (2016) Tetrao urogallus. BirdLife International: The IUCN Red List of Threatened Species 2016:e.T22679487A85942729. https://doi.org/10.2305/IUCN.UK.2016-3.RLTS.T22679487A85942729.en
Jacob G, Debrunner R, Gugerli F et al (2010) Field surveys of capercaillie (Tetrao urogallus) in the Swiss Alps underestimated local abundance of the species as revealed by genetic analyses of non-invasive samples. Conserv Genet 11:33–44. https://doi.org/10.1007/s10592-008-9794-8
Jombart T (2008) Adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24:1403–1405. https://doi.org/10.1093/bioinformatics/btn129
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2:e281. https://doi.org/10.7717/peerj.281
Kassambara A (2023) rstatix: pipe-friendly framework for basic statistical tests. Version 0.7.2. https://CRAN.R-project.org/package=rstatix
Keller V, Herrando S, Voříšek P et al (2020) European breeding Bird Atlas 2: distribution, abundance and change. European Bird Census Council & Lynx Edicions, Barcelona
Lehne F (2014) Habitatmodellierung und -charakterisierung der Lebensraumeigenschaften von Raufußhühnern in den Tiroler Alpen. Universität Innsbruck
Lentner R, Lehne F, Vallant S, Masoner A (2017) Raufußhühner-Monitoring Tirol—Monitoringperiode 2011–2014. Bericht Land Tirol
Lentner R, Masoner A, Lehne F (2018) Sind Zählungen an Balzplätzen von Auer- und Birkhühnern noch zeitgemäß? Ergebnisse aus dem Raufußhühner-Monitoring tirol. Ornithol Beob 115:24
Lentner R, Lehne F, Danzl A, Eberhard E (2022) Atlas der Brutvögel Tirols. Berenkamp Verlag, Wattens
Lynch M, Ritland K (1999) Estimation of pairwise relatedness with molecular markers. Genetics 152:1753–1766. https://doi.org/10.1093/genetics/152.4.1753
Mabry KE, Shelley EL, Davis KE et al (2013) Social mating system and sex-biased dispersal in mammals and birds: a phylogenetic analysis. PLoS ONE 8:e57980. https://doi.org/10.1371/journal.pone.0057980
Mäki-Petäys H, Corander J, Aalto J et al (2007) No genetic evidence of sex-biased dispersal in a lekking bird, the capercaillie (Tetrao urogallus). J Evol Biol 20:865–873. https://doi.org/10.1111/j.1420-9101.2007.01314.x
Marko PB, Hart MW (2011) The complex analytical landscape of gene flow inference. Trends Ecol Evol 26:448–456. https://doi.org/10.1016/j.tree.2011.05.007
Masoner A (2012) Wissenschaftliche Begleituntersuchungen am Auerhuhn (Tetrao urogallus) im Rahmen des Raufusshuhn-Monitoringprojektes Tirol (östliches und westlichen Achental). Univ. Innsbruck
Mollet P, Kéry M, Gardner B et al (2015) Estimating population size for capercaillie (Tetrao urogallusL.) with spatial capture-recapture models based on genotypes from one field sample. PLoS ONE 10:e0129020. https://doi.org/10.1371/journal.pone.0129020
Morán-Luis M, Fameli A, Blanco-Fontao B et al (2014) Demographic status and genetic tagging of endangered vapercaillie in NW Spain. PLoS ONE 9:e99799. https://doi.org/10.1371/journal.pone.0099799
Pakkala T, Pellikka J, Lindén H (2003) Capercaillie Tetrao urogallus—a good candidate for an umbrella species in taiga forests. Wildl Biol 9:309–316. https://doi.org/10.2981/wlb.2003.019
Pennell MW, Stansbury CR, Waits LP, Miller CR (2013) Capwire: a R package for estimating population census size from non-invasive genetic sampling. Mol Ecol Resour 13:154–157. https://doi.org/10.1111/1755-0998.12019
Pew J, Muir PH, Wang J, Frasier TR (2015) Related: an R package for analysing pairwise relatedness from codominant molecular markers. Mol Ecol Resour 15:557–561. https://doi.org/10.1111/1755-0998.12323
Phillips SJ, Dudík M, Schapire RE (2004) A maximum entropy approach to species distribution modeling. In: Proceedings of the twenty-first international conference on machine learning. Association for Computing Machinery, New York, p 83
Piertney SB, Höglund J (2001) Polymorphic microsatellite DNA markers in black grouse (Tetrao tetrix). Mol Ecol Notes 1:303–304. https://doi.org/10.1046/j.1471-8278.2001.00118.x
Pritchard JK, Stephens M, Donnelly P (2000) Inference of Population structure using multilocus genotype data. Genetics 155:945–959. https://doi.org/10.1093/genetics/155.2.945
Prugnolle F, de Meeus T (2002) Inferring sex-biased dispersal from population genetic tools: a review. Heredity 88:161–165. https://doi.org/10.1038/sj.hdy.6800060
Pusey AE (1987) Sex-biased dispersal and inbreeding avoidance in birds and mammals. Trends Ecol Evol 2:295–299. https://doi.org/10.1016/0169-5347(87)90081-4
Queller DC, Goodnight KF (1989) Estimating relatedness using genetic markers. Evolution 43:258–275. https://doi.org/10.2307/2409206
R Core Team (2023) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna. https://www.R-project.org/
Rabasa SG, Gutiérrez D, Escudero A (2007) Metapopulation structure and habitat quality in modelling dispersal in the butterfly Iolana iolas. Oikos 116:793–806. https://doi.org/10.1111/j.0030-1299.2007.15788.x
Roberge J-M, Angelstam P (2004) Usefulness of the umbrella species concept as a conservation tool. Conserv Biol 18:76–85
