Abstract
Purpose
We compared a simple breast cancer risk prediction model, BRISK (which includes mammographic density, polygenic risk and clinical factors), against a similar model with more risk factors (simplified Rosner) and against two commonly used clinical models (Gail and IBIS).
Methods
Using nested case–control data from the Nurses’ Health Study, we compared the models’ association, discrimination and calibration. Classification performance was compared between Gail and BRISK for 5-year risks and between IBIS and BRISK for remaining lifetime risk.
Results
The odds ratio per standard deviation was 1.43 (95% CI 1.32, 1.55) for BRISK 5-year risk, 1.07 (95% CI 0.99, 1.14) for Gail 5-year risk, 1.72 (95% CI 1.59, 1.87) for simplified Rosner 10-year risk, 1.51 (95% CI 1.41, 1.62) for BRISK remaining lifetime risk and 1.26 (95% CI 1.16, 1.36) for IBIS remaining lifetime risk. The area under the receiver operating characteristic curve (AUC) was improved for BRISK over Gail for 5-year risk (AUC = 0.636 versus 0.511, P < 0.0001) and for BRISK over IBIS for remaining lifetime risk (AUC = 0.647 versus 0.571, P < 0.0001). BRISK was well calibrated for the estimation of both 5-year risk (expected/observed [E/O] = 1.03; 95% CI 0.73, 1.46) and remaining lifetime risk (E/O = 1.01; 95% CI 0.86, 1.17). The Gail 5-year risk (E/O = 0.85; 95% CI 0.58, 1.24) and IBIS remaining lifetime risk (E/O = 0.73; 95% CI 0.60, 0.87) were not well calibrated, with both under-estimating risk. BRISK improves classification of risk compared to Gail 5-year risk (NRI = 0.31; standard error [SE] = 0.031) and IBIS remaining lifetime risk (NRI = 0.287; SE = 0.035).
Conclusion
BRISK performs better than two commonly used clinical risk models and no worse compared to a similar model with more risk factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
For decades, women have been told that they have an increased risk of breast cancer if they have a family history of the disease [1], and more recently, women have been told that they have increased risk of breast cancer if they have mammographically dense breasts [2]. Genetic risk derived from common variants throughout the genome can also be used to predict risk of breast cancer [3] but this information is not readily available to women. This fragmented approach to risk prediction can result in women being given inaccurate risk information and can mean that women get conflicting and confusing advice.
Our goal is to accurately predict breast cancer risk for unaffected women by combining information on the key risk factors (including polygenic risk, mammographic density and family history) into a single simple model rather than by considering the risk factors separately [4,5,6]. This approach is of practical importance for both clinical and population health. In clinical practice, women at very high risk of breast cancer can be identified and given risk reduction options including medication or alternative screening modalities. While clinical implementation of reduced screening is not yet standard of care, women at very low risk of breast cancer can be identified as candidates for less frequent screening, with a concomitant financial benefit to population-based screening programs.
About 15% of women report having at least one first-degree relative with breast cancer [7,8,9,10], and a further 20% report having at least one affected second-degree relative [11]. The magnitude of familial risk depends on the closeness of the genetic relationship, the age at diagnosis of the affected relative and the age of the at-risk woman [1]. Familial risk is best captured by algorithms that model major genes and underlying genetic risk using multi-generational pedigree data [12,13,14,15]. Cancer family clinics use these tools when managing the small proportion of women who have several affected relatives and where there is the time and commitment to collect and validate extensive family cancer data. This approach is not applicable on a population basis.
Deeper understanding of breast cancer family history based on genetics led to the discovery of the first major susceptibility genes for breast cancer (BRCA1 and BRCA2) over 20 years ago [16]. Since then, additional highly and moderately penetrant risk variants have been identified in other genes [16]. These hereditary high-risk variants are rare. For example, BRCA1 and BRCA2 variants are found in only 1 in 400 women [17], but are associated with a full lifetime breast cancer risk of over 50% [18]. While it is important to identify women with these variants, they represent only about 5% of breast cancer cases [19]. The rarity of these mutations means they have limited impact on population health.
