Abstract
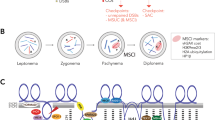
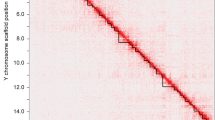
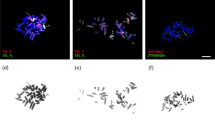
Double-strand break repair during meiosis is normally achieved using the homologous chromosome as a repair template. Heteromorphic sex chromosomes share little sequence homology, presenting unique challenges to the repair of double-strand breaks. Our understanding of how heteromorphic sex chromosomes behave during meiosis has been focused on ancient sex chromosomes, where the X and Y differ markedly in overall structure and gene content. It remains unclear how more recently evolved sex chromosomes that share considerably more sequence homology with one another pair and form double-strand breaks. One possibility is barriers to pairing evolve rapidly. Alternatively, recently evolved sex chromosomes may exhibit pairing and double-strand break repair that more closely resembles that of their autosomal ancestors. Here, we use the recently evolved X and Y chromosomes of the threespine stickleback fish (Gasterosteus aculeatus) to study patterns of pairing and double-stranded break formation using molecular cytogenetics. We found that the sex chromosomes of threespine stickleback fish did not pair exclusively in the pseudoautosomal region. Instead, the chromosomes fully paired in a non-homologous fashion. To achieve this, the X chromosome underwent synaptic adjustment during pachytene to match the axis length of the Y chromosome. Double-strand break formation and repair rate also matched that of the autosomes. Our results highlight that recently evolved sex chromosomes exhibit meiotic behavior that is reminiscent of autosomes and argues for further work to identify the homologous templates that are used to repair double-strand breaks on the X and Y chromosomes.






Similar content being viewed by others
Abbreviations
- DSB:
-
Double-strand breaks
- FISH:
-
Fluorescent in situ hybridization
- Idh :
-
Isocitrate dehydrogenase
- RAD51:
-
RAD51 recombinase
- SMC3:
-
Structural maintenance of chromosomes 3
References
Ashley T, Plug AW, Xu J et al (1995) Dynamic changes in Rad51 distribution on chromatin during meiosis in male and female vertebrates. Chromosoma 104:19–28. https://doi.org/10.1007/bf00352222
Bachtrog D (2013) Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat Rev Genet 14:113–124. https://doi.org/10.1038/nrg3366
Barlow AL, Benson FE, West SC, Hultén MA (1997) Distribution of the Rad51 recombinase in human and mouse spermatocytes. Embo J 16:5207–5215. https://doi.org/10.1093/emboj/16.17.5207
Baudat F, Manova K, Yuen JP et al (2000) Chromosome synapsis defects and sexually dimorphic meiotic progression in mice lacking Spo11. Mol Cell 6:989–998. https://doi.org/10.1016/s1097-2765(00)00098-8
Bellott DW, Hughes JF, Skaletsky H et al (2014) Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508:494–499. https://doi.org/10.1038/nature13206
Bergero R, Gardner J, Bader B et al (2019) Exaggerated heterochiasmy in a fish with sex-linked male coloration polymorphisms. Proc National Acad Sci 116:6924–6931. https://doi.org/10.1073/pnas.1818486116
Blokhina YP, Nguyen AD, Draper BW, Burgess SM (2019) The telomere bouquet is a hub where meiotic double-strand breaks, synapsis, and stable homolog juxtaposition are coordinated in the zebrafish. Danio Rerio Plos Genet 15:e1007730. https://doi.org/10.1371/journal.pgen.1007730
Borg B (1982) Seasonal effects of photoperiod and temperature on spermatogenesis and male secondary sexual characters in the three-spined stickleback, Gasterosteus aculeatus L. Can J Zool 60:3377–3386
Borodin PM, Basheva EA, Torgasheva AA et al (2011) Multiple independent evolutionary losses of XY pairing at meiosis in the grey voles. Chromosome Res 20:259–268. https://doi.org/10.1007/s10577-011-9261-0
