Abstract
Periodic planewave and molecular cluster density functional theory (DFT) calculations were performed on three potential arrangements of 18 chain chain cellulose microfibrils (CMFs). To determine the most probable arrangement in plant cell walls, the molecular structure, 13C NMR chemical shifts and WAXS diffractograms resulting from the DFT model calculations were compared to experimental data. In addition, the relative potential energies of the 18-chain model CMFs were considered as evidence for the most likely arrangement. The preponderance of evidence for the CMF arrangement that is most probable in plant cell walls is a 6-layer CMF in an arrangement of 234432 glucan chains where each integer represents the number of chains in a given layer. An accurate model for the habit of the CMF in plant cell walls is necessary for further modeling of CMF interactions with other plant cell wall components and studies of cellulose degradation.





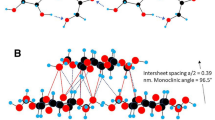
adapted from ref (Hill et al. 2014)). The 234432 arrangement is the only habit also consisting of a hexamer of trimers (b)
Similar content being viewed by others
References
Adamo C, Barone V (1998) Exchange functionals with improved long-range behavior and adiabatic connection methods without adjustable parameters: the mPW and mPW1PW models. J Chem Phys 108:664–675
Agarwal UP, Ralph SA, Reiner RS, Baez C (2016) Probing crystallinity of never-dried wood cellulose with Raman spectroscopy. Cellulose 23:125–144
Banks JL et al (2005) Integrated modeling program, applied chemical theory (IMPACT). J Comput Chem 26:1752–1780
Becke AD (1993) A new mixing of Hartree–Fock and local density-functional theories. J Chem Phys 98:1372–1377
Besombes S, Mazeau K (2005) The cellulose/lignin assembly assessed by molecular modeling. Part 2: seeking for evidence of organization of lignin molecules at the interface with cellulose. Plant Physiol Biochem 43:277–286
Cancés E, Mennucci B, Tomasi J (1997) A new integral equation formalism for the polarizable continuum model: theoretical background and applications to isotropic and anisotropic dielectrics. J Chem Phys 107:3032–3041
Case DA, Babin V, Berryman JT, Betz RM, Cai Q, Cerutti DS, Cheatham TE, III, Darden TA, Duke RE, Gohlke H, Goetz AW, Gusarov S, Homeyer N, Janowski P, Kaus J, Kolossváry I, Kovalenko A, Lee TS, LeGrand S, Luchko T, Luo R, Madej B, Merz KM, Paesani F, Roe DR, Roitberg A, Sagui C, Salomon-Ferrer R, Seabra G, Simmerling CL, Smith W, Swails J, Walker RC, Wang J, Wolf RM, Wu X, Kollman PA (2014) AMBER 14. University of California, San Francisco
Chen P-C, Hub JS (2014) Validating solution ensembles from molecular dynamics simulation by wide-angle X-ray scattering data. Biophys J 107:435–447
Cosgrove DJ (2018) Nanoscale structure, mechanics and growth of epidermal cell walls. Curr Opin Plant Biol 46:77–86
Crawshaw J, Bras W, Mant G, Cameron R (2002) Simultaneous SAXS and WAXS investigations of changes in native cellulose fiber microstructure on swelling in aqueous sodium hydroxide. J Appl Polym Sci 83:1209–1218
Dick-Pérez M, Zhang Y, Hayes J, Salazar A, Zabotina OA, Hong M (2011) Structure and interactions of plant cell-wall polysaccharides by two-and three-dimensional magic-angle-spinning solid-state NMR. Biochemistry 50:989–1000
Dri FL, Hector LG Jr, Moon RJ, Zavattieri PD (2013) Anisotropy of the elastic properties of crystalline cellulose Iß from first principles density functional theory with Van der Waals interactions. Cellulose 20:2703–2718. https://doi.org/10.1007/s10570-013-00718
Fernandes AN et al (2011) Nanostructure of cellulose microfibrils in spruce wood. Proc Natl Acad Sci 108:E1195–E1203
Frisch MJ et al (2009) Gaussian 09, Revision D. 01. Gaussian, Inc, Wallingford
Gottlieb HE, Kotlyar V, Nudelman A (1997) NMR chemical shifts of common laboratory solvents as trace impurities. J Org Chem 62:7512–7515
Grimme S, Ehrlich S, Goerigk L (2011) Effect of the damping function in dispersion corrected density functional theory. J Comput Chem 32:1456–1465
Gubitosi M, Nosrati P, Hamid MK, Kuczera S, Behrens MA, Johansson EG, Olsson U (2017) Stable, metastable and unstable cellulose solutions. Royal Soc Open Sci 4:170487
Guerriero G, Fugelstad J, Bulone V (2010) What do we really know about cellulose biosynthesis in higher plants? J Integr Plant Biol 52:161–175