There are genetic implications at the population level if we look beyond highly or moderately penetrant variants. Large collaborative studies have identified single-nucleotide polymorphisms (SNPs) that are independently associated with breast cancer risk [20,21,22]. The breast cancer associations of these SNPs are small, with the odds ratio (OR) per risk allele typically being 1.1 or less [22], and empirically these risks appear to multiply [23]. These SNPs can be used to create a polygenic risk score (PRS) that can predict risk of breast cancer. Mavaddat et al. [3] first identified a 77-SNP PRS that has an OR per standard deviation (SD) of 1.46. More recently, Mavaddat et al. [24] identified a 313-SNP PRS that has an OR per SD of 1.61.
Mammographic density, visualized by the bright regions on a mammogram, can be measured on a continuous scale using a computer-assisted method (e.g. Cumulus) [25] or a fully automated method (e.g. Volpara) [26]. Mammographic density can also be assigned to categories by radiologists after visual inspection (e.g. BI-RADS classification) [27]. Percent mammographic density (adjusted for age and body mass index) is associated with risk of breast cancer [28]. Over 40% of women in the USA have dense breasts, more so at young ages [2]. Unfortunately, most women are unaware of risk factors other than family history and there is limited and often conflicting guidance for healthcare providers to recommend alternative screening modalities based on mammographic density alone.
These three major risk factors for breast cancer, and other risk factors such as body mass index and menopausal status, act largely independently [29]. Therefore, in terms of absolute risk, the combination of risk factors is extremely important for accurately determining risk. Information on these risk factors can be collected with a simple questionnaire or from electronic health records, while DNA analysis involves a simple buccal collection. This simplified approach allows for scalability of risk assessment in the general population without affecting the already-limited clinician-patient interaction.
In this paper we use a nested case–control dataset from the Nurses’ Health Study to assess the performance of a new risk model (BRISK) compared with the simplified Rosner model [30] and the standard clinical models Gail [31] and IBIS (version 7) [13]. Traditionally, the Gail model has been used for determining 5-year risks in the general population and IBIS has been used to predict remaining lifetime risk for women with a strong family history. BRISK was designed to combine data that is simple to collect in clinical practice with measures of mammographic density and polygenic risk that can be provided by other services.
Materials and methods
BRISK model
The model was constructed by bringing together evidence for the major risk factors for breast cancer. The input variables include age, number of affected first-degree relatives, age of youngest first-degree relative, number of affected second-degree relatives, percent mammographic density (or BI-RADs category), body mass index, and menopausal status. A description of the model is provided in the supplementary data.
The family history risks were based on those in the Collaborative Group on Hormonal Factors in Breast Cancer analysis [1]. These risks were smoothed and centered to have a population average risk of 1. The 313 SNPs in the PRS were those from Mavaddat et al. [24] and were combined to form a PRS using the approach of Mealiffe et al. [32]. The estimates for body mass index for pre- and post-menopausal women were taken from Hopper et al. [29]. The estimates for percent mammographic density were taken from unpublished analyses of the Australian Breast Cancer Family Registry [33] and the Australian Mammographic Density Twins and Sisters Study [34] and included estimates for percent mammographic density as a continuous measure and an alternative categorical measure to approximate BI-RADS classifications.
Participants
Our analyses used nested case–control data from the Nurses’ Health Study, which was established in 1976 and included 121,700 female registered nurses aged 30–55 years [35]. Questionnaires were mailed to women biennially to collect information on breast cancer risk factors, including age at menarche, age at first birth, parity, family history of breast cancer, height, weight, menopausal status, age at menopause, and hormone replacement therapy use. The nested case–control dataset comprised 1131 breast cancer cases and 1700 controls for whom questionnaire, mammographic density, and genotyping data were available. A subset of cases (n = 881) and controls (n = 1327) also had IBIS (version 7) risk predictions available for analysis. A further subset of the cases had estrogen receptor (ER) status available: 562 ER-positive and 106 ER-negative.
