Abstract
Purpose
In a phase II trial in patients with metastatic triple-negative breast cancer (mTNBC; NCT02978716), administering trilaciclib prior to gemcitabine plus carboplatin (GCb) enhanced T-cell activation and improved overall survival versus GCb alone. The survival benefit was more pronounced in patients with higher immune-related gene expression. We assessed immune cell subsets and used molecular profiling to further elucidate effects on antitumor immunity.
Methods
Patients with mTNBC and ≤ 2 prior chemotherapy regimens for locally recurrent TNBC or mTNBC were randomized 1:1:1 to GCb on days 1 and 8, trilaciclib prior to GCb on days 1 and 8, or trilaciclib alone on days 1 and 8, and prior to GCb on days 2 and 9. Gene expression, immune cell populations, and Tumor Inflammation Signature (TIS) scores were assessed in baseline tumor samples, with flow cytometric analysis and intracellular and surface cytokine staining used to assess immune cell populations and function.
Results
After two cycles, the trilaciclib plus GCb group (n = 68) had fewer total T cells and significantly fewer CD8+ T cells and myeloid-derived suppressor cells compared with baseline, with enhanced T-cell effector function versus GCb alone. No significant differences were observed in patients who received GCb alone (n = 34). Of 58 patients in the trilaciclib plus GCb group with antitumor response data, 27 had an objective response. RNA sequencing revealed a trend toward higher baseline TIS scores among responders versus non‑responders.
Conclusion
The results suggest that administering trilaciclib prior to GCb may modulate the composition and response of immune cell subsets to TNBC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Triple-negative breast cancer (TNBC) is a specific subtype of breast cancer that is associated with high invasiveness, high metastatic potential, proneness to relapse, and poor prognosis. Patients with TNBC have fewer treatment options available to them than those with other types of invasive breast cancer [1,2,3,4]. Cytotoxic chemotherapy is the primary treatment option for patients with programmed death-ligand 1 (PD-L1)-negative disease [4, 5], whereas the combination of chemotherapy plus pembrolizumab is the preferred first-line treatment option for patients with PD-L1-positive tumors (the overall rate of PD-L1 positivity in TNBC ranges from 20 to 60%, depending on the assays and methods used) [6,7,8,9]. Other treatment options include poly(adenosine diphosphate-ribose) polymerase inhibitors for patients with BRCA-mutated tumors (reported prevalence rates varying from 10 to 20%) [10, 11], and sacituzumab govitecan, a Trop-2-directed antibody–drug conjugate, for the treatment of patients with unresectable locally advanced or metastatic TNBC (mTNBC) who have received two or more prior lines of systemic therapy [12]. Despite the emergence of these new therapies, many patients with locally advanced TNBC or mTNBC have no options other than standard chemotherapy, which is commonly associated with toxicities that can adversely impact quality of life [13].
Trilaciclib is an intravenous myeloprotection therapy that is administered as a 30-min infusion within 4 h prior to the start of chemotherapy on each day chemotherapy is administered. Trilaciclib transiently arrests cyclin-dependent kinase 4/6 (CDK4/6)-dependent hematopoietic stem and progenitor and immune cells in the G1 phase of the cell cycle during chemotherapy exposure, protecting these cells from chemotherapy-induced damage [14,15,16]. In 2019, trilaciclib received breakthrough designation from the US Food and Drug Administration (FDA) and, in 2021, was approved by the FDA to decrease the incidence of chemotherapy-induced myelosuppression in adult patients when administered prior to a platinum/etoposide-containing regimen or topotecan-containing regimen for extensive-stage small cell lung cancer (ES-SCLC) on the basis of the results from three randomized, placebo-controlled phase II studies [17,18,19].
Trilaciclib has been shown to favorably alter the tumor immune microenvironment in in vivo murine syngeneic models [15, 20, 21], in an ex vivo patient-derived organotypic tumor spheroid culture system [20], and in a clinical setting in patients with ES-SCLC [15, 17]. Specifically, trilaciclib has been shown to enhance T-cell activation and the production of cytokines and chemokines [15, 20], to promote a favorable tumor immune microenvironment by increasing the intratumoral ratio of effector T cells to regulatory T cells (Tregs) and the number of activated T cells in the periphery [15], to inhibit immunosuppression by Tregs [15, 20], to significantly increase the expansion of T-cell clones [15, 17], and to enhance the induction of memory cluster of differentiation (CD)8+ T cells [21]. Consistent with its known mechanism of action, trilaciclib may elicit some of these immune effects by protecting lymphocytes from chemotherapy-induced damage [22].
The efficacy and safety of trilaciclib in patients with mTNBC have been investigated in a randomized phase II trial (NCT02978716) [23, 24]. Treatment with trilaciclib prior to gemcitabine plus carboplatin (GCb) did not lead to a significant improvement in duration and occurrence of severe neutropenia (primary endpoint); however, overall survival (OS; secondary endpoint) was improved for patients who received trilaciclib plus GCb compared with those who received GCb alone (median 19.8 vs. 12.6 months, respectively) [23]. In subgroup analyses, OS was prolonged irrespective of CDK4/6 dependence and PD-L1 status, but benefit was greater in the PD-L1-positive population. OS was also more pronounced in, but not exclusive to, patients with higher immune-related gene expression [24]. Lastly, administering trilaciclib enhanced T-cell activation, as evidenced by an enrichment of new T-cell clones and decreased Simpson clonality in peripheral blood [23, 24].
