Abstract
By employing Plackett–Burman design (PBD) to investigate the qualitative interactions between eleven parameters [potato shell weight, incubation time, glucose, lactose, baker's yeast, peptone, (NH4)2SO4, CuSO4, FeSO4, CaCl2, KCl], and central composite design (CCD), the production of α-amylase by a local bacterial isolate Bacillus spp. NRC1 using agro-industrial wastes was improved. Using a variety of waste materials, including corn cobs (CC), onion peels (OP), rice straw (RS), potato shells (PS), Molokhia stem (MS), okra suppression (OS), lemon peels (LP), and pea peels (PP), Bacillus spp. NRC1 was tested in the production of amylase. Out of the eight agro-industrial wastes tried, the highest amylase yield (6.99 U/ml) was reached using potato peels. The statistical optimization of enzyme production was carried out using PBD followed by CCD design, causing 2.06-fold and 1.51-fold increase, respectively. The overall increase was 3.11-fold. In PBD design, potato peels, peptone and (NH4)2SO4 were positive factors for amylase production. The crude enzyme was tested for its ability to desize cotton fabric and subsequently studying its effect on fabric dyeability. The produced amylase proved its potentials in textile industry.
Graphical Abstract

Similar content being viewed by others
1 Introduction
Enzymes are biocatalysts that have been exploited industrially since decades. They have the superiority of being highly specific, ecofriendly and economic thus decreasing the problems associated with dependence on chemical catalysts in industry [1].
Amylases are commercially crucial enzymes as they are contributing 25–30% of worldwide market of catalyst [2]. Alpha-amylase (E.C. 3.2.1.1) plays a crucial part in the conversion of starch into low molecular weight sugars. Also, it is applied in removing environmental pollutant, bakery, fermentation, animal feed, detergent, paper, and desizing of textiles industries [3,4,5].
Amylase can be found in a variety of sources, including plants, animals, and microorganisms; however, only microbial amylases are suitable for use in industry. Microbial amylase are either obtained from fungi [6, 7] or bacteria [1, 5, 8, 9]. Agro-industrial waste is a valuable resource for enzyme and valuable products production [5]. Potato peels are one of worthwhile agro-industrial wastes since a typical potato processing facility can lose between 21 and 25% of the potatoes from peeling, trimming, and cutting. Potato peels are a suitable choice for fermentation to yield amylase because they are rich in numerous reusable chemicals. Among its various nutrients are proteins (18%), carbohydrates (55.0%) which contain starch (25.0%) and non-starch polysaccharides like xylans, cellulose, and glucan (30%), fats (1.0%), ash (6.0%), minerals (Ca 0.12%, Mg 0.23%, P 0.57%, K 0.42%, Na 0.06%), and vitamins (0.021 mg of thiamine, 0.038 mg of riboflavin, 1.033 mg of niacin, and 11.4 mg of ascorbic acid) [4, 10].
Medium optimization is a method to enhance enzyme production. One factor at a time is the traditional optimization method which has the disadvantage of being time consuming and ignoring the interactions between variable factors. However, statistical optimization methods overcome these drawbacks as they are able to screen larger number of significant factors, evaluate the interaction between parameters and finally select the optimum conditions to maximize enzyme production [4, 11].
In textile industry, a sizing agent is added to yarn prior to fabric production. This sizing agent protects the yarn against breaking during weaving. Starch is considered a perfect sizing agent as it is cheap, available and easily removed in textile finishing industry. Desizing process involves the removal of starch from fabric which is carried out either chemically or enzymatically. Amylase desizing has the leverage of selectivity, speed, specificity and ecofriendly [12, 13]. Enzymatic biotreatment of cotton fabrics enhances the physico-chemical properties of the surface and introduces functional groups on the fibers’ surface. Thus, it boosts the affinity of fabric dying [14, 15].