Burgoyne PS (1982) Genetic homology and crossing over in the X and Y chromosomes of Mammals. Hum Genet 61:85–90
Carrasco LAP, Penman DJ, Bromage N (1999) Evidence for the presence of sex chromosomes in the Nile tilapia (Oreochromis niloticus) from synaptonemal complex analysis of XX, XY and YY genotypes. Aquaculture 173:207–218. https://doi.org/10.1016/s0044-8486(98)00488-8
Chandley AC, Goetz P, Hargreave TB et al (1984) On the nature and extent of XY pairing at meiotic prophase in man. Cytogenet Genome Res 38:241–247. https://doi.org/10.1159/000132070
Charlesworth D, Bergero R, Graham C et al (2021) How did the guppy Y chromosome evolve? Plos Genet 17:e1009704. https://doi.org/10.1371/journal.pgen.1009704
Checchi PM, Engebrecht J (2011) Heteromorphic sex chromosomes: navigating meiosis without a homologous partner. Mol Reprod Dev 78:623–632. https://doi.org/10.1002/mrd.21369
Connallon T, Clark AG (2010) Gene duplication, gene conversion and the evolution of the Y chromosome. Genetics 186:277–286. https://doi.org/10.1534/genetics.110.116756
Conte MA, Gammerdinger WJ, Bartie KL et al (2017) A high quality assembly of the Nile Tilapia (Oreochromis niloticus) genome reveals the structure of two sex determination regions. BMC Genomics 18:341. https://doi.org/10.1186/s12864-017-3723-5
Cortez D, Cortez D, Marin R et al (2014) Origins and functional evolution of Y chromosomes across mammals. Nature 508:488–493. https://doi.org/10.1038/nature13151
Craig-Bennett A (1931) The reproductive cycle of the three-spined stickleback, Gasterosteus aculeatus. Linn Phil Trans Roy Soc B 219:197–279. https://doi.org/10.2307/92176
Cuñado N, Barrios J, Miguel ES et al (2002) Synaptonemal complex analysis in oocytes and spermatocytes of threespine stickleback Gasterosteus aculeatus (Teleostei, Gasterosteidae). Genetica 114:53–56. https://doi.org/10.1023/a:1014685114092
Darolti I, Wright AE, Mank JE (2020) Guppy Y chromosome integrity maintained by incomplete recombination suppression. Genome Biol Evol 12:evaa099. https://doi.org/10.1093/gbe/evaa099
Dawe RK, Lowry EG, Gent JI et al (2018) A kinesin-14 motor activates neocentromeres to promote meiotic drive in maize. Cell 173:839-850.e18. https://doi.org/10.1016/j.cell.2018.03.009
Dumont BL, Payseur BA (2011) Genetic analysis of genome-scale recombination rate evolution in house mice. PLoS Genet 7:e1002116. https://doi.org/10.1371/journal.pgen.1002116
Dumont BL, Williams CL, Ng BL et al (2018) Relationship between sequence homology, genome architecture, and meiotic behavior of the sex chromosomes in North American voles. Genetics 210:83–97. https://doi.org/10.1534/genetics.118.301182
Enguita-Marruedo A, Martín-Ruiz M, García E et al (2019) Transition from a meiotic to a somatic-like DNA damage response during the pachytene stage in mouse meiosis. Plos Genet 15:e1007439. https://doi.org/10.1371/journal.pgen.1007439
Federici F, Mulugeta E, Schoenmakers S et al (2015) Incomplete meiotic sex chromosome inactivation in the domestic dog. BMC Genomics 16:291. https://doi.org/10.1186/s12864-015-1501-9
de la Fuente R, Parra MT, Viera A et al (2007) Meiotic pairing and segregation of achiasmate sex chromosomes in eutherian mammals: the role of SYCP3 protein. Plos Genet 3:e198. https://doi.org/10.1371/journal.pgen.0030198
de la Fuente R, Sánchez A, Marchal JA et al (2012) A synaptonemal complex-derived mechanism for meiotic segregation precedes the evolutionary loss of homology between sex chromosomes in arvicolid mammals. Chromosoma 121:433–446. https://doi.org/10.1007/s00412-012-0374-9
Gammerdinger WJ, Conte MA, Acquah EA et al (2014) Structure and decay of a proto-Y region in tilapia, Oreochromis niloticus. Bmc Genomics 15:975. https://doi.org/10.1186/1471-2164-15-975