The aim of the current analysis was to further investigate potential immune mechanisms of trilaciclib in mTNBC through the analysis of immune cell subsets and molecular profiling in peripheral blood and tumor samples, respectively.
Materials and methods
Study design and participants
This analysis is based on data from a multicenter, randomized, open-label, phase II trial including patients aged ≥ 18 years with mTNBC who had received up to two prior chemotherapy regimens for locally recurrent TNBC or mTNBC (NCT02978716) [23, 24]. Patients were randomized (1:1:1) to 21-day treatment cycles: GCb (gemcitabine 1000 mg/m2, carboplatin area under the curve 2) alone on days 1 and 8; trilaciclib 240 mg/m2 within 4 h prior to GCb on days 1 and 8; or trilaciclib alone on days 1 and 8 and trilaciclib within 4 h prior to GCb on days 2 and 9.
Antitumor efficacy endpoints and assessments
Efficacy and survival outcomes were analyzed as prespecified secondary endpoints and included objective response rate (confirmed complete or partial response) assessed in response-evaluable patients, and progression-free survival and OS, assessed in the intention-to-treat population. Objective response rate and progression-free survival were investigator assessed according to Response Evaluation Criteria in Solid Tumours (RECIST) version 1.1, based on the May 15, 2020, data cut-off. For tumor assessment, computed tomography or magnetic resonance imaging was performed at screening and at protocol-specified intervals (every 9 weeks for the first 6 months, then every 12 weeks thereafter) until disease progression, withdrawal of consent, or receipt of subsequent anticancer therapy. OS was analyzed following the final database lock on July 17, 2020. Genetic and/or expression markers in blood and tumors and immunologic markers, including PD-L1 expression, were analyzed as post hoc exploratory objectives. Baseline PD-L1 status was measured using the Ventana SP142 PD-L1 assay; tumors were scored as PD-L1 positive if the proportion of PD-L1-expressing tumor-infiltrating immune cells was ≥ 1% and PD-L1 negative if < 1%, per the assay interpretation guide for TNBC tumors [25].
Peripheral immune cell population and function analysis
Peripheral blood was collected prior to and during treatment for flow cytometric analysis; for the purposes of this analysis, samples were collected prior to treatment on the first day of the first and third treatment cycles (C1D1 and C3D1, respectively). Blood was collected in Cyto-Chex® (Streck) and sodium heparin tubes and shipped at ambient temperature on the day of collection for processing. Whole blood was used for intracellular cytokine staining and surface staining. Staining and flow cytometric assays were performed by a contract research organization (Covance Central Laboratory Services; Indianapolis, Indiana, USA).
Tumor gene expression analysis
Genomic DNA and total RNA were simultaneously purified and sequenced as previously described from formalin-fixed, paraffin-embedded (FFPE) diagnostic tumor samples collected at baseline [23]. Purification was performed using the AllPrep DNA/RNA FFPE kit (QIAGEN; Germantown, Maryland, USA). Libraries were prepared using TruSeq RNA and DNA Exome kits for RNA-Seq and DNA-Seq, respectively (Illumina; San Diego, California, USA). Cluster generation and sequencing of libraries were performed on the Illumina HiSeq system. Gene expression read counts and fragments per kilobase of exon per million mapped reads (FPKM) were quantified using RNA-Seq by Expectation Maximization (RSEM) software [26]. RNA-Seq samples in which < 30% of RNA fragments were > 200 nucleotides in length (DV200) were excluded from the analysis. Differentially expressed genes between trilaciclib responders (complete or partial response) and non‑responders (stable or progressive disease), at an adjusted P value of < 0.05, were identified using the DESeq2 package [27]. Gene set enrichment analysis (GSEA) was performed using GSEA_4.1.0 software (number of permutations: 10,000; permutation type: phenotype) [28, 29] using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database (c2.cp.kegg.v7.4).
Tumor immune microenvironment analysis
The Tumor Inflammation Signature (TIS) [30] was used to assess the tumor immune microenvironment at baseline. The TIS is an investigational 18-gene signature that detects a pre-existing but suppressed adaptive immune response within tumors by measuring the expression of genes associated with antigen-presentation cell abundance (PSMB10, HLA-DQA1, HLA-DRB1, CMKLR1), T-cell and natural killer (NK)-cell abundance (HLA-E, NKG7, CD8A), interferon (IFN) activity (CCL5, CXCL9, CD27, CXCR6, IDO1, STAT1), and T-cell exhaustion (TIGIT, LAG3, CD274, PDCDILG2, CD276) [30]. TIS or signature scores were calculated as an average of the expression values (quantile-normalized and log10-transformed) of the respective gene sets.