This study targeted microbial amylase production through fermentation of agro industrial residues by a local bacterial isolate. Two steps statistical design [Plackett–Burman design (PBD) followed by Central Composite Design (CCD)] was employed to maximize enzyme production. The resulted crude enzyme was tested for its cotton fabric desizing ability and the dying and pigmentation of biotreated fabrics to evaluate enzyme applicability in textile industry.
2 Materials and Methods
2.1 Raw Material
Agro- industrial wastes [corn cobs (CC), onion peel (OP), rice straw (RS), potato shells (PS), Molokhia stem (MS), okra suppression (OS), lemon peels (LP), and pea peels (PP)] were collected from Egyptian local market. They were cleaned and dried for 24 h at 70 °C in an oven. In order to use the dried materials as the substrate in submerged fermentation (SMF), they were powdered, sieved into coarse particles (1 cm), and then stored in airtight containers.
2.2 Strain Isolation and Identification
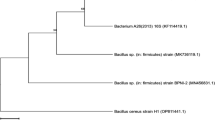
Bacillus spp. NRC1 was isolated from potato peels purchased from local Egyptian market. Two grams of potato peels were added to 50 ml sterile distilled water in 250 Erlenmeyer flask, shacked for 1 h then 200 µl was spread on nutrient agar plates. The plates were incubated for 24 h at 37 °C. Single colonies were picked and streaked on nutrient agar plates to check culture purity and incubated again (24 h, 37° C). The pure cultures were transformed to nutrient agar slants for further tests. Molecular identification based on the 16S-rRNA sequence of the selected strains was conducted. The nucleotide sequence of the isolated bacterium was submitted to the GenBank and assigned accession number MN396623 and the phylogenetic tree of Bacillus spp. NRC1 was shown in Supplementary Fig. 1.
2.3 Production of α-Amylase Enzyme Under Submerged Fermentation (SMF)
Two grams of each abovementioned eight dried substrates were added to 50 ml distilled water in 250 ml Erlenmeyer flasks, autoclaved (121 °C, 20 min) and cooled. Each flask was inoculated with 2 ml of cell suspension of 24 h old slant (OD600∼0.4). The flasks were incubated in shacking incubator (150 rpm, 37 °C) for 2 days.
2.4 α-Amylase Assay
According to Sajjad and Choudhry [16], this was accomplished by mixing 0.5 ml of 1% starch solution produced in acetate buffer (0.05 M, pH 5.0) with 0.5 ml of clear culture filtrate (crude enzyme). The combination was incubated for 20 min at 40 °C, and the Somogyi method [17] was used to calculate the amount of released reducing sugars. One unit of α-amylase (IU) was defined as the amount of enzyme which librated 1 µmol of reducing sugar per min under the assay conditions.
2.5 Optimization of Bacillus spp. NRC1 α-Amylase Production by Statistical Factorial Design
2.5.1 Plackett–Burman Design (PBD)
In this design, we looked into the qualitative impact of eleven parameters on the generation of α-amylase, such as A: potato shells weight (g/flask), B: incubation time (h), C: glucose (g/l), D: lactose (g/l), E: baker’s yeast (g/l), F: peptone (g/l), G: (NH4)2SO4 (g/l), H: CuSO4 (g/l), J: FeSO4, K: CaCl2 (g/l), L: KCl (g/l).
Low level (− 1) and high level (+ 1) studies were conducted on each parameter. Twelve experiments in total were conducted using the n + 1 rule, where n: is the total number of elements being studied. Each column in the experimental design corresponded to an independent variable, and each row to an experiment (Table 1). The multiple coefficient of determination, or R2, provides the percentage of variance explained by the model developed and the F-value provides the statistical significance (Table 2). Experimental responses were analyzed by first order model by the following equation: α-amylase activity (U/ml) = β° + ΣβiXi.
β° is the model intercept and βi is the linear coefficient, and Xi is the level of the independent variable.