Statistical methods
We used standard univariate methods based on t-tests for continuous variables and contingency table methods for categorical variables to compare risk scores and covariates between cases vs. controls with results presented as OR per SD of risk. We used the area under the receiver operating characteristic curve (AUC) to compare the risk score distributions between cases vs. controls [36]. We compared the AUCs for competing risk models using the methods of DeLong [37]. For calibration, we compared the expected number of incident cases over 5 years for controls (median 5-year age-specific BRISK score ✕ number of women in 5-year age groups) to the number of incident cases that would be observed based on 5-year population incidence rates. Calibration was performed using controls because they are representative of the general population. Similar methods were used for estimated remaining lifetime risk. The median (rather than the mean) was used because the age-specific distribution of BRISK 5-year and remaining lifetime risk scores were strongly right-skewed. We used reclassification tables to compare pairs of risk models in terms of their ability to assign women to categories of risk based upon clinical thresholds used to guide chemoprevention or increased screening. National Comprehensive Cancer Network guidelines for breast cancer risk reduction recommend that 5-year risks be assessed using the Gail model, with patients exceeding a threshold of 1.67% be offered risk-reducing medication. The United States Preventive Services Task Force and the American Society of Clinical Oncology use 5-year risk threshold of 3%, based on evidence that women over this threshold are most likely to benefit from endocrine prevention therapy. Therefore, the reclassification for 5-year risk estimation compared Gail versus the BRISK model using both thresholds. Clinical guidelines for breast cancer screening recommend that lifetime risk estimates be assessed using models that are largely based upon family history, with women exceeding a risk threshold of 20% in the USA (or 25% in other countries) to be offered increased screening for breast cancer, therefore the reclassification for lifetime risk estimation compared IBIS versus the BRISK model. We categorized the estimated 5-year risk using categories (< 1%, ≥ 1 to < 1.67%, ≥ 1.67% to < 3%, and ≥ 3%), and used the net reclassification index (NRI) from Pencina and Steyerberg [38] to cross-classify the risk score distribution of alternative risk models, separately for cases and controls [39]. A similar approach was used to cross-classify estimated remaining lifetime risk using the categories (< 6%, ≥ 6% to < 12%, ≥ 12% to < 20%, ≥ 20% to < 25%, and ≥ 25%). NRI analyses for cases were performed for overall breast cancer as well as by ER and PR subtype (ER positive and PR positive; ER negative and PR negative) and stage (stage 1; stage ≥ 2).
Results
Participants
The risk factors for cases and controls are summarized in Table 1 and are in broad agreement with previously published data [40, 41].
Association and discrimination
The OR per SD and AUC for each of the models are presented in Table 2. Discrimination was improved for BRISK 5-year risk over Gail 5-year risk (χ2 = 88.29, degrees of freedom [d.f.] = 1, P < 0.0001) and for BRISK remaining lifetime risk over IBIS remaining lifetime risk (χ2 = 22.91, d.f. = 1, P < 0.0001). There was no difference in AUC between the simplified Rosner 10-year risk and either BRISK 5-year risk (χ2 = 3.69, d.f. = 1, P = 0.06) or BRISK remaining lifetime risk (χ2 = 1.18, d.f. = 1, P = 0.3). Similar results were obtained for the alternate specification of BRISK that uses approximated BI-RADS categories for the mammographic density measure (see Supplementary Table 1).
Model calibration
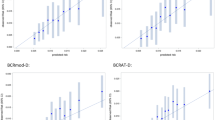
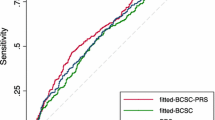
BRISK was very well calibrated for the estimation of both 5-year risk (E/O = 1.03; 95% CI 0.73, 1.46) and remaining lifetime risk (E/O = 1.01; 95% CI 0.86, 1.17). The Gail 5-year risk (E/O = 0.85; 95% CI 0.58, 1.24) and IBIS remaining lifetime risk estimates (E/O = 0.73; 95% CI 0.60, 0.87) were not well calibrated in this dataset, with both significantly under-estimating risk (Table 3). The distributions of cases and controls according to the 5-year or lifetime risk estimates provided by the relevant model are presented in Fig. 1, demonstrating separation of controls (orange) to the left and cases (blue) to the right. The data illustrate the assignment of controls to lower risk and the concomitant assignment of cases to higher risk by the BRISK model compared to Gail and IBIS.