Statistical methods
Statistical comparisons of cell numbers/ratios and TIS scores for different time points and patient groups were performed using the Wilcoxon signed-rank test. Plots were created using the ggplot2 and EnhancedVolcano R packages [31, 32].
Results
Participants and treatment
As of July 17, 2020, median (range) duration of follow-up was 8.4 (0.1–25.7) months for the 34 patients who received GCb alone, 14.0 (1.3–33.6) months for the 33 patients who received trilaciclib prior to GCb on days 1 and 8, and 15.3 (3.5–33.7) months for the 35 patients who received trilaciclib alone on days 1 and 8 and trilaciclib prior to GCb on days 2 and 9. Antitumor response status was available for 58 of the 68 patients who received trilaciclib prior to GCb: 27 patients (46.6%) had an antitumor response with trilaciclib plus GCb (trilaciclib responders), and 31 (53.4%) had no response (non‑responders).
Analysis of immune subsets and T-cell function at C1D1 versus C3D1 in patients receiving trilaciclib prior to GCb or GCb alone
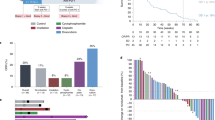
Patients who received trilaciclib prior to GCb had fewer total T cells (P = 0.064) and significantly fewer CD8+ T cells (P = 0.013) and myeloid-derived suppressor cells (MDSCs; P < 0.0001) at C3D1 compared with C1D1, whereas no significant differences were observed between these time points in patients who received GCb alone (Fig. 1a). Administering trilaciclib prior to GCb greatly enhanced T-cell effector function compared with administering GCb alone, as evidenced by significant increases in the number of cytokine-producing CD4+ and CD8+ T cells (Fig. 1b) from C1D1 to C3D1. No significant differences from baseline in T-cell effector functions were observed in patients who received GCb alone.
Changes to a immune cell populations and b T-cell function in peripheral blood over two cycles (C1D1 vs. C3D1) in patients receiving trilaciclib prior to GCb or GCb alone. C1D1 cycle 1, day 1 C3D1 cycle 3, day 1 CD cluster of differentiation, GCb gemcitabine plus carboplatin, IFNγ interferon gamma, IL interleukin, MDSC myeloid-derived suppressor cell, Treg regulatory T cell
Immune cell populations and T-cell function analysis among trilaciclib responders versus non-responders
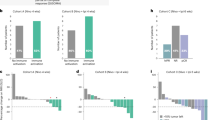
To determine if the impact on T-cell effector function was attributable to clinical outcomes, data from responders and non‑responders who were treated with trilaciclib were compared. After two cycles, T-cell numbers were maintained in trilaciclib responders but significantly reduced in non‑responders (P = 0.0034; Fig. 2), with significant reductions in CD4+ T cells (P = 0.009) and CD8+ T cells (P = 0.0066) contributing to the overall reduction. Tregs were maintained among both responders and non‑responders, whereas MDSCs were significantly reduced among both responders (P = 0.0046) and non‑responders (P = 0.013; Fig. 2). T-cell function was maintained or improved in responders but was maintained or reduced in non‑responders. Human leukocyte antigen–DR isotype (HLA-DR) expression, a marker of T-cell activation, was also downregulated in trilaciclib non‑responders (Fig. 3).
Tumor gene expression analysis (trilaciclib responders vs. non-responders at baseline)
Analysis of tumor samples revealed 69 differentially expressed genes (adjusted P < 0.05) between trilaciclib responders (n = 15) and non‑responders (n = 17) at baseline (Fig. 4a). In total, 23 genes were upregulated and 46 genes were downregulated (Table 1).
a Differential gene expression analysis (adjusted P value < 0.05; red dots, │log2FC│ > 1; blue dots, │log2FC│ ≤ 1) and b tumor inflammation signatures in tumor samples from trilaciclib responders and non-responders. False discovery rate < 0.25. FC fold change, IFNγ interferon gamma, NK natural killer, TIS tumor inflammation score
KEGG pathways upregulated in trilaciclib responders (FDR < 0.25) were T-cell receptor signaling, antigen processing and presentation, NK-cell–mediated cytotoxicity, nucleotide-binding oligomerization domain (NOD)-like receptor signaling, Toll-like receptor signaling, cytosolic DNA sensing, graft-versus-host disease, and glycosphingolipid biosynthesis. Analysis of immune gene signatures revealed trends toward a higher overall TIS score at baseline among responders versus non‑responders (Fig. 4b). Trends toward increased TIS score were observed for T-cell/NK-cell abundance, IFN activity, and T-cell exhaustion in trilaciclib responders.
Discussion
Data from this exploratory analysis provide further evidence of a trilaciclib-mediated antitumor immune response among patients with mTNBC [23, 24]. Patients who received trilaciclib prior to GCb had fewer, but more functional, peripheral T cells and fewer MDSCs after two treatment cycles than patients who received GCb alone. Furthermore, peripheral T-cell numbers, function, and activation were maintained in trilaciclib responders after two treatment cycles but reduced in non‑responders, whereas Treg numbers were maintained and MDSCs were significantly reduced in both groups.