2.5.2 Central Composite Design (CCD)
In this design, we examined the quantitative impact of the Plackett–Burman design's most effective variables, including (A) the weight of potato shells (g/flask), (B) peptone (g/l), and (C) (NH4)2SO4 (g/l).The five levels − 1.316, − 1, 0, + 1, and + 1.316 employed in CCD to study the variables resulted in a total of 20 trials as indicated in Table 3. The analysis of variance (ANOVA) was used in the statistical analysis of the model (Table 4).
2.6 Applications of Bacillus spp. NRC1 Culture Filtrate
2.6.1 Enzymatic Desizing and Half Bleached Treatment
In this experiment an aqueous desizing solution was applied to cotton fabric. It was composed of enzyme solution (30 ml l−1), non-ionic wetting agent (1 g l−1) and NaClO2 (5 g l−1) using a material to liquor ratio of 1:30 and the reaction was carried out in buffer solution. The treatment was carried out in an SK3310HP KUDOS ultrasonic water bath cleaner with a frequency of 45 kHz for 30 min at 60 °C, after that, the fabrics were padded, patched and put in oven at 60 °C for 30 min then washed twice with hot water, then with cold water and finally dried. Wettability was estimated as time required for water drop to be absorbed into the fabric. Desizing efficiency was estimated by the TEGEWA scale method and violet scale shade [18].
2.6.2 Coloration of the Biotreated Cotton Samples
2.6.2.1 Dyeing
A reactive red dye was used to dye treated cotton fabric according to Li et al. [18]. The dyeing was conducted with sodium sulfate (60 g l−1) at 40 °C for 30 min, then fixation for 60 min in sodium carbonate (20 g l−1) at 60 °C. The dyed samples were washed and dried at room temperature.
2.6.2.2 Pigment Printing Technique
Cotton fabric was printed using pigment printing technique. The printing paste was prepared according to Hebeish et al. [19]. It was composed of curcumin (3 g %), urea (2.5 g %), synthetic thickener (2.5 g %), binder (2.5 g %) and NaH2PO4 (0.5 g %) in distilled water. According to the guidelines set out in the test AATCC test method [20], the samples were washed. According to the AATCC test methods [21,22,23] the color fastness to rubbing, perspiration, and light was assessed.
3 Results and Discussion
3.1 Production of α-Amylase
Agricultural wastes are rich in carbon, nitrogen and minerals which make them a perfect target for microbial fermentation to produce enzymes [24]. Out of the eight substrates tried, potato peels gave the highest enzyme yield (6.99 U/ml). This result was similar to that reported by Mukherjee et al. [4] who used potato peels to produce α-amylase. Potato peels have the advantages of being rich in nutrients needed for microbial growth and enzyme production beside their availability from potato processing industries [4]. On the other hand, other agricultural wastes were used for enzyme production; olive oil cake [6], wheat bran [25], rice husk [26], moong husk, maize straw, sugarcane bagasse, wheat straw, orange and pomegranate [27].
3.2 Optimization of Bacillus spp. NRC1 α-Amylase Production
3.2.1 Plackett–Burman Design
We optimized and studied the qualitative effect of 11 factors as shown in Table 1 on α-amylase production. The highest α-amylase production was obtained in trail 8 (14.40 U/ml) causing 2.06-fold increase in production. Showing that the maximum production was obtained after 48 h of incubation similar to that reported by Issac and Prince [28] and Saad et al. [11].