Haigler CH, Roberts AW (2019) Structure/function relationships in the rosette cellulose synthesis complex illuminated by an evolutionary perspective. Cellulose 26:227–247
Hayashi T (1989) Xyloglucans in the primary cell wall. Annu Rev Plant Biol 40:139–168
Hill JL, Hammudi MB, Tien M (2014) The Arabidopsis cellulose synthase complex: a proposed hexamer of CESA trimers in an equimolar stoichiometry. Plant Cell 26:4834–4842
Horii F, Hirai A, Kitamaru R (1983) Solid-state 13C-NMR study of conformations of oligosaccharides and cellulose. Polym Bull 10:357–361
Jarvis MC (2013) Cellulose biosynthesis: counting the chains. Plant Physiol 163:1485–1486
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Kimura S, Laosinchai W, Itoh T, Cui X, Linder CR, Brown RM (1999) Immunogold labeling of rosette terminal cellulose-synthesizing complexes in the vascular plant Vigna angularis. Plant Cell 11:2075–2085
Kirschner KN, Yongye AB, Tschampel SM, González-Outeiriño J, Daniels CR, Foley BL, Woods RJ (2008) GLYCAM06: a generalizable biomolecular force field. Carbohydr J Comput Chem 29:622–655
Klimeš J, Bowler DR, Michaelides A (2009) Chemical accuracy for the van der Waals density functional. J Phys Condens Matter 22:022201
Klimeš J, Bowler DR, Michaelides A (2011) Van der Waals density functionals applied to solids. Phys Rev B 83:195131
Knight CJ, Hub JS (2015) WAXSiS: a web server for the calculation of SAXS/WAXS curves based on explicit-solvent molecular dynamics. Nucleic Acids Res 43:W225–W230
Kono H, Numata Y (2006) Structural investigation of cellulose Iα and Iβ by 2D RFDR NMR spectroscopy: determination of sequence of magnetically inequivalent d-glucose units along cellulose chain. Cellulose 13:317–326
Kono H, Erata T, Takai M (2003) Determination of the through-bond carbon–carbon and carbon–proton connectivities of the native celluloses in the solid state. Macromolecules 36:5131–5138
Kresse G, Furthmüller J (1996) Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys Rev B 54:11169
Kresse G, Hafner J (1993) Ab initio molecular dynamics for open-shell transition metals. Phys Rev B 48:13115
Kresse G, Hafner J (1994) Ab initio molecular-dynamics simulation of the liquid-metal–amorphous-semiconductor transition in germanium. Phys Rev B 49:14251
Kresse G, Furthmüller J, Hafner J (1994) Theory of the crystal structures of selenium and tellurium: the effect of generalized-gradient corrections to the local-density approximation. Phys Rev B 50:13181
Kubicki JD, Mohamed MN-A, Watts HD (2013) Quantum mechanical modeling of the structures, energetics and spectral properties of Iα and Iβ cellulose. Cellulose 20:9–23
Kubicki JD, Yang H, Sawada D, O’Neill H, Oehme D, Cosgrove D (2018) The shape of native plant cellulose microfibrils. Sci Rep 8:13983
Lee C, Yang W, Parr RG (1988) Development of the Colle–Salvetti correlation-energy formula into a functional of the electron density. Phys Rev B 37:785
Makarem M, Lee CM, Kafle K, Huang S, Chae I, Yang H, Kubicki JD, Kim SH (2019) Probing cellulose structures with vibrational spectroscopy. Cellulose 26:35–79. https://doi.org/10.1007/s10570-018-2199-z
Nishiyama Y, Langan P, Chanzy H (2002) Crystal structure and hydrogen-bonding system in cellulose Iβ from synchrotron X-ray and neutron fiber diffraction. J Am Chem Soc 124:9074–9082
Nishiyama Y, Johnson GP, French AD, Forsyth VT, Langan P (2008) Neutron crystallography, molecular dynamics, and quantum mechanics studies of the nature of hydrogen bonding in cellulose Iβ. Biomacromol 9:3133–3140
Nixon BT et al (2016) Comparative structural and computational analysis supports eighteen cellulose synthases in the plant cellulose synthesis complex. Sci Rep 6:28696
Oehme DP, Yang H, Kubicki JD (2018) An evaluation of the structures of cellulose generated by the CHARMM force field: Comparisons to in planta cellulose. Cellulose 25:3755–3777
Perdew JP, Wang Y (1992) Accurate and simple analytic representation of the electron-gas correlation energy. Phys Rev B 45:13244
Pereira CS, Silveira RL, Dupree P, Skaf MS (2017) Effects of xylan side-chain substitutions on xylan–cellulose interactions and implications for thermal pretreatment of cellulosic biomass. Biomacromol 18:1311–1321