These findings support prior research indicating that, following transient G1 arrest, the proportion of immunosuppressive cells in the tumor microenvironment is decreased and effector T-cell function is enhanced [15]. The benefits of transient T-cell inhibition have also been shown in a drug-regulatable platform, wherein transient inhibition of chimeric antigen receptor expression directed T cells to a memory-like phenotype and restored antitumor functionality in a cell population that had already developed features of exhaustion [33]. CDK4/6 inhibition has previously been shown to enhance T-cell function via the de-repression of the Nuclear Factor of Activated T cell (NFAT) family of proteins. De-repression leads to the activation of downstream genes that regulate T-cell function, resulting in reduced proliferation but increased tumor infiltration and activation of effector T cells [20]. A reduction in the number of MDSCs, which are critical drivers of immune suppression in the tumor microenvironment, suggests that trilaciclib can reduce suppression of the host antitumor immune response to enhance immune-mediated antitumor responses [34].
Data from diagnostic tumor samples collected at baseline showed that genes involved in immune cell activation were upregulated among trilaciclib responders, and there was a trend toward higher TIS scores. In general, although higher TIS scores are not associated with increased OS in breast cancer, they are associated with improved prognosis in those patients with the highest 10% of TIS scores [30]. All KEGG pathways that were upregulated at baseline in trilaciclib responders were pathways that are involved in the generation of an immune response and, except for the graft-versus-host disease pathway, are also necessary for an antitumor response.
A trend for higher T-cell exhaustion at baseline may indicate that patients had a greater existing immune response and, consequently, higher existing T-cell infiltration into the tumor. Exhausted T-cell profiles in the tumor microenvironment [35], or in peripheral blood [36], have previously been associated with better responses. Preclinical studies have shown that, following CDK4/6 inhibition, intratumoral CD8+ T cells display markedly reduced expression of the inhibitory immune receptors PD-1, Tim-3, CTLA-4, and LAG-3—all markers of T-cell exhaustion [37]—potentially enhancing the susceptibility of such tumors to antitumor immune responses. It is possible, therefore, that differential gene expression profiles at baseline, including T-cell exhaustion, may be predictive of response to trilaciclib-containing regimens.
Additional analyses were conducted to compare results by PD-L1 status at baseline. Increased levels of peripheral memory CD8+ T cells and naïve CD8+ T cells were observed after two cycles in trilaciclib responders, regardless of PD-L1 status. However, greater peripheral immune responses and a trend toward an enriched TIS were identified in PD-L1-positive trilaciclib responders at baseline compared with non‑responders. These data support previous research showing that CDK4/6 inhibition promotes the formation of memory CD8+ T cells, which is proposed to occur via upregulation of MXD4 and resultant downregulation of Myc activity during T-cell activation [38]. Furthermore, because an enriched tumor microenvironment suggests better immune cell infiltration, collectively, these data may explain, at least in part, why subgroup analysis of the final OS results from this trial demonstrated larger OS benefit in the PD-L1-positive population [24].
Limitations of this study include the small sample size, particularly in the responder subsets. Moreover, antitumor efficacy outcomes were not the primary study endpoints. The sample size was powered to show superiority among patients who received trilaciclib prior to GCb over those who received GCb alone for at least one primary endpoint (duration of severe neutropenia in cycle 1 or occurrence of severe neutropenia during the treatment period). As such, comparisons of secondary endpoints, including antitumor responses, should be considered exploratory and interpreted with caution. However, the findings of this hypothesis-generating analysis were consistent with, and supportive of, a previous exploratory analysis of the same study [24] and suggest that the improvement in OS observed in patients with mTNBC could be due to increased antitumor immunity, mediated by trilaciclib [24]. Clinical trials are ongoing to explore this further. The phase III PRESERVE 2 trial, investigating trilaciclib or placebo administered prior to GCb in patients with locally advanced unresectable TNBC or mTNBC, has OS as the primary endpoint (NCT04799249). In this study, the impact of trilaciclib on tumor-associated immune responses will be evaluated by comparing immunophenotypic changes between tumor biopsies from patients receiving trilaciclib or placebo. In addition, a phase II mechanism-of-action trial in the neoadjuvant TNBC setting is underway (NCT05112536). The primary objective is to evaluate the immune-based mechanism of action of trilaciclib after a single dose, as measured by changes in the CD8+/Treg ratio in tumor tissue. Pathologic complete response, safety and tolerability, and additional exploratory immune biomarker endpoints will also be assessed.
Overall, these data contribute to a growing body of evidence that transient administration of trilaciclib prior to GCb may enhance antitumor efficacy by both protecting immune cells from chemotherapy-induced damage and modulating the composition and response of immune cell subsets. Data from ongoing clinical studies are critical to confirming the underlying immune mechanisms and to identifying biomarkers that will clearly distinguish between trilaciclib responders and non‑responders.
Data availability
All data generated or analyzed during this study are included in this published article.