The α-amylase activity (U/ml) can by calculated from the following equation:
The validation of the design is expressed by α-amylase production. Each medium component's impact on the production of α-amylase was described by the magnitude and direction of the factor coefficient. The coefficient's (−) and (+) symbols denote the tested factor's respective negative and positive effects on the production of α-amylase. The weight of the potato shells and the addition of additional nitrogen sources, such as peptone and (NH4)2SO4, had the highest positive effects on the production of α-amylase, as shown in Fig. 1, according to some authors [28,29,30,31,32,33,34] found that yeast extract was the most effective source of nitrogen. Contrarily, as reported by Monga et al. [29] and Ahmed et al. [3], the addition of additional carbon sources such as glucose and lactose had a negative impact on the production of α-amylase. In contrast, it was found by Irfan et al. [30], Saleem and Ebrahim [31], Issac and Prince [28] and Hasan et al. [33] that the addition of glucose and lactose increased the production of α-amylase. According to Ahmed et al. [3], incubation time, FeSO4, and CaCl2 had a detrimental influence on the production of α-amylase in Bacillus spp. NRC1. The results of the analysis of variance are displayed in Table 2 and showed the extent to which the design worked to produce α-amylase. The design's success was demonstrated by the R2 value of 0.9948, which showed that the model could account for 99.48% of the response's variability. The model is successful since the values of the predicted R2 of 0.9176 and the adjusted R2 of 0.9811 are closer to one another. For the production of α-amylase in this analysis, adequate precision of 28.326 indicated adequate signal. The value of the coefficient of variation (CV), which measures the accuracy with which experiments were carried out, was discovered to be 7.42. Low CV value indicates that the experiments were accurate and reliable [35].
3.2.2 Central Composite Design
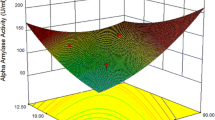
The quantitative effect and interaction between the most effective factors [PS weight, peptone, and (NH4)2SO4] on α-amylase production determined from PB design were shown by Contour plots (Fig. 2a–c). Also, here the validation of CCD is expressed by α-amylase production. According to Table 3, trial number 18 had the highest α-amylase production, 21.76 U/ml, which resulted in increases of 1.51 and 3.11 times when compared to the previous step of optimization and the unoptimized medium, respectively. This yield of α-amylase was greater than those that other writers had reported [28, 30, 36,37,38]. The α-amylase activity (U/ml) can by calculated from the following equation:
The relationship between the experimental and predicted values for observed and predicted α-amylase activity was displayed in Fig. 3, and the data points located close to the diagonal line, suggesting a satisfactory correlation. By measuring the coefficient R2, which has a value of 0.9904, the goodness of fit of the model was evaluated. The closer the R2 is to 1.0, the more robust the model is and the more accurately it predicts the generation of α-amylase. This outcome surpassed that of Singh et al. [5], which used CCD to produce α-amylase from Bacillus subtilis 1934 on potato and apple peel. The findings revealed that for potato and apple peel, respectively, R2 is 0.97 and 0.95.
The adjusted R2 value of 0.9860 and the predicted R2 value of 0.9443 were reasonably consistent. Model significance is indicated by the model's F-value of 224.00 (Table 4). The significance of the model terms is indicated by Prob > F values less than 0.0500. Low CV% score of 2.52 suggested experiment accuracy and reliability [35]. The effectiveness of this design for α-amylase production which indicated by mentioned R2, predicated R2 and adjusted R2 was higher than that reported by other authors [38, 39].
In a numerical optimization, the quadratic model predicted the maximum α-amylase activity of 21.76 U/ml by SMF, which can be achieved with optimal values of media components g/l: 1.00 baker’s yeast, 1.00 peptone, 5.00 (NH4)2SO4, 0.01 CuSO4, 0.01 CaCl2, 0.01 KCl and 4 g potato shells/flask, after 48 h of incubation.
3.3 Textile Applications
3.3.1 Enzymatic Desizing and Half Bleached Treatment
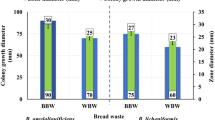
In textile industry, desizing step is important since it removes starch from fabrics to prepare it for dying. The use of ultrasonic increases the hydrophobic interactions between amylase and fabric, thus intensifying enzyme action leading to increased desizing efficacy. Moreover, the ultrasonic enhances loosened products removal from the fabric bulk [14, 40]. The results demonstrated in Table 5 revealed that the desizing activity increased remarkably by increasing enzyme concentration from 2 to 3%. The enzyme enhanced the bleaching activity of sodium chlorite which was confirmed by increased whiteness index from 41.83 to 48.66. Also, maximum weight loss (WL) was at 3% enzyme concentration (6.1%). In this context, our results were better than Hao et al. [40] who reported 3.5% weight loss at 60 °C and similar to Chand et al. [41] who used pure culture of Bacillus sp. KR-8104 for desizing resulting in TEGEWA scale 6. Moreover, Aggarwal et al. [42] applied different conditions to desize grey cotton fabric and the results showed TEGEWA scale range of 4–7. On the other hand, Chand et al. [12] reported weight loss of 14% and TEGEWA violet scale 7–8, while, Nair and Bhat [13] succeeded to desize starched white cotton cloth pieces using amylase from marine isolate.
