Abstract
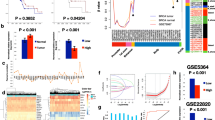
Deletions of chromosome 1p36 are common in malignancies; however, there is limited information regarding the biological and prognostic implications of 1p36 in cancer. Steroid Receptor–Associated and Regulated Protein (SRARP) is a tumor suppressor on chromosome 1p36.13 that its inactivation predicts poor cancer outcome, indicating that the 1p36.13 segment requires further studies. Therefore, a comprehensive multi-omics analysis of The Cancer Genome Atlas (TCGA), the Pan-Cancer Analysis of Whole Genomes (PCAWD), the International Cancer Genome Consortium (ICGC), and the Genomic Data Commons (GDC) Pan-Cancer datasets was conducted to investigate the prognostic implications of 1p36.13 in malignancies. This study revealed that expression and DNA methylation of multiple genes on 1p36.13 are significantly associated with survival in primary tumors and normal adjacent tissues. In addition, copy-number loss in every gene on 1p36.13 predicts poor cancer outcome. Importantly, copy-number loss and somatic mutations of chromosome 1p36.13 segment are associated with worse survival in primary tumors, and DNA hypermethylation of 1p36.13 predicts poor outcome in normal adjacent tissues. Therefore, genomic and epigenetic aberrations of chromosome 1p36.13 have promising prognostic implications in cancer.











Similar content being viewed by others
Abbreviations
- SRARP:
-
Steroid Receptor–Associated and Regulated Protein
- AR:
-
Androgen receptor
- Mb:
-
Megabases
- TCGA:
-
The Cancer Genome Atlas
- RNAseq:
-
RNA sequencing
- RSEM:
-
RNA Seq by Expectation Maximization
- PCAWG:
-
The Pan-Cancer Analysis of Whole Genomes
- GDC:
-
The Genomic Data Commons
- FPKM-UQ:
-
Fragments per kilobase of transcript per million mapped reads upper quartile
- ICGC:
-
International Cancer Genome Consortium
- CNV:
-
Copy-number variation
- FDR:
-
False discovery rate
- DNA-VAF:
-
DNA-variant allele frequency.
References
Anders S, Pyl PT, Huber W (2015) HTSeq-A Python framework to work with high-throughput sequencing data. Bioinformatics 31:166–169. https://doi.org/10.1093/bioinformatics/btu638
Andries V, Vandepoele K, Staes K, Berx G, Bogaert P, van Isterdael G, Ginneberge D, Parthoens E, Vandenbussche J, Gevaert K, van Roy F (2015) NBPF1, a tumor suppressor candidate in neuroblastoma, exerts growth inhibitory effects by inducing a G1 cell cycle arrest. BMC Cancer 15:391. https://doi.org/10.1186/s12885-015-1408-5
Bagchi A, Mills AA (2008) The quest for the 1p36 tumor suppressor. Cancer Res 68:2551–2556. https://doi.org/10.1158/0008-5472.CAN-07-2095
Baylin SB (2005) DNA methylation and gene silencing in cancer. Nat Clin Pract Oncol 2:S4–S11. https://doi.org/10.1038/ncponc0354
Brodeur GM, Sekhon G, Goldstein MN (1977) Chromosomal aberrations in human neuroblastomas. Cancer 40:2256–2263. https://doi.org/10.1002/1097-0142(197711)40:5<2256::aid-cncr2820400536>3.0.co;2-1
Calabrese C, Davidson NR, Demircioğlu D et al (2020) Genomic basis for RNA alterations in cancer. Nature 578:129–136. https://doi.org/10.1038/s41586-020-1970-0
Chen Z, Wu W, Huang Y, Xie L, Li Y, Chen H, Li W, Yin D, Hu K (2019) RCC2 promotes breast cancer progression through regulation of Wnt signaling and inducing EMT. J Cancer 10:6837–6847. https://doi.org/10.7150/jca.36430
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES, Getz G (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31:213–219. https://doi.org/10.1038/nbt.2514
Esteller M (2007) Epigenetic gene silencing in cancer: the DNA hypermethylome. Hum Mol Genet 16:R50–R59. https://doi.org/10.1093/hmg/ddm018