Abbreviations
- C1D1:
-
Cycle 1, day 1
- C3D1:
-
Cycle 3, day 1
- CD:
-
Cluster of differentiation
- CDK4/6:
-
Cyclin-dependent kinase 4/6
- ES-SCLC:
-
Extensive-stage small cell lung cancer
- FDR:
-
False discovery rate
- FFPE:
-
Formalin fixed, paraffin embedded
- FPKM:
-
Fragments per kilobase of exon per million mapped reads
- GCb:
-
Gemcitabine plus carboplatin
- GSEA:
-
Gene set enrichment analysis
- HLA-DR:
-
Human leukocyte antigen–DR isotype
- IFN(γ):
-
Interferon (gamma)
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MDSC:
-
Myeloid-derived suppressor cell
- mTNBC:
-
Metastatic triple-negative breast cancer
- NK:
-
Natural killer
- NOD:
-
Nucleotide-binding oligomerization domain
- OS:
-
Overall survival
- PD-L1:
-
Programmed death-ligand 1
- RECIST:
-
Response Evaluation Criteria in Solid Tumours
- RSEM:
-
RNA-Seq by Expectation Maximization
- TIS:
-
Tumor Inflammation Signature
- TNBC:
-
Triple-negative breast cancer
- Treg:
-
Regulatory T cell
References
Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA, Lickley LA, Rawlinson E, Sun P, Narod SA (2007) Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res 13:4429–4434. https://doi.org/10.1158/1078-0432.CCR-06-3045
Foulkes WD, Smith IE, Reis-Filho JS (2010) Triple-negative breast cancer. N Engl J Med 363:1938–1948. https://doi.org/10.1056/NEJMra1001389
Li X, Yang J, Peng L, Sahin AA, Huo L, Ward KC, O’Regan R, Torres MA, Meisel JL (2017) Triple-negative breast cancer has worse overall survival and cause-specific survival than non-triple-negative breast cancer. Breast Cancer Res Treat 161:279–287. https://doi.org/10.1007/s10549-016-4059-6
Li Y, Zhan Z, Yin X, Fu S, Deng X (2021) Targeted therapeutic strategies for triple-negative breast cancer. Front Oncol 11:731535. https://doi.org/10.3389/fonc.2021.731535
Twelves C, Jove M, Gombos A, Awada A (2016) Cytotoxic chemotherapy: still the mainstay of clinical practice for all subtypes metastatic breast cancer. Crit Rev Oncol Hematol 100:74–87. https://doi.org/10.1016/j.critrevonc.2016.01.021
Beckers RK, Selinger CI, Vilain R, Madore J, Wilmott JS, Harvey K, Holliday A, Cooper CL, Robbins E, Gillett D, Kennedy CW, Gluch L, Carmalt H, Mak C, Warrier S, Gee HE, Chan C, McLean A, Walker E, McNeil CM, Beith JM, Swarbrick A, Scolyer RA, O’Toole SA (2016) Programmed death ligand 1 expression in triple-negative breast cancer is associated with tumour-infiltrating lymphocytes and improved outcome. Histopathology 69:25–34. https://doi.org/10.1111/his.12904
Mittendorf EA, Philips AV, Meric-Bernstam F, Qiao N, Wu Y, Harrington S, Su X, Wang Y, Gonzalez-Angulo AM, Akcakanat A, Chawla A, Curran M, Hwu P, Sharma P, Litton JK, Molldrem JJ, Alatrash G (2014) PD-L1 expression in triple-negative breast cancer. Cancer Immunol Res 2:361–370. https://doi.org/10.1158/2326-6066.CIR-13-0127
Cortes J, Cescon DW, Rugo HS, Nowecki Z, Im SA, Yusof MM, Gallardo C, Lipatov O, Barrios CH, Holgado E, Iwata H, Masuda N, Otero MT, Gokmen E, Loi S, Guo Z, Zhao J, Aktan G, Karantza V, Schmid P, Investigators K (2020) Pembrolizumab plus chemotherapy versus placebo plus chemotherapy for previously untreated locally recurrent inoperable or metastatic triple-negative breast cancer (KEYNOTE-355): a randomised, placebo-controlled, double-blind, phase 3 clinical trial. Lancet 396:1817–1828. https://doi.org/10.1016/S0140-6736(20)32531-9
Cortes J, Rugo HS, Cescon DW, Im SA, Yusof MM, Gallardo C, Lipatov O, Barrios CH, Perez-Garcia J, Iwata H, Masuda N, Torregroza Otero M, Gokmen E, Loi S, Guo Z, Zhou X, Karantza V, Pan W, Schmid P, Investigators K (2022) Pembrolizumab plus chemotherapy in advanced triple-negative breast cancer. N Engl J Med 387:217–226. https://doi.org/10.1056/NEJMoa2202809
Robson M, Im SA, Senkus E, Xu B, Domchek SM, Masuda N, Delaloge S, Li W, Tung N, Armstrong A, Wu W, Goessl C, Runswick S, Conte P (2017) Olaparib for metastatic breast cancer in patients with a germline BRCA mutation. N Engl J Med 377:523–533. https://doi.org/10.1056/NEJMoa1706450