3.3.2 Coloration of the Biotreated Cotton Samples
The effect of Bacillus spp. NRC1 α-amylase on the coloration of cotton fabric was evaluated by studying the fastness properties of dyed and printed samples in comparison to commercial enzyme. The data demonstrated in Table 6 revealed that washing fastness (WF) and rubbing fastness (Rub) of biotreated fabric ranged from fairly good to good (3–4). The perspiration fastness (Per) ranged from moderate to good for dyed fabric (2–4) and fairly good to good (3–4) for printed sample. On the other hand, light fastness (LF) ranged from good to very good (5–6). The light fastness of printed sample was better than that treated with the commercial enzyme. In this context, our results were similar to that showed by Vankar et al. [43] who used natural dye of Terminalia arjuna to color cotton fabric. They reported WF and Peracidic of 4 and Perbasic Rubdry Rubwet of 3–4, however, our results of LF were better. Enzyme treatment of cotton fabric enhanced dying; this can be attributed to surface modification leading to enhanced dye adsorption capacity. Moreover, Vankar and Shanker [44] studied the amylase on dying of cotton fabrics, their results for WF and LF were 4–5. Öner and Sahinbaskan [45] recorded WF and Rub of 4–5 for different synthetic dyes.
4 Conclusion
Enzyme production optimization through factorial designs is preferred as it studies factors interaction, lowering production cost and saving time. The present study revealed that industrially applicable α-amylase can be produced via safe, low-cost, and environment-green technologies via utilizing Bacillus spp. NRC1 as a potent amylase producer hydrolyzing different agricultural wastes, especially potato peels. By applying PBD and CCD designs, the enzyme yield increased to 2.06-fold and 3.11-fold compared to unoptimized medium, respectively. The enzyme was efficient in cotton desizing and coloration of fabric which makes it a potential candidate for applying in textile sector.
Data Availability
All data generated or analysed during this study are included in this published article.
References
Mondal S, Mondal K, Halder SK, Thakur N, Mondal KC (2022) Microbial Amylase: old but still at the forefront of all major industrial enzymes. Biocatal Agric Biotechnol. https://doi.org/10.1016/j.bcab.2022.102509
Sharif S, Shah AH, Fariq A, Jannat S, Rasheed S, Yasmin A (2023) Optimization of amylase production using response surface methodology from newly isolated thermophilic bacteria. Heliyon. https://doi.org/10.1016/j.heliyon.2023.e12901
Ahmed SA, Mostafa FA, Helmy WA, Abdel-Naby MA (2017) Improvement of bacterial α-amylase production and application using two steps statistical factorial design. Biocatal Agric Biotechnol 10:224–233. https://doi.org/10.1016/j.bcab.2017.03.004
Mukherjee R, Paul T, Soren JP, Halder SK, Mondal KC, Pati BR, Das Mohapatra PK (2019) Acidophilic α-amylase production from Aspergillus niger RBP7 using potato peel as substrate: a waste to value added approach. Waste Biomass Valoriz 10:851–863. https://doi.org/10.1007/s12649-017-0114-8
Singh R, Langyan S, Sangwan S, Gaur P, Khan FN, Yadava P, Rohatgi B, Shrivastava M, Khandelwal A, Darjee S, Sahu PK (2022) Optimization and production of alpha-amylase using Bacillus subtilis from apple peel: comparison with alternate feedstock. Food Biosci 49:101978. https://doi.org/10.1016/j.fbio.2022.101978
