Abstract
Understanding the spatial context of genetic variation for species at risk is important for effective management and long-term survival of the species. We use multilocus microsatellite data to investigate the population genetic structure of the spotted gar (Lepisosteus oculatus) across its northern range edge in Canada. We then compare these northern individuals with samples taken from the southern core of the species range. For the northern samples, significant genetic differentiation among groups of individuals forming two major genetically distinct populations, and as many as 7–9 smaller subpopulations, was recovered using hierarchical Bayesian assignment methods and non-equilibrial discriminant function analyses. Spatial genetic variation is present, particularly at higher hierarchical groupings; however, some population admixture at sites is evident and is indicative of dispersal and gene flow among some locations or shared ancestry. Gene flow estimates among populations and subpopulations is very low, ranging from essentially complete isolation to as high as 5 %—suggesting that mechanisms in addition to geographic isolation are operating to create genetic structure. In Lake Erie, the physical isolation of Point Pelee appears to confer distinct genetic differentiation for those populations and provide a source of genetic variation for Lake Erie proper when breaches to the barrier beach occur. Results indicate that the northern edge populations are distinct from southern populations and should be conserved to maintain the overall genetic diversity of this species. Additionally, the asymmetrical genetic connectivity among the Point Pelee and Rondeau Bay sites highlights the sensitivity of Point Pelee to environmental perturbation and habitat degradation.


Similar content being viewed by others
References
Bailey RM, Smith GR (1981) Origin and geography of the fish fauna of the Laurentian Great Lakes. Can J Fish Aquat Sci 38:1539–1561
Bouvier LD, Mandrak NE (2010) Information in support of a recovery potential assessment of spotted gar (Lepisosteus oculatus) in Canada. DFO Can Sci Advis Sec Res Doc. 2010/079 vi+22 p
COSEWIC (2005) COSEWIC assessment and update status report on the spotted gar Lepisosteus oculatus in Canada. Committee on the status of endangered wildlife in Canada, Ottawa
COSEWIC (2010) COSEWIC’s assessment process and criteria. Committee on the status of endangered wildlife in Canada, Ottawa
Dent EA, vonHoldt BM (2012) Structure harvester: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. doi:10.1007/s12686-011-9548-7
Dieringer D, Schlötterer C (2003) Microsatellite analyzer (MSA): a platform independent analysis tool for large microsatellite datasets. Mol Ecol Notes 3:167–169
Elphinstone MS, Hinten GN, Anderson MJ, Nock CJ (2003) An inexpensive and high-throughput procedure to extract and purify total genomic DNA for population studies. Mol Ecol Notes 3:317–320
Evanno G, Regnant S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetic data analysis. Evol Bioinform Online 1:47–50
Faubet P, Gaggiotti OE (2008) A new Bayesian method to identify the environmental factors that influence recent migration. Genetics 178:1491–1504
Ferrara A (2001) Life history strategy of Lepisosteidae: implications for the conservation and management of alligator gar. PhD Thesis, Auburn University, Auburn
Franklin IR (1980) Evolutionary change in small populations. In: Soule ME, Wilcox BA (eds) Conservation biology—an evolutionary—ecological perspective. Sinauer Asscociates, Sunderland, pp 135–149
Garcia-Ramos G, Kirkpatrick M (1997) Models of adaptation and gene flow in peripheral populations. Evolution 5:21–28
Glass WR (2012) Living on the edge: conservation of fish species at risk in Canada. PhD Thesis, University of Windsor, Windsor, Canada
Glass WR, Mandrak NE (2014) Distribution of spotted gar (Lepisosteus oculatus) adults and juveniles in the Rondeau Bay, Long Point Bay, and Hamilton Harbour watersheds. Can Manuscr Rep Fish Aquat Sci 3048:iii+21
Glass WR, Corkum LD, Mandrak NE (2011) Pectoral fin ray aging: an evaluation of a non-lethal method for aging gars and its application to a population of the threatened spotted gar. Environ Biol Fish 90:235–242
Glass WR, Corkum LD, Mandrak NE (2012) Spring and summer distribution and habitat use by adult threatened spotted gar in Rondeau Bay, Ontario, using radiotelemetry. Trans Am Fish Soc 141:1026–1035