Classification performance
5-year risk
We used reclassification tables to assess the assignment of women to the following 5-year risk categories: < 1%, > 1 to < 1.67%, > 1.67 to < 3% and > 3%, which cover the two most widely used thresholds for offering chemopreventive medication. For 5-year risk, BRISK improved classification performance for cases and controls over Gail 5-year risk with an NRI of 0.31 (standard error [SE] = 0.031). The reclassification improvement for cases was 0.415 (SE = 0.023) and − 0.103 (SE = 0.021) for controls (Table 4). The BRISK model assigned 72.2% of cases above the 1.67% National Comprehensive Cancer Network threshold for risk-reducing medication compared to 43.9% by Gail. At the United States Preventive Services Task Force threshold of 3%, the BRISK model assigned 51% of cases above the threshold, with only 8.75% being identified at this high-risk category by Gail (Supplementary Table 2).
The classification performance in cases was consistent for both ER positive cases (reclassification improvement = 0.44, SE 0.03) and ER negative cases (reclassification improvement = 0.31, SE 0.07) (Tables 5 and 6). The BRISK model assigned 53% of ER positive cases and 39.9% of ER negative cases above the 3% 5-year risk threshold.
Remaining lifetime risk
We also used reclassification tables to assess the assignment of women to the following lifetime risk categories: < 6%, > 6 to < 12%, > 12 to < 20%, > 20 to < 25% and > 25%, which cover the two most widely used thresholds for offering increased screening by MRI. BRISK remaining lifetime risk improved classification performance for cases and controls over IBIS remaining lifetime risk with an overall NRI of 0.287 (SE = 0.035). The reclassification improvement for cases was 0.381 (SE = 0.026) and − 0.094 (SE = 0.022) for controls (Table 7). The BRISK model assigned 41.3% of cases above the 20% National Comprehensive Cancer Network lifetime risk threshold for offering MRI screening compared to just 4.1% by IBIS. At the higher threshold of 25%, the BRISK model assigned 33.5% of cases above the threshold, with only 0.9% being identified at this high-risk category by IBIS (Supplementary Table 3). The improved classification performance in cases was consistent for both ER positive cases (reclassification improvement = 0.425; SE = 0.032) and ER negative cases (reclassification improvement = 0.34; SE = 0.079; Tables 8 and 9). The reclassification of lifetime risk is also favorable when analyzed by stage at diagnosis with the reclassification improvement for stage 1 cases = 0.397 (SE = 0.036) and for later stages (2 +) reclassification improvement = 0.423 (SE = 0.050; Tables 10 and 11).
Discussion
Many models have been developed to estimate a women’s risk of developing breast cancer; these can be summarized as simple clinical models such as the Gail model [31], which was designed for the general population, comprehensive models such as simplified Rosner [30], or complex pedigree-based models designed for use in a familial genetics setting such as IBIS [13]. In this paper, we have assessed the performance of a new risk model, BRISK, which incorporates family history, body mass index, menopausal status, polygenic risk and mammographic density in a format that does not require a complex pedigree assessment or multi-page questionnaire. The DNA required for genotyping can readily be obtained by buccal or saliva sample. This simple design specification is important in a general practice or mammography clinic setting where patient contact time is necessarily short, but where a risk assessment is useful and actionable. This concept is supported by developments in colorectal cancer risk assessment where reducing the questionnaire complexity has also been shown to not adversely affect model performance, but does increase ease of use [42,43,44,45].
We used a nested case–control dataset from the Nurses’ Health Study to compare the model’s performance to existing models. Our results are consistent with studies that have investigated the association of mammographic density and PRS with breast cancer risk [46,47,48], including those that have incorporated a risk prediction model [49, 50]. We used the classic version of IBIS (without SNPs or mammographic density) because this was the only version available in the Nurses’ Health Study. We aim to compare BRISK against IBIS version 8 at the earliest opportunity.