Karam EA, Wahab WAA, Saleh SAA, Hassan ME, Kansoh AL, Esawy MA (2017) Production, immobilization and thermodynamic studies of free and immobilized Aspergillus awamori amylase. Int J Biol Macromol 102:694–703. https://doi.org/10.1016/j.ijbiomac.2017.04.033
Mahmood S, Shahid MG, Irfan M, Nadeem M, Syed Q (2018) Partial characterization of α-amylase produced from Aspergillus niger using potato peel as substrate. Punjab Univ J Zool 33(1):22–27. https://doi.org/10.17582/pujz/2018.33.1.22.27
Priyadarshini S, Pradhan SK, Ray P (2020) Production, characterization and application of thermostable, alkaline α-amylase (AA11) from Bacillus cereus strain SP-CH11 isolated from Chilika Lake. Int J Biol Macromol 145:804–812. https://doi.org/10.1016/j.ijbiomac.2019.11.149
Abo-Kamer AM, Abd-El-Salam IS, Mostafa FA, Mustafa AERA, Al-Madboly LA (2023) A promising microbial α-amylase production, and purification from Bacillus cereus and its assessment as antibiofilm agent against Pseudomonas aeruginosa pathogen. Microb Cell Factories 22(1):141. https://doi.org/10.1186/s12934-023-02139-6
Joshi A, Sethi S, Arora B, Azizi AF, Thippeswamy B (2020) Potato peel composition and utilization. In: Raigond P, Singh B, Dutt S, Chakrabarti SK (eds) Potato. Springer, Singapore. https://doi.org/10.1007/978-981-15-7662-1_13
Saad WF, Othman AM, Abdel-Fattah M, Ahmad MS (2021) Response surface methodology as an approach for optimization of α-amylase production by the new isolated thermotolerant Bacillus licheniformis WF67 strain in submerged fermentation. Biocatal Agric Biotechnol 32:101944. https://doi.org/10.1016/j.bcab.2021.101944
Chand N, Nateri AS, Sajedi RH, Mahdavi A, Rassa M (2012) Enzymatic desizing of cotton fabric using a Ca2+-independent α-amylase with acidic pH profile. J Mol Catal B 83:46–50. https://doi.org/10.1016/j.molcatb.2012.07.003
Nair HP, Bhat SG (2020) Arabian Sea metagenome derived-α-amylase P109 and its potential applications. Ecol Genet Genomics 16:100060. https://doi.org/10.1016/j.egg.2020.100060
Zhang Y, Rather LJ, Li Q (2022) Recent advances in the surface modification strategies to improve functional finishing of cotton with natural colourants—a review. J Clean Prod 335:130313. https://doi.org/10.1016/j.jclepro.2021.130313
Abou Taleb M, Gomaa SK, Wahba MI, Zaki RA, El-Fiky AF, El-Refai HA, El-Sayed H (2022) Bioscouring of wool fibres using immobilized thermophilic lipase. Int J Biol Macromol 1(194):800–810. https://doi.org/10.1016/j.ijbiomac.2021.11.128
Sajjad M, Choudhry S (2012) Effect of starch containing organic substrates on alpha amylase production in Bacillus strains. Afr J Microbiol Res 6(45):7285–7291. https://doi.org/10.5897/AJMR12.181
Somogyi MJ (1952) Notes on sugars determinations. J Biol Chem 195:19–23. https://doi.org/10.1016/S0021-9258(19)50870-5
Li S, Boyter H, Stewart N (2004) Ultraviolet (UV) curing for textile coloration. AATCC Rev 4(8):44–49
Hebeish A, Ragheb AA, Rekaby M, El-Hennawi HM, Shahin AA (2019) Chitosan/disperse dye nanoparticles for concomitant printing and antibacterial finishing. Nanotechnol Russ 14(9):462–470. https://doi.org/10.1134/S1995078019050069
AATCC (1993) AATCC Test Method (36-1972) 68 (1993) Colorfastness to washing: characterization of textile colorants, technical manual method of the American Association of Textile Chemists and Colorists, USA