Fabregat A, Jupe S, Matthews L, Sidiropoulos K, Gillespie M, Garapati P, Haw R, Jassal B, Korninger F, May B, Milacic M, Roca CD, Rothfels K, Sevilla C, Shamovsky V, Shorser S, Varusai T, Viteri G, Weiser J, Wu G, Stein L, Hermjakob H, D’Eustachio P (2018) The Reactome pathway knowledgebase. Nucleic Acids Res 48:D498–D503. https://doi.org/10.1093/nar/gkx1132
Goldman M, Craft B, Hastie M et al (2019) The UCSC Xena platform for public and private cancer genomics data visualization and interpretation. bioRxiv. https://doi.org/10.1101/326470
Grossman RL, Heath AP, Ferretti V, Varmus HE, Lowy DR, Kibbe WA, Staudt LM (2016) Toward a shared vision for cancer genomic data. N Engl J Med 375:1109–1112. https://doi.org/10.1056/NEJMp1607591
He HL, Lee YE, Shiue YL, Lee SW, Lin LC, Chen TJ, Wu TF, Li CF (2015) PLA2G2A overexpression is associated with poor therapeutic response and inferior outcome in rectal cancer patients receiving neoadjuvant concurrent chemoradiotherapy. Histopathology 66:991–1002. https://doi.org/10.1111/his.12613
Henrich KO, Schwab M, Westermann F (2012) 1p36 tumor suppression - a matter of dosage? Cancer Res 72:6079–6088. https://doi.org/10.1158/0008-5472.CAN-12-2230
Jensen MA, Ferretti V, Grossman RL, Staudt LM (2017) The NCI Genomic Data Commons as an engine for precision medicine. Blood 130:453–459. https://doi.org/10.1182/blood-2017-03-735654
Knuutila S, Aalto Y, Autio K, Björkqvist AM, el-Rifai W’, Hemmer S, Huhta T, Kettunen E, Kiuru-Kuhlefelt S, Larramendy ML, Lushnikova T, Monni O, Pere H, Tapper J, Tarkkanen M, Varis A, Wasenius VM, Wolf M, Zhu Y (1999) DNA copy number losses in human neoplasms. Am J Pathol 155:683–691. https://doi.org/10.1016/S0002-9440(10)65166-8
Légaré S, Cavallone L, Mamo A et al (2015) The estrogen receptor cofactor SPEN functions as a tumor suppressor and candidate biomarker of drug responsiveness in hormone-dependent breast cancers. Cancer Res 75:4351–4363. https://doi.org/10.1158/0008-5472.CAN-14-3475
Légaré S, Chabot C, Basik M (2017) SPEN, a new player in primary cilia formation and cell migration in breast cancer. Breast Cancer Res 104:104. https://doi.org/10.1186/s13058-017-0897-3
Li B, Zhu J, Meng L (2019) High expression of ACTL8 is poor prognosis and accelerates cell progression in head and neck squamous cell carcinoma. Mol Med Rep 19:877–884. https://doi.org/10.3892/mmr.2018.9716
Lin J, Deng Z, Tanikawa C et al (2014) Downregulation of the tumor suppressor HSPB7, involved in the p53 pathway, in renal cell carcinoma by hypermethylation. Int J Oncol 44:1490–1498. https://doi.org/10.3892/ijo.2014.2314
Martini G, Cardone C, Vitiello PP, Belli V, Napolitano S, Troiani T, Ciardiello D, Della Corte CM, Morgillo F, Matrone N, Sforza V, Papaccio G, Desiderio V, Paul MC, Moreno-Viedma V, Normanno N, Rachiglio AM, Tirino V, Maiello E, Latiano TP, Rizzi D, Signoriello G, Sibilia M, Ciardiello F, Martinelli E (2019) EphA2 is a predictive biomarker of resistance and a potential therapeutic target for improving antiepidermal growth factor receptor therapy in colorectal cancer. Mol Cancer Ther 18:845–855. https://doi.org/10.1158/1535-7163.MCT-18-0539
Mayrhofer M, Kultima HG, Birgisson H, Sundström M, Mathot L, Edlund K, Viklund B, Sjöblom T, Botling J, Micke P, Påhlman L, Glimelius B, Isaksson A (2014) 1p36 deletion is a marker for tumour dissemination in microsatellite stable stage II-III colon cancer. BMC Cancer 14:872. https://doi.org/10.1186/1471-2407-14-872
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12:R41. https://doi.org/10.1186/gb-2011-12-4-r41
Moore LD, Le T, Fan G (2013) DNA methylation and its basic function. Neuropsychopharmacology 38:23–38. https://doi.org/10.1038/npp.2012.112
Naderi A (2017) C1orf64 is a novel androgen receptor target gene and coregulator that interacts with 14-3-3 protein in breast cancer. Oncotarget 8:57907–57933. https://doi.org/10.18632/oncotarget.17826