Rösner S, Brandl R, Segelbacher G et al (2014) Noninvasive genetic sampling allows estimation of capercaillie numbers and population structure in the Bohemian Forest. Eur J Wildl Res 60:789–801. https://doi.org/10.1007/s10344-014-0848-6
Rousset F (2008) Genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Mol Ecol Resour 8:103–106. https://doi.org/10.1111/j.14718286.2007.01931.x
Segelbacher G, Storch I (2002) Capercaillie in the Alps: genetic evidence of metapopulation structure and population decline. Mol Ecol 11:1669–1677. https://doi.org/10.1046/j.1365-294X.2002.01565.x
Segelbacher G, Paxton RJ, Steinbrück G, et al (2000) Characterization of microsatellites in capercaillie Tetrao urogallus (AVES). Mol Ecol 9:1934–1935. https://doi.org/10.1046/j.1365-294x.2000.0090111934.x
Segelbacher G, Wegge P, Sivkov AV, Höglund J (2007) Kin groups in closely spaced capercaillie leks. J Ornithol 148:79–84. https://doi.org/10.1007/s10336-006-0103-3
Segelbacher G, Manel S, Tomiuk J (2008) Temporal and spatial analyses disclose consequences of habitat fragmentation on the genetic diversity in capercaillie (Tetrao urogallus). Mol Ecol 17:2356–2367. https://doi.org/10.1111/j.1365-294X.2008.03767.x
Simberloff D (1998) Flagships, umbrellas, and keystones: is single-species management passé in the landscape era? Biol Conserv 83:247–257. https://doi.org/10.1016/S0006-3207(97)00081-5
Storch I (1997) Male territoriality, female range use, and spatial organisation of capercaillie Tetrao urogallus leks. Wildl Biol 3:149–161. https://doi.org/10.2981/wlb.1997.019
Storch I (2007) Grouse: status survey and conservation action plan 2006–2010. IUCN, Gland
Suter W, Graf RF, Hess R (2002) Capercaillie (Tetrao urogallus) and avian biodiversity: testing the umbrella-species concept. Conserv Biol 16:778–788. https://doi.org/10.1046/j.1523-1739.2002.01129.x
Taberlet P, Griffin S, Goossens B et al (1996) Reliable genotyping of samples with very low DNA quantities using PCR. Nucleic Acids Res 24:3189–3194. https://doi.org/10.1093/nar/24.16.3189
Vallant S, Niederstätter H, Berger B et al (2018) Increased DNA typing success for feces and feathers of capercaillie (Tetrao urogallus) and black grouse (Tetrao tetrix). Ecol Evol 8:3941–3951. https://doi.org/10.1002/ece3.3951
Vangestel C, Callens T, Vandomme V, Lens L (2013) Sex-biased dispersal at different geographical scales in a cooperative breeder from fragmented rainforest. PLoS ONE 8:e71624. https://doi.org/10.1371/journal.pone.0071624
Waits LP, Luikart G, Taberlet P (2001) Estimating the probability of identity among genotypes in natural populations: cautions and guidelines. Mol Ecol 10:249–256. https://doi.org/10.1046/j.1365-294x.2001.01185.x
Wilberg MJ, Dreher BP (2004) GENECAP: a program for analysis of multilocus genotype data for non-invasive sampling and capture-recapture population estimation. Mol Ecol Notes 4:783–785. https://doi.org/10.1111/j.1471-8286.2004.00797.x
Yannic G, Basset P, Büchi L et al (2012) Scale-specific sex-biased dispersal in the Valais shrew unveiled by genetic variation on the Y chromosome, autosomes, and mitochondrial DNA. Evol Int J Org Evol 66:1737–1750. https://doi.org/10.1111/j.1558-5646.2011.01554.x
Acknowledgements
We thank Franz Goller, Gunther Gressmann, Martin Kurzthaler, Felix Lassacher, Florian Lehne, Alois Masoner, Gregor Schartner, Ramona Steixner, and Gerald Wille for help in collecting the samples. We are also thankful to Michael Haupolter, who provided the habitat suitability models, and Harald Niederstätter, who designed the primers for sex determination. We thank Philipp Andesner, Florian Reischer, Theresia Telser, and Elisabeth Zangerl for assistance in the laboratory. The Bayesian clustering analysis was done using the LEO HPC infrastructure of the University of Innsbruck. We thank the Provincial Government of Tyrol, Department for Hunting and Fishing, and the Tyrolean Hunters’ Association for the cooperation in the grouse-monitoring project and providing data.
Funding
Open access funding provided by University of Innsbruck and Medical University of Innsbruck. The monitoring is financed by the Provincial Government of Tyrol, Department for Hunting and Fishing, and the Tyrolean Hunters’ Association.
Author information
Authors and Affiliations
Contributions
TZ and MH analysed data. TZ wrote the first draft of the manuscript. MH finalised the manuscript with contributions from all other authors. SV contributed to the analysis. TZ, SV, and RL contributed to fieldwork. RL, FMS and BCS initiated the study. FMS and BCS supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All analyses were in accord with the nature-conservation regulations for Tyrol, Austria.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeni, T., Haider, M., Vallant, S. et al. Towards a standardised set of data analyses for long-term genetic monitoring of grouse using non-invasive sampling: a case study on western capercaillie. Conserv Genet 25, 75–86 (2024). https://doi.org/10.1007/s10592-023-01552-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-023-01552-z