AATCC (1993) AATCC Test Method (8-1989) 68 (1993) Colorfastness to crocking: AATCC crock meter method, technical manual method of the American Association of Textile Chemists and Colorists, USA
AATCC (1993) AATCC Test Method (15-1989) 68 (1993) Colorfastness to perspiration, technical manual method of the American Association of Textile Chemists and Colorists, USA
AATCC (1993) AATCC Test Method (16A-1989) 68 (1993) Colorfastness to light: outdoor, technical manual method of the American Association of Textile Chemists and Colorists, USA
Mojumdar A, Deka J (2019) Recycling agro-industrial waste to produce amylase and characterizing amylase–gold nanoparticle composite. Int J Recycl Org Waste Agric 8:263–269. https://doi.org/10.1007/s40093-019-00298-4
Almanaa TN, Vijayaraghavan P, Alharbi NS, Kadaikunnan S, Khaled JM, Alyahya SA (2020) Solid state fermentation of amylase production from Bacillus subtilis D19 using agro-residues. J King Saud Univ Sci 32(2):1555–1561. https://doi.org/10.1016/j.jksus.2019.12.011
Rathi A, Gupta N, Dhruw V, Beliya E, Tiwari S, Paul JS, Jadhav SK (2022) Valorization of rice milled by-products (rice husk and de-oiled rice bran) into α-amylase with its process optimization, partial purification and kinetic study. Process Biochem 120:101–113. https://doi.org/10.1016/j.procbio.2022.06.006
Bhatt K, Lal S, Srinivasan R, Joshi B (2020) Bioconversion of agriculture wastes to produce α-amylase from Bacillus velezensis KB 2216: purification and characterization. Biocatal Agric Biotechnol 28:101703. https://doi.org/10.1016/jbcab2020101703
Issac R, Prince R (2015) Production of alpha-amylase by solid state fermentation using Bacillus cereus MTCC 7524 and Bacillus licheniformis MTCC 7445 from dairy sludge—a comparative study. Int J Pharmtech Res 8(9):111–117
Monga M, Goyal M, Kalra KL, Soni G (2011) Production and stabilization of amylases from Aspergillus niger. Mycosphere 2(2):129–134
Irfan M, Nadeem M, Syed Q (2012) Media optimization for amylase production in solid state fermentation of wheat bran by fungal strains. J Cell Mol Biol 10(1):55–64
Saleem A, Ebrahim MK (2014) Production of amylase by fungi isolated from legume seeds collected in Almadinah Almunawwarah. Saudi Arabia J Taibah Univ Sci 8(2):90–97. https://doi.org/10.1016/j.jtusci.2013.09.002
Akhter P, Mishra A, Mishra V, Raghav A (2017) Alpha amylase production from Aspergillus terreus UF39 using oil cakes. Eur J Biotechnol Biosci 5(6):15–21
Hasan MM, Marzan LW, Hosna A, Hakim A, Azad AK (2017) Optimization of some fermentation conditions for the production of extracellular amylases by using Chryseobacterium and Bacillus isolates from organic kitchen wastes. J Genet Eng Biotechnol 15(1):59–68. https://doi.org/10.1016/j.jgeb.2017.02.009
Ojha SK, Singh PK, Mishra S, Pattnaik R, Dixit S, Verma SK (2020) Response surface methodology based optimization and scale-up production of amylase from a novel bacterial strain, Bacillus aryabhattai KIIT BE-1. Biotechnol Rep 27:e00506. https://doi.org/10.1016/j.btre.2020.e00506
Saeed AM, El-Shatoury EH, Sayed HA (2021) Statistical factorial designs for optimum production of thermostable α-amylase by the degradative bacterium Parageobacillus thermoglucosidasius Pharon1 isolated from Sinai. Egypt J Genet Eng Biotechnol 19:1–9. https://doi.org/10.1186/s43141-021-00123-4