Overall, BRISK has superior discrimination and is better calibrated than both Gail (for 5-year risk) and IBIS (for remaining lifetimes risk), which are in widespread clinical usage (Tables 2, 3, 4). BRISK identifies 51% of cases and 32% of controls who are over the United States Preventive Services Task Force 5-year risk threshold where chemoprevention is recommended. BRISK also identifies 41% of cases and 22% of controls who are over the National Comprehensive Cancer Network 20% remaining lifetime risk threshold where increased screening would be offered.
Reclassification analysis of both 5-year and remaining lifetime risk suggested that the BRISK model improves classification for both ER positive and ER negative disease; however, the model still better predicts ER-positive disease. The SNPs in this model have ER-dependent ORs that, if applied, might further improve the tumor subtype prediction. We aim to further validate BRISK in this capacity because it could have even greater implications on clinician recommendation and patient uptake of risk-reducing medication and could enable future risk reduction options for those with ER-negative disease. Limitations to risk assessment remain despite improvements upon current standards shown within these data; the model is still a prediction. We still categorize a proportion of women within average risk categories (< 1.67%) who still go on to develop breast cancer, suggesting there remain unknown risk factors that can contribute to risk that models currently do not capture. However, the enhancement of current risk stratification is significant, and provides impactful clinical benefit.
When considering population level risk reduction measures, the proportion of the population identified as at-risk is reasonable. If used in clinical practice the model would identify between a third and a fifth of women aged 40–70 years as being at increased risk, either by 5-year risk (≥ 3.0%) or remaining lifetime risk (≥ 20%), respectively, meaning that we would subject approximately one-third of all women to increased surveillance or risk-reducing discussions with a clinician. This is a substantial increase over the current 2–10% identified by Gail or IBIS.
Risk-reducing medication is effective at reducing 38–65% of breast cancer incidence depending on the selective ER modulator or aromatase inhibitor clinical trial [51,52,53,54,55]. Despite having several risk-reducing medications that have been shown to reduce the risk of breast cancer, uptake has been dismal [56]. If risk-reducing medication recommendation could be improved through the use of a risk prediction model such as BRISK, a significant reduction in breast cancer incidence could be achieved.
Implications for screening modifications based on the BRISK model at 20% or 25% actionable thresholds for at-risk women are substantial. Based on this case–control study, we show that BRISK is capable of identifying equivalent proportions of stage 1 and stage 2 + breast cancer cases above an actionable risk threshold (Supplementary Table 3. These data suggest that if the stage 2 + women had been assessed with BRISK prior to diagnosis, they would have been identified as at-risk and provided additional screening based on their actionable risk level; they therefore could have been diagnosed with an earlier stage breast cancer.
Screening and risk-reducing options exist for breast cancer; however, implementation of the already accessible tools remains a challenge. Stratification of the general population is the first step towards enabling a more structured conversation on risk reductions for an at-risk woman, whether it be centered on lifestyle habits, surveillance or medication. Furthermore, inconsistent medical body recommendations on screening mammography and mammographic density notification can make it challenging for clinicians to make decisions, especially at a primary care level when they are required to know screening guidance for a wide variety of diseases. Enabling a basic conversation about breast cancer risk with the help of a simple risk model can support clinician use of risk assessment without the burden of specialized knowledge in the area.
In summary, we have developed an easy to implement risk model that enables clinicians to have increased visibility of at-risk patients compared with current clinical models, independent of the estrogen receptor status of future tumors or potential aggressive nature of the disease. This risk model was constructed with real-world implementation concerns in mind. It incorporates only the most impactful epidemiological factors. Focusing on a few risk factors that are simple to collect will save time and remove potential ambiguity due to incomplete questionnaires. Ultimately this improves risk stratification beyond current model performance without impeding on physician time management.
Data availability
The data generated in this study are not publicly available due to participant confidentiality and privacy concerns but are available upon request. Data access must be approved by the institutional review boards of the Brigham and Women’s Hospital and Harvard T.H. Chan School of Public Health. Further information including the procedures to obtain and access data from the Nurses’ Health Studies is described at: https://www.nurseshealthstudy.org/researchers.