Litton JK, Rugo HS, Ettl J, Hurvitz SA, Goncalves A, Lee KH, Fehrenbacher L, Yerushalmi R, Mina LA, Martin M, Roche H, Im YH, Quek RGW, Markova D, Tudor IC, Hannah AL, Eiermann W, Blum JL (2018) Talazoparib in patients with advanced breast cancer and a germline BRCA mutation. N Engl J Med 379:753–763. https://doi.org/10.1056/NEJMoa1802905
Bardia A, Hurvitz SA, Tolaney SM, Loirat D, Punie K, Oliveira M, Brufsky A, Sardesai SD, Kalinsky K, Zelnak AB, Weaver R, Traina T, Dalenc F, Aftimos P, Lynce F, Diab S, Cortes J, O’Shaughnessy J, Dieras V, Ferrario C, Schmid P, Carey LA, Gianni L, Piccart MJ, Loibl S, Goldenberg DM, Hong Q, Olivo MS, Itri LM, Rugo HS, Investigators ACT (2021) Sacituzumab govitecan in metastatic triple-negative breast cancer. N Engl J Med 384:1529–1541. https://doi.org/10.1056/NEJMoa2028485
Gupta GK, Collier AL, Lee D, Hoefer RA, Zheleva V, Siewertsz van Reesema LL, Tang-Tan AM, Guye ML, Chang DZ, Winston JS, Samli B, Jansen RJ, Petricoin EF, Goetz MP, Bear HD, Tang AH (2020) Perspectives on triple-negative breast cancer: current treatment strategies, unmet needs, and potential targets for future therapies. Cancers (Basel) 12:2392. https://doi.org/10.3390/cancers12092392
He S, Roberts PJ, Sorrentino JA, Bisi JE, Storrie-White H, Tiessen RG, Makhuli KM, Wargin WA, Tadema H, van Hoogdalem EJ, Strum JC, Malik R, Sharpless NE (2017) Transient CDK4/6 inhibition protects hematopoietic stem cells from chemotherapy-induced exhaustion. Sci Transl Med 9:eaa13986. https://doi.org/10.1126/scitranslmed.aal3986
Lai AY, Sorrentino JA, Dragnev KH, Weiss JM, Owonikoko TK, Rytlewski JA, Hood J, Yang Z, Malik RK, Strum JC, Roberts PJ (2020) CDK4/6 inhibition enhances antitumor efficacy of chemotherapy and immune checkpoint inhibitor combinations in preclinical models and enhances T-cell activation in patients with SCLC receiving chemotherapy. J Immunother Cancer 8:e000847. https://doi.org/10.1136/jitc-2020-000847
Li C, Hart L, Owonikoko TK, Aljumaily R, Rocha Lima CM, Conkling PR, Webb RT, Jotte RM, Schuster S, Edenfield WJ, Smith DA, Sale M, Roberts PJ, Malik RK, Sorrentino JA (2021) Trilaciclib dose selection: an integrated pharmacokinetic and pharmacodynamic analysis of preclinical data and Phase Ib/IIa studies in patients with extensive-stage small cell lung cancer. Cancer Chemother Pharmacol 87:689–700. https://doi.org/10.1007/s00280-021-04239-9
Daniel D, Kuchava V, Bondarenko I, Ivashchuk O, Reddy S, Jaal J, Kudaba I, Hart L, Matitashvili A, Pritchett Y, Morris SR, Sorrentino JA, Antal JM, Goldschmidt J (2020) Trilaciclib prior to chemotherapy and atezolizumab in patients with newly diagnosed extensive-stage small cell lung cancer: a multicentre, randomised, double-blind, placebo-controlled phase II trial. Int J Cancer 148:2557–2570. https://doi.org/10.1002/ijc.33453
Hart LL, Ferrarotto R, Andric ZG, Beck JT, Subramanian J, Radosavljevic DZ, Zaric B, Hanna WT, Aljumaily R, Owonikoko TK, Verhoeven D, Xiao J, Morris SR, Antal JM, Hussein MA (2021) Myelopreservation with trilaciclib in patients receiving topotecan for small cell lung cancer: results from a randomized, double-blind, placebo-controlled phase II study. Adv Ther 38:350–365. https://doi.org/10.1007/s12325-020-01538-0
Weiss JM, Csoszi T, Maglakelidze M, Hoyer RJ, Beck JT, Domine Gomez M, Lowczak A, Aljumaily R, Rocha Lima CM, Boccia RV, Hanna W, Nikolinakos P, Chiu VK, Owonikoko TK, Schuster SR, Hussein MA, Richards DA, Sawrycki P, Bulat I, Hamm JT, Hart LL, Adler S, Antal JM, Lai AY, Sorrentino JA, Yang Z, Malik RK, Morris SR, Roberts PJ, Dragnev KH, Group GTS (2019) Myelopreservation with the CDK4/6 inhibitor trilaciclib in patients with small-cell lung cancer receiving first-line chemotherapy: a phase Ib/randomized phase II trial. Ann Oncol 30:1613–1621. https://doi.org/10.1093/annonc/mdz278
Deng J, Wang ES, Jenkins RW, Li S, Dries R, Yates K, Chhabra S, Huang W, Liu H, Aref AR, Ivanova E, Paweletz CP, Bowden M, Zhou CW, Herter-Sprie GS, Sorrentino JA, Bisi JE, Lizotte PH, Merlino AA, Quinn MM, Bufe LE, Yang A, Zhang Y, Zhang H, Gao P, Chen T, Cavanaugh ME, Rode AJ, Haines E, Roberts PJ, Strum JC, Richards WG, Lorch JH, Parangi S, Gunda V, Boland GM, Bueno R, Palakurthi S, Freeman GJ, Ritz J, Haining WN, Sharpless NE, Arthanari H, Shapiro GI, Barbie DA, Gray NS, Wong KK (2018) CDK4/6 inhibition augments antitumor immunity by enhancing T-cell activation. Cancer Discov 8:216–233. https://doi.org/10.1158/2159-8290.CD-17-0915