Goedecke W, Eijpe M, Offenberg HH et al (1999) Mre11 and Ku70 interact in somatic cells, but are differentially expressed in early meiosis. Nat Genet 23:194–198. https://doi.org/10.1038/13821
Grelon M, Vezon D, Gendrot G, Pelletier G (2001) AtSPO11-1 is necessary for efficient meiotic recombination in plants. EMBO J 20:589–600
Gruhn JR, Rubio C, Broman KW et al (2013) Cytological studies of human meiosis: sex-specific differences in recombination originate at, or prior to, establishment of double-strand breaks. PLoS ONE 8:e85075. https://doi.org/10.1371/journal.pone.0085075
Guioli S, Lovell-Badge R, Turner JMA (2012) Error-prone ZW pairing and no evidence for meiotic sex chromosome inactivation in the chicken germ line. Plos Genet 8:e1002560. https://doi.org/10.1371/journal.pgen.1002560
Hassold T, Hunt P (2001) To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2:280–291. https://doi.org/10.1038/35066065
Hassold TJ, Sherman SL, Pettay D et al (1991) XY chromosome nondisjunction in man is associated with diminished recombination in the pseudoautosomal region. Am J Hum Genet 49:253–260
Hunter N, Kleckner N (2001) The single-end invasion an asymmetric intermediate at the double-strand break to double-holliday junction transition of meiotic recombination. Cell 106:59–70. https://doi.org/10.1016/s0092-8674(01)00430-5
Inagaki A, Schoenmakers S, Baarends WM (2010) DNA double strand break repair, chromosome synapsis and transcriptional silencing in meiosis. Epigenetics 5:255–266. https://doi.org/10.4161/epi.5.4.11518
Kauppi L, Barchi M, Baudat F et al (2011) Distinct properties of the XY pseudoautosomal region crucial for male meiosis. Science 331:916–920. https://doi.org/10.1126/science.1195774
Kauppi L, Jasin M, Keeney S (2012) The tricky path to recombining X and Y chromosomes in meiosis. Ann Ny Acad Sci 1267:18–23. https://doi.org/10.1111/j.1749-6632.2012.06593.x
Kleckner N, Storlazzi A, Zickler D (2003) Coordinate variation in meiotic pachytene SC length and total crossover/chiasma frequency under conditions of constant DNA length. Trends Genet 19:623–628. https://doi.org/10.1016/j.tig.2003.09.004
Lange J, Yamada S, Tischfield SE et al (2016) The landscape of mouse meiotic double-strand break formation, processing, and repair. Cell 167:695–708. https://doi.org/10.1016/j.cell.2016.09.035
Lao JP, Hunter N (2010) Trying to avoid your sister. Plos Biol 8:e1000519. https://doi.org/10.1371/journal.pbio.1000519
Lisachov AP, Zadesenets KS, Rubtsov NB, Borodin PM (2015) Sex chromosome synapsis and recombination in male guppies. Zebrafish 12:174–180. https://doi.org/10.1089/zeb.2014.1000
Lu L-Y, Yu X (2015) Double-strand break repair on sex chromosomes: challenges during male meiotic prophase. Cell Cycle 14:516–525. https://doi.org/10.1080/15384101.2014.998070
Lukassen S, Bosch E, Ekici AB, Winterpacht A (2018) Characterization of germ cell differentiation in the male mouse through single-cell RNA sequencing. Sci Rep 8:6521. https://doi.org/10.1038/s41598-018-24725-0
MacQueen AJ, Phillips CM, Bhalla N et al (2005) Chromosome sites play dual roles to establish homologous synapsis during meiosis in C. elegans. Cell 123:1037–1050. https://doi.org/10.1016/j.cell.2005.09.034
Mahadevaiah SK, Turner JMA, Baudat F et al (2001) Recombinational DNA double-strand breaks in mice precede synapsis. Nat Genet 27:271–276. https://doi.org/10.1038/85830
Marais GAB, Marais GAB, Campos PRA et al (2010) Can intra-Y gene conversion oppose the degeneration of the human Y chromosome? A simulation study. Genome Biol Evol 2:347–357. https://doi.org/10.1093/gbe/evq026