Naderi A (2018) SRARP and HSPB7 are epigenetically regulated gene pairs that function as tumor suppressors and predict clinical outcome in malignancies. Mol Oncol 12:724–755. https://doi.org/10.1002/1878-0261.12195
Shahriyari L (2017) Effect of normalization methods on the performance of supervised learning algorithms applied to HTSeq-FPKM-UQ data sets: 7SK RNA expression as a predictor of survival in patients with colon adenocarcinoma. Brief Bioinform 20:985–994. https://doi.org/10.1093/bib/bbx153
Wang JK, Wang WJ, Cai HY, du BB, Mai P, Zhang LJ, Ma W, Hu YG, Feng SF, Miao GY (2018) MFAP2 promotes epithelial–mesenchymal transition in gastric cancer cells by activating TGF-β/SMAD2/3 signaling pathway. Onco Targets Ther 11:4001–4017. https://doi.org/10.2147/OTT.S160831
Weinstein JN, Akbani R, Broom BM et al (2014) Comprehensive molecular characterization of urothelial bladder carcinoma. Nature 507:315–322. https://doi.org/10.1038/nature12965
Wu WKK, Li X, Wang X, Dai RZW, Cheng ASL, Wang MHT, Kwong T, Chow TC, Yu J, Chan MTV, Wong SH (2017) Oncogenes without a neighboring tumor-suppressor gene are more prone to amplification. Mol Biol Evol 34:903–907. https://doi.org/10.1093/molbev/msw295
Yates AD, Achuthan P, Akanni W, Allen J, Allen J, Alvarez-Jarreta J, Amode MR, Armean IM, Azov AG, Bennett R, Bhai J, Billis K, Boddu S, Marugán JC, Cummins C, Davidson C, Dodiya K, Fatima R, Gall A, Giron CG, Gil L, Grego T, Haggerty L, Haskell E, Hourlier T, Izuogu OG, Janacek SH, Juettemann T, Kay M, Lavidas I, le T, Lemos D, Martinez JG, Maurel T, McDowall M, McMahon A, Mohanan S, Moore B, Nuhn M, Oheh DN, Parker A, Parton A, Patricio M, Sakthivel MP, Abdul Salam AI, Schmitt BM, Schuilenburg H, Sheppard D, Sycheva M, Szuba M, Taylor K, Thormann A, Threadgold G, Vullo A, Walts B, Winterbottom A, Zadissa A, Chakiachvili M, Flint B, Frankish A, Hunt SE, IIsley G, Kostadima M, Langridge N, Loveland JE, Martin FJ, Morales J, Mudge JM, Muffato M, Perry E, Ruffier M, Trevanion SJ, Cunningham F, Howe KL, Zerbino DR, Flicek P (2020) Ensembl 2020. Nucleic Acids Res 48:D682–D688. https://doi.org/10.1093/nar/gkz966
Yuan L, Lv Y, Li H, Gao H, Song S, Zhang Y, Xing G, Kong X, Wang L, Li Y, Zhou T, Gao D, Xiao ZX, Yin Y, Wei W, He F, Zhang L (2015) Deubiquitylase OTUD3 regulates PTEN stability and suppresses tumorigenesis. Nat Cell Biol 17:1169–1181. https://doi.org/10.1038/ncb3218
Zack TI, Schumacher SE, Carter SL, Cherniack AD, Saksena G, Tabak B, Lawrence MS, Zhang CZ, Wala J, Mermel CH, Sougnez C, Gabriel SB, Hernandez B, Shen H, Laird PW, Getz G, Meyerson M, Beroukhim R (2013) Pan-cancer patterns of somatic copy number alteration. Nat Genet 45:1134–1140. https://doi.org/10.1038/ng.2760
Zhang J, Baran J, Cros A, et al (2011) International cancer genome consortium data portal-a one-stop shop for cancer genomics data. Database bar026. doi: https://doi.org/10.1093/database/bar026
Author information
Authors and Affiliations
Contributions
AN conceived the study, carried out the bioinformatics and biostatistics analyses of datasets, interpreted the analyzed data, and drafted the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author reports no conflict of interest.
Ethics approval
This article does not contain any studies with human participants or animals performed by the author. All datasets used for this study are publicly available and anonymized.
Additional information
Responsible Editor: Matthew Breen
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Naderi, A. Genomic and epigenetic aberrations of chromosome 1p36.13 have prognostic implications in malignancies. Chromosome Res 28, 307–330 (2020). https://doi.org/10.1007/s10577-020-09638-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10577-020-09638-x