Lelliott EJ, Kong IY, Zethoven M, Ramsbottom KM, Martelotto LG, Meyran D, Zhu JJ, Costacurta M, Kirby L, Sandow JJ, Lim L, Dominguez PM, Todorovski I, Haynes NM, Beavis PA, Neeson PJ, Hawkins ED, McArthur GA, Parish IA, Johnstone RW, Oliaro J, Sheppard KE, Kearney CJ, Vervoort SJ (2021) CDK4/6 inhibition promotes antitumor immunity through the induction of T-cell memory. Cancer Discov 11:2582–2601. https://doi.org/10.1158/2159-8290.CD-20-1554
Lelliott EJ, Sheppard KE, McArthur GA (2022) Harnessing the immunotherapeutic potential of CDK4/6 inhibitors in melanoma: is timing everything? NPJ Precis Oncol 6:26. https://doi.org/10.1038/s41698-022-00273-9
Tan AR, Wright GS, Thummala AR, Danso MA, Popovic L, Pluard TJ, Han HS, Vojnovic Z, Vasev N, Ma L, Richards DA, Wilks ST, Milenkovic D, Yang Z, Antal JM, Morris SR, O’Shaughnessy J (2019) Trilaciclib plus chemotherapy versus chemotherapy alone in patients with metastatic triple-negative breast cancer: a multicentre, randomised, open-label, phase 2 trial. Lancet Oncol 20:1587–1601. https://doi.org/10.1016/S1470-2045(19)30616-3
Tan AR, Wright GS, Thummala AR, Danso MA, Popovic L, Pluard TJ, Han HS, Vojnovic Z, Vasev N, Ma L, Richards DA, Wilks ST, Milenkovic D, Xiao J, Sorrentino J, Horton J, O’Shaughnessy J (2022) Trilaciclib prior to chemotherapy in patients with metastatic triple-negative breast cancer: final efficacy and subgroup analysis from a randomized phase II study. Clin Cancer Res 28:629–636. https://doi.org/10.1158/1078-0432.CCR-21-2272
US Food and Drug Administration (2019) Ventana PD-L1 (SP142) assay. https://www.accessdata.fda.gov/cdrh_docs/pdf16/p160002s009c.pdf. Accessed June 28, 2022
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics 12:323. https://doi.org/10.1186/1471-2105-12-323
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Mootha VK, Lindgren CM, Eriksson KF, Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E, Ridderstrale M, Laurila E, Houstis N, Daly MJ, Patterson N, Mesirov JP, Golub TR, Tamayo P, Spiegelman B, Lander ES, Hirschhorn JN, Altshuler D, Groop LC (2003) PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet 34:267–273. https://doi.org/10.1038/ng1180
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES, Mesirov JP (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550. https://doi.org/10.1073/pnas.0506580102
Danaher P, Warren S, Lu R, Samayoa J, Sullivan A, Pekker I, Wallden B, Marincola FM, Cesano A (2018) Pan-cancer adaptive immune resistance as defined by the tumor inflammation signature (TIS): results from the cancer genome atlas (TCGA). J Immunother Cancer 6:63. https://doi.org/10.1186/s40425-018-0367-1
Wickham H (2009) ggplot2: Elegant Graphics for Data Analysis. https://ggplot2.tidyverse.org. Accessed June 28, 2022
Blighe K, Sharmila R, Lewis M (2022) EnhancedVolcano: publication-ready volcano plots with enhanced colouring and labeling. https://bioconductor.org/packages/devel/bioc/vignettes/EnhancedVolcano/inst/doc/EnhancedVolcano.html. Accessed November 25, 2022
Weber EW, Parker KR, Sotillo E, Lynn RC, Anbunathan H, Lattin J, Good Z, Belk JA, Daniel B, Klysz D, Malipatlolla M, Xu P, Bashti M, Heitzeneder S, Labanieh L, Vandris P, Majzner RG, Qi Y, Sandor K, Chen LC, Prabhu S, Gentles AJ, Wandless TJ, Satpathy AT, Chang HY, Mackall CL (2021) Transient rest restores functionality in exhausted CAR-T cells through epigenetic remodeling. Science 372:eaba1786. https://doi.org/10.1126/science.aba1786
Shou D, Wen L, Song Z, Yin J, Sun Q, Gong W (2016) Suppressive role of myeloid-derived suppressor cells (MDSCs) in the microenvironment of breast cancer and targeted immunotherapies. Oncotarget 7:64505–64511. https://doi.org/10.18632/oncotarget.11352
Daud AI, Loo K, Pauli ML, Sanchez-Rodriguez R, Sandoval PM, Taravati K, Tsai K, Nosrati A, Nardo L, Alvarado MD, Algazi AP, Pampaloni MH, Lobach IV, Hwang J, Pierce RH, Gratz IK, Krummel MF, Rosenblum MD (2016) Tumor immune profiling predicts response to anti-PD-1 therapy in human melanoma. J Clin Invest 126:3447–3452. https://doi.org/10.1172/JCI87324