Moens PB, Chen DJ, Shen Z et al (1997) Rad51 immunocytology in rat and mouse spermatocytes and oocytes. Chromosoma 106:207–215. https://doi.org/10.1007/s004120050241
Moses MJ, Poorman PA, Roderick TH, Davisson MT (1982) Synaptonemal complex analysis of mouse chromosomal rearrangements. IV. Synapsis and synaptic adjustment in two paracentric inversions. Chromosoma 84:457–474. https://doi.org/10.1007/bf00292848
Naftaly AS, Pau S, White MA (2021) Long-read RNA sequencing reveals widespread sex-specific alternative splicing in threespine stickleback fish. Genome Res 31:1486–1497. https://doi.org/10.1101/gr.274282.120
Nanda I, Schories S, Simeonov I et al (2022) Evolution of the degenerated Y-chromosome of the swamp guppy. Micropoecilia Picta Cells 11:1118. https://doi.org/10.3390/cells11071118
Page J, Berríos S, Parra MT et al (2005) The program of sex chromosome pairing in meiosis is highly conserved across marsupial species implications for sex chromosome evolution. Genetics 170:793–799. https://doi.org/10.1534/genetics.104.039073
Page J, de la Fuente R, Gómez R et al (2006) Sex chromosomes, synapsis, and cohesins: a complex affair. Chromosoma 115:250–259. https://doi.org/10.1007/s00412-006-0059-3
Peichel CL, McCann SR, Ross JA et al (2020) Assembly of the threespine stickleback Y chromosome reveals convergent signatures of sex chromosome evolution. Genome Biol 21:177. https://doi.org/10.1186/s13059-020-02097-x
Peichel CL, Ross JA, Matson CK et al (2004) the master sex-determination locus in threespine sticklebacks is on a nascent Y chromosome. Curr Biol 14:1416–1424. https://doi.org/10.1016/j.cub.2004.08.030
Pratto F, Brick K, Khil P et al (2014) Recombination initiation maps of individual human genomes. Science 346:1256442. https://doi.org/10.1126/science.1256442
Rasmussen SW, Holm PB (1978) Human meiosis II. Chromosome pairing and recombination nodules in human spermatocytes. Carlsberg Res Commun 43:275. https://doi.org/10.1007/bf02906106
Roesti M, Moser D, Berner D (2013) Recombination in the threespine stickleback genome-patterns and consequences. Mol Ecol 22:3014–3027. https://doi.org/10.1111/mec.12322
Romanienko PJ, Camerini-Otero RD (2000) The mouse Spo11 gene is required for meiotic chromosome synapsis. Mol Cell 6:975–987. https://doi.org/10.1016/s1097-2765(00)00097-6
Ross JA, Peichel CL (2008) Molecular cytogenetic evidence of rearrangements on the Y chromosome of the threespine stickleback fish. Genetics 179:2173–2182. https://doi.org/10.1534/genetics.108.088559
Ross JA, Urton JR, Boland J et al (2009) Turnover of sex chromosomes in the stickleback fishes (Gasterosteidae). PLOS Genet 5:e1000391. https://doi.org/10.1371/journal.pgen.1000391
Rosser ZH, Balaresque P, Jobling MA (2009) Gene conversion between the X chromosome and the male-specific region of the Y chromosome at a translocation hotspot. Am J Hum Genetics 85:130–134. https://doi.org/10.1016/j.ajhg.2009.06.009
Rouyer F, Simmler M-C, Johnsson C et al (1986) A gradient of sex linkage in the pseudoautosomal region of the human sex chromosomes. Nature 319:291–295. https://doi.org/10.1038/319291a0
Sakamoto T, Innan H (2021) Muller’s ratchet of the Y chromosome with gene conversion. Genetics 220:iyab204. https://doi.org/10.1093/genetics/iyab204
Sardell JM, Cheng C, Dagilis AJ et al (2018) Sex differences in recombination in sticklebacks. G3 8:1971–1983. https://doi.org/10.1534/g3.118.200166
Sardell JM, Kirkpatrick M (2019) Sex differences in the recombination landscape. Am Nat 195:361–379. https://doi.org/10.1086/704943
Schoenmakers S, Wassenaar E, Hoogerbrugge JW et al (2009) Female meiotic sex chromosome inactivation in chicken. Plos Genet 5:e1000466. https://doi.org/10.1371/journal.pgen.1000466
Solari AJ (1992) Equalization of Z and W axes in chicken and quail oocytes. Cytogenet Genome Res 59:52–56. https://doi.org/10.1159/000133199