Vidyalakshmi R, Paranthaman R, Indhumathi J (2009) Amylase production on submerged fermentation by Bacillus spp. World J Chem 4(1):89–91
Raplong HH, Odeleye PO, Hammuel C, Idoko MO, Asanato JI, Odeke EH (2014) Production of alpha amylase by Bacillus cereus in submerged fermentation. Aceh Int J Sci Technol 3(3):124–130. https://doi.org/10.13170/aijst.3.3.1592
Tallapragada P, Dikshit R, Jadhav A, Sarah U (2017) Partial purification and characterization of amylase enzyme under solid state fermentation from Monascus sanguineus. J Genet Eng Biotechnol 15(1):95–101. https://doi.org/10.1016/j.jgeb.2017.02.003
Pereira WJ, Alves GL, Purcena LL, Bataus LAM, Fernandes KF, Batista KA (2017) Statistical optimization of α-amylase production by Escherichia coli using extruded bean as nitrogen and carbon source. Int J Environ Agric Biotechnol 2(4):238886. https://doi.org/10.22161/ijeab/2.4.74
Hao L, Wang R, Fang K, Liu J (2013) Ultrasonic effect on the desizing efficiency of α-amylase on starch-sized cotton fabrics. Carbohydr Polym 96(2):474–480. https://doi.org/10.1016/j.carbpol.2013.04.003
Chand N, Sajedi RH, Nateri AS, Khajeh K, Rassa M (2014) Fermentative desizing of cotton fabric using an α-amylase-producing Bacillus strain: optimization of simultaneous enzyme production and desizing. Process Biochem 49(11):1884–1888. https://doi.org/10.1016/j.procbio.2014.07.007
Aggarwal R, Dutta T, Sheikh J (2019) Extraction of amylase from the microorganism isolated from textile mill effluent vis a vis desizing of cotton. Sustain Chem Pharm 14:100178. https://doi.org/10.1016/j,scp.2019.100178
Vankar PS, Shanker R, Verma A (2007) Enzymatic natural dyeing of cotton and silk fabrics without metal mordants. J Clean Prod 15(15):1441–1450. https://doi.org/10.1016/j.jclepro.2006.05.004
Vankar PS, Shanker R (2008) Ecofriendly ultrasonic natural dyeing of cotton fabric with enzyme pretreatments. Desalination 230(1–3):62–69. https://doi.org/10.1016/j.desal.2007.11.016
Öner E, Sahinbaskan BY (2011) A new process of combined pretreatment and dyeing: REST. J Clean Prod 19(14):1668–1675. https://doi.org/10.1016/j.jclepro.2011.05.008
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was financially supported by the Research Projects Sector in the National Research Centre, Dokki, Giza, Egypt. The work was a part of an Internal Project (No. 11070202) entitled “Production of some microbial enzymes using agricultural residues and their applications in textile and food industries”.
Author information
Authors and Affiliations
Contributions
FAM: Conceptualization, Methodology, Performing the experiments, Software, Validation, Formal analysis, Investigation, Resources, Data, Writing original draft, Editing, Visualization. HRW: Performing the experiments, Resources, Data, Visualization. HME-H: Textile dying and printing, Data. SAM: Textile dying and printing, Data. SS: Textile enzymatic treatment, Data. SAAS: Conceptualization, Methodology, Performing the experiments, Validation, Investigation, Resources, Data, Writing and editing, Visualization.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mostafa, F.A., Wehaidy, H.R., El-Hennawi, H.M. et al. Statistical Optimization of α-Amylase Production from Novel Local Isolated Bacillus spp. NRC1 and Its Textile Applications. Catal Lett (2024). https://doi.org/10.1007/s10562-023-04545-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10562-023-04545-2