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Jelks HL, Walsh SJ, Burkhead NM, Contreras-Balders S, Diaz-Pard E, Hendrickson DA, Lyons J, Mandrak NE, McCormick F, Nelson JS, Platania SP, Porter BA, Renaud CB, Schmitter-Soto JJ, Taylor EB, Warren ML Jr (2008) Conservation of imperiled North American freshwater and diadromous fishes. Fisheries 33:372–407
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet 11:94
Larson G, Schaetzl R (2001) Origin and evolution of the Great Lakes. J Great Lakes Res 27(518):546
Moyer GR, Sloss BL, Kreiser BR, Feldheim KA (2009) Isolation and characterization of microsatellite loci for alligator gar (Atractosteus spatula) and their variability in two other species (Lepisosteus oculatus and L. osseus) of Lepisosteidae. Mol Ecol Res 9:963–966
Munwes I, Geffen E, Roll U, Friedmann A, Daya A, Tikochinski Y, Gafny S (2010) The change in genetic diversity down the core-edge gradient in the eastern spadefoot toad (Pelobates syriacus). Mol Ecol 19:2675–2689
NatureServe (2014) www.natureserve.org. Accessed July 2, 2014
Page LM, Burr BM (2011) A field guide to freshwater fishes of North America north of Mexico, 2nd edn. Houghton Mifflin Company, Boston
Pritchard JK, Stevens M, Donnely P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rambaut A, Drummond AJ (2007) Tracer v1.4. Available from http://beast.bio.ed.ac.uk/Tracer
Redmond LC (1964) Ecology of the spotted gar (Lepisosteus oculatus Winchell) in southeastern Missouri. MSc thesis, University of Missouri, Columbia
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rodriguez-Munoz R, Mirol PM, Segelbacher G, Fernandez A, Tregenza T (2007) Genetic differentiation of an endangered capercaillie (Tetrao urogallus) population at the Southern edge of the species range. Conserv Genet 8:659–670
Rosenberg N (2004) Distruct: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Staton SK, Boyko AL, Dunn SE, Burridge M (2012) Recovery strategy for the spotted gar (Lepisosteus oculatus) in Canada. Species at risk act recovery strategy series. Fisheries and Oceans Canada, Ottawa
Stepien CA, Faber JE (1998) Population genetic structure, phylogeography and spawning philopatry in Walleye (Stizostedion vitreum) from mitochondrial DNA control region sequences. Mol Ecol 7:1757–1769
Surette HJ (2006) Processes influencing temporal variation in fish species composition in Point Pelee National Park. MSc thesis, University of Guelph, Geulph
Waples RS (2005) Genetic estimates of effective population size: to what time periods do the estimates apply? Mol Ecol 14:3335–3352
Waples RS, Do C (2008) LDNE: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Resour 8:753–756
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evolut Appl 3:244–262
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Wilson GA, Rannala B (2003) Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163:1177–1191
Young, JAM, Koops MA (2010) Recovery potential modelling of spotted gar (Lepisosteus oculatus) in Canada. DFO Can Sci Advis Sec Res Doc 2010/078
Acknowledgments
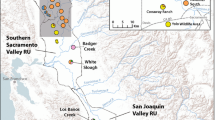
We thank Hank Bart, David Rowe, and Mathew Spickard for providing us with tissue samples. We also thank Aaron Simpson, Ben Meunier, Krista Lajevardi, Solomon David, Tim Clay, Mark Suchy, Cynthia Fox, Allyse Ferrara, Bob Hrabik, and Jason Crites for assistance with field collections and Thom Heiman for assistance with aging of the specimens. Funding was provided by the Fisheries and Oceans Canada Species at Risk Program awarded to NEM and NSERC funding to DDH. The basemap used to create Fig. 1 was provided by freevectormaps.com. We thank our anonymous reviewers for their constructive comments that have helped to improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10592_2015_708_MOESM1_ESM.tif
Supplementary Fig. 1. Population genetic structure of spotted gar determined by Bayesian clustering assignment using STRUCTURE without location information, showing 10 resolved populations throughout North America. Southern group population substructure for K = 2 and K = 3, and northern group substructure for K = 2 and K = 3 are also shown along with Lake Erie (Point Pelee and Rondeau Bay) substructure. The coloured bubbles correspond to the respective recovered population, with effective number of breeding individuals (N b) in italics; arrows indicate direction of gene flow with accompanying migration estimates
Rights and permissions
About this article
Cite this article
Glass, W.R., Walter, R.P., Heath, D.D. et al. Genetic structure and diversity of spotted gar (Lepisosteus oculatus) at its northern range edge: implications for conservation. Conserv Genet 16, 889–899 (2015). https://doi.org/10.1007/s10592-015-0708-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-015-0708-2