Phyo P, Wang T, Yang Y, O’Neill H, Hong M (2018) Direct determination of hydroxymethyl conformations of plant cell wall cellulose using 1H polarization transfer solid-state NMR. Biomacromol 19:1485–1497
Rassolov VA, Ratner MA, Pople JA, Redfern PC, Curtiss LA (2001) 6–31G* basis set for third-row atoms. J Comput Chem 22:976–984
Sarotti AM, Pellegrinet SC (2009) A multi-standard approach for GIAO 13C NMR calculations. J Org Chem 74:7254–7260
Schrödinger L (2014) Schrödinger release 2014-1: Maestro, version 9.7. New York, NY
Shklyaev OE, Kubicki JD, Watts HD, Crespi VH (2014) Constraints on Iβ cellulose twist from DFT calculations of 13C NMR chemical shifts. Cellulose 21:3979–3991
Simmons TJ et al (2016) Folding of xylan onto cellulose fibrils in plant cell walls revealed by solid-state NMR. Nat Commun 7:13902
Suzuki S, Horii F, Kurosu H (2009) Theoretical investigations of 13C chemical shifts in glucose, cellobiose, and native cellulose by quantum chemistry calculations. J Mol Struct 921:219–226
Thomas LH, Forsyth VT, Martel A, Grillo I, Altaner CM, Jarvis MC (2015) Diffraction evidence for the structure of cellulose microfibrils in bamboo, a model for grass and cereal celluloses. BMC Plant Biol 15:153
Vandavasi VG et al (2016) A structural study of CESA1 catalytic domain of Arabidopsis cellulose synthesis complex: evidence for CESA trimers. Plant Physiol 170:123–135
Vietor RJ, Newman RH, Ha MA, Apperley DC, Jarvis MC (2002) Conformational features of crystal-surface cellulose from higher plants. Plant J 30:721–731
Wang T, Zabotina O, Hong M (2012) Pectin–cellulose interactions in the Arabidopsis primary cell wall from two-dimensional magic-angle-spinning solid-state nuclear magnetic resonance. Biochemistry 51:9846–9856
Wang T, Yang H, Kubicki JD, Hong M (2016) Cellulose structural polymorphism in plant primary cell walls investigated by high-field 2D solid-state NMR spectroscopy and density functional theory calculations. Biomacromol 17:2210–2222
Watts HD, Mohamed MNA, Kubicki JD (2011) Comparison of multistandard and TMS-standard calculated NMR shifts for coniferyl alcohol and application of the multistandard method to lignin dimers. J Phys Chem B 115:1958–1970
Watts HD, Mohamed MNA, Kubicki JD (2014) A DFT study of vibrational frequencies and 13C NMR chemical shifts of model cellulosic fragments as a function of size. Cellulose 21:53–70
Yang H, Zimmer J, Yingling YG, Kubicki JD (2015) How cellulose elongates—a QM/MM study of the molecular mechanism of cellulose chainization in bacterial CESA. J Phys Chem B 119:6525–6535
Yang H, Wang T, Oehme D, Petridis L, Hong M, Kubicki JD (2018) Structural factors affecting 13C NMR chemical shifts of cellulose: a computational study. Cellulose 25:23–36
Yang H, McManus JB, Oehme D, Singh A, Yingling YG, Tien M, Kubicki JD (2019) Simulations of cellulose synthesis initiation and termination in bacteria. J Phys Chem B 123:3699–3705
Zhao Y, Schultz NE, Truhlar DG (2006) Design of density functionals by combining the method of constraint satisfaction with parametrization for thermochemistry, thermochemical kinetics, and noncovalent interactions. J Chem Theory Comput 2:364–382
Zhao Z, Crespi VH, Kubicki JD, Cosgrove DJ, Zhong L (2014) Molecular dynamics simulation study of xyloglucan adsorption on cellulose surfaces: effects of surface hydrophobicity and side-chain variation. Cellulose 21:1025–1039
Acknowledgments
This work was supported by the Center for Lignocellulose Structure and Formation, an Energy Frontier Research Center funded by the U.S. Department of Energy, Office of Science, Basic Energy Sciences under Award # DE-SC0001090. Portions of this research were conducted with Advanced Cyber Infrastructure computational resources provided by the Institute for Cyber Science at The Pennsylvania State University (http://ics.psu.edu). This research also used resources of NERSC, supported by the Office of Science of DOE under Contract No. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, H., Kubicki, J.D. A density functional theory study on the shape of the primary cellulose microfibril in plants: effects of C6 exocyclic group conformation and H-bonding. Cellulose 27, 2389–2402 (2020). https://doi.org/10.1007/s10570-020-02970-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10570-020-02970-9