Terranova-Barberio M, Pawlowska N, Dhawan M, Moasser M, Chien AJ, Melisko ME, Rugo H, Rahimi R, Deal T, Daud A, Rosenblum MD, Thomas S, Munster PN (2020) Exhausted T cell signature predicts immunotherapy response in ER-positive breast cancer. Nat Commun 11:3584. https://doi.org/10.1038/s41467-020-17414-y
Goel S, DeCristo MJ, Watt AC, BrinJones H, Sceneay J, Li BB, Khan N, Ubellacker JM, Xie S, Metzger-Filho O, Hoog J, Ellis MJ, Ma CX, Ramm S, Krop IE, Winer EP, Roberts TM, Kim HJ, McAllister SS, Zhao JJ (2017) CDK4/6 inhibition triggers anti-tumour immunity. Nature 548:471–475. https://doi.org/10.1038/nature23465
Heckler M, Ali LR, Clancy-Thompson E, Qiang L, Ventre KS, Lenehan P, Roehle K, Luoma A, Boelaars K, Peters V, McCreary J, Boschert T, Wang ES, Suo S, Marangoni F, Mempel TR, Long HW, Wucherpfennig KW, Dougan M, Gray NS, Yuan GC, Goel S, Tolaney SM, Dougan SK (2021) Inhibition of CDK4/6 promotes CD8 T-cell memory formation. Cancer Discov 11:2564–2581. https://doi.org/10.1158/2159-8290.CD-20-1540
Acknowledgements
We acknowledge Aaron Stevens and Jessica Sorrentino, former employees of G1 Therapeutics, Inc., for their contributions toward this analysis and interpretation of data. We also thank all the patients, their families, and study personnel for participating in the study.
Funding
This work was supported by G1 Therapeutics, Inc. The study sponsor was involved in the study design, in the collection, analysis, and interpretation of the data, in the writing of the report, and in the decision to submit the paper for publication. Medical writing assistance was provided by Alligent Europe (Envision Pharma Group), funded by G1 Therapeutics, Inc.
Author information
Authors and Affiliations
Contributions
ART and JOS contributed to the design and implementation of the clinical research and to the acquisition of data. SC, SA, and JSY conceived and designed the analysis. All authors were responsible for the analysis and interpretation of results; provided critical feedback and helped shape the research, analysis, and development of the manuscript; and read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Antoinette R. Tan reports institutional clinical trial support and personal fees from G1 Therapeutics, Inc.; outside of the submitted work, institutional clinical trial support from Arvinas, Genentech/Roche, Merck, and Pfizer, and personal fees from AstraZeneca, Genentech/Roche, Jazz Pharmaceuticals, Novartis, Puma, Seagen, and Stemline Therapeutics. Joyce O’Shaughnessy reports institutional clinical trial support and personal fees from G1 Therapeutics, Inc.; outside of the submitted work, personal fees from AbbVie, Agendia, Amgen, Aptitude Health, AstraZeneca, BMS, Celgene, Eisai, Genentech, Immunomedics, Ipsen, Jounce Therapeutics, Lilly, Merck, Myriad, Novartis, Odonate Therapeutics, Pfizer, Prime Oncology, Puma Biotechnology, Roche, Seattle Genetics, and Syndax Pharmaceuticals. Subing Cao and Sarah Ahn (at the time of the study), and John S. Yi are paid employees and shareowners of G1 Therapeutics, Inc.
Ethical approval
The study (NCT02978716) was designed and conducted in accordance with the Declaration of Helsinki and the Good Clinical Practice guidelines of the International Council for Harmonisation. The protocol and all study-related materials were approved by the institutional review board or independent ethics committee of each investigational site.
Consent to participate
Written informed consent was obtained from all individual participants included in the study.
Consent to publish
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tan, A.R., O’Shaughnessy, J., Cao, S. et al. Investigating potential immune mechanisms of trilaciclib administered prior to chemotherapy in patients with metastatic triple-negative breast cancer. Breast Cancer Res Treat 201, 307–316 (2023). https://doi.org/10.1007/s10549-023-07009-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-023-07009-8