Solari AJ (1974) The behavior of the XY pair in mammals. Int Rev Cytol 38:273–317. https://doi.org/10.1016/s0074-7696(08)60928-6
Solari AJ (1970) The spatial relationship of the X and Y chromosomes during meiotic prophase in mouse spermatocytes. Chromosoma 29:217–236. https://doi.org/10.1007/bf00326080
Soriano P, Keitges EA, Schorderet DF et al (1987) High rate of recombination and double crossovers in the mouse pseudoautosomal region during male meiosis. Proc National Acad Sci 84:7218–7220. https://doi.org/10.1073/pnas.84.20.7218
Tease C, Hultén MA (2004) Inter-sex variation in synaptonemal complex lengths largely determine the different recombination rates in male and female germ cells. Cytogenet Genome Res 107:208–215. https://doi.org/10.1159/000080599
Traut W, Winking H (2001) Meiotic chromosomes and stages of sex chromosome evolution in fish: zebrafish, platyfish and guppy. Chromosome Res 9:659–672. https://doi.org/10.1023/a:1012956324417
Tres LL (1977) Extensive pairing of the XY bivalent in mouse spermatocytes as visualized by whole-mount electron microscopy. J Cell Sci 25:1–15
Trombetta B, Cruciani F, Underhill PA et al (2010) Footprints of X-to-Y gene conversion in recent human evolution. Mol Biol Evol 27:714–725. https://doi.org/10.1093/molbev/msp231
Trombetta B, Fantini G, D’Atanasio E et al (2016) Evidence of extensive non-allelic gene conversion among LTR elements in the human genome. Sci Rep 6:28710. https://doi.org/10.1038/srep28710
Trombetta B, Sellitto D, Scozzari R, Cruciani F (2014) Inter- and intraspecies phylogenetic analyses reveal extensive X-Y gene conversion in the evolution of gametologous sequences of human sex chromosomes. Mol Biol Evol 31:2108–2123. https://doi.org/10.1093/molbev/msu155
Turner JMA (2007) Meiotic sex chromosome inactivation. Development (Cambridge, England) 134:1823–1831. https://doi.org/10.1242/dev.000018
Turner JMA, Mahadevaiah SK, Ellis PJI et al (2006) Pachytene asynapsis drives meiotic sex chromosome inactivation and leads to substantial postmeiotic repression in spermatids. Dev Cell 10:521–529. https://doi.org/10.1016/j.devcel.2006.02.009
Wright AE, Darolti I, Bloch NI et al (2017) Convergent recombination suppression suggests role of sexual selection in guppy sex chromosome formation. Nat Commun 8:14251. https://doi.org/10.1038/ncomms14251
Zhou Q, Zhang J, Bachtrog D et al (2014) Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 346:1246338. https://doi.org/10.1126/science.1246338
Zickler D, Kleckner N (1999) Meiotic chromosomes: integrating structure and function. Annu Rev Genet 33:603–754. https://doi.org/10.1146/annurev.genet.33.1.603
Acknowledgements
The authors thank Shaugnessy McCann, Jackie Cory, Jim Cory, Damean McCann, and Bill Frothingham for assistance collecting threespine stickleback fish. Kelly Dawe, Beth Dumont, and Kyle Swentowsky provided assistance in troubleshooting and useful discussions of our results. Brittany Dorsey helped count RAD51 foci.
Funding
This work was supported by the National Science Foundation MCB 1943283 to M.A.W.
Author information
Authors and Affiliations
Contributions
M.A.W. and S.N. conceived and designed the study. S.N., L.A.W., and M.K.F. performed the research. S.N. and M.A.W. analyzed the data. S.N. and M.A.W. wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Responsible Editor: Beth Sullivan.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Nath, S., Welch, L.A., Flanagan, M.K. et al. Meiotic pairing and double-strand break formation along the heteromorphic threespine stickleback sex chromosomes. Chromosome Res 30, 429–442 (2022). https://doi.org/10.1007/s10577-022-09699-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-022-09699-0