References
Collaborative Group on Hormonal Factors in Breast Cancer (2001) Familial breast cancer: collaborative reanalysis of individual data from 52 epidemiological studies including 58,209 women with breast cancer and 101,986 women without the disease. Lancet 358:1389–1399
Cappello NM, Richetelli D, Lee CI (2019) The impact of breast density reporting laws on women’s awareness of density-associated risks and conversations regarding supplemental screening with providers. J Am Coll Radiol 16:139–146
Mavaddat N, Pharoah PD, Michailidou K, Tyrer J, Brook MN et al (2015) Prediction of breast cancer risk based on profiling with common genetic variants. J Natl Cancer Inst. https://doi.org/10.1093/jnci/djv036
Allman R, Dite GS, Hopper JL, Gordon O, Starlard-Davenport A et al (2015) SNPs and breast cancer risk prediction for African American and Hispanic women. Breast Cancer Res Treat 154:583–589
Dite GS, MacInnis RJ, Bickerstaffe A, Dowty JG, Allman R et al (2016) Breast cancer risk prediction using clinical models and 77 independent risk-associated SNPs for women aged under 50 years: Australian breast cancer family registry. Cancer Epidemiol Biomark Prev 25:359–365
Dite GS, Mahmoodi M, Bickerstaffe A, Hammet F, Macinnis RJ et al (2013) Using SNP genotypes to improve the discrimination of a simple breast cancer risk prediction model. Breast Cancer Res Treat 139:887–896
Ziogas A, Gildea M, Cohen P, Bringman D, Taylor TH et al (2000) Cancer risk estimates for family members of a population-based family registry for breast and ovarian cancer. Cancer Epidemiol Biomark Prev 9:103–111
Braithwaite D, Miglioretti DL, Zhu W, Demb J, Trentham-Dietz A et al (2018) Family history and breast cancer risk among older women in the breast cancer surveillance consortium cohort. JAMA Intern Med 178:494–501
Shiyanbola OO, Arao RF, Miglioretti DL, Sprague BL, Hampton JM et al (2017) Emerging trends in family history of breast cancer and associated risk. Cancer Epidemiol Biomark Prev 26:1753–1760
Brewer HR, Jones ME, Schoemaker MJ, Ashworth A, Swerdlow AJ (2017) Family history and risk of breast cancer: an analysis accounting for family structure. Breast Cancer Res Treat 165:193–200
Jacobi CE, Jonker MA, Nagelkerke NJ, van Houwelingen JC, de Bock GH (2003) Prevalence of family histories of breast cancer in the general population and the incidence of related seeking of health care. J Med Genet 40:e83
Antoniou AC, Cunningham AP, Peto J, Evans DG, Lalloo F et al (2008) The BOADICEA model of genetic susceptibility to breast and ovarian cancers: updates and extensions. Br J Cancer 98:1457–1466
Tyrer J, Duffy SW, Cuzick J (2004) A breast cancer prediction model incorporating familial and personal risk factors. Stat Med 23:1111–1130
Berry DA, Parmigiani G, Sanchez J, Schildkraut J, Winer E (1997) Probability of carrying a mutation of breast-ovarian cancer gene BRCA1 based on family history. J Natl Cancer Inst 89:227–238
Parmigiani G, Berry D, Aguilar O (1998) Determining carrier probabilities for breast cancer-susceptibility genes BRCA1 and BRCA2. Am J Hum Genet 62:145–158
Wendt CMS (2019) Identifying breast cancer susceptibility genes—a review of the genetic background in familial breast cancer. Acta Oncol 58:135
Maxwell KN, Domchek SM, Nathanson KL, Robson ME (2016) Population frequency of germline BRCA1/2 mutations. J Clin Oncol 34:4183–4185
Kuchenbaecker KB, Hopper JL, Barnes DR, Phillips KA, Mooij TM et al (2017) Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. JAMA 317:2402–2416
van der Groep P, van der Wall E, van Diest PJ (2011) Pathology of hereditary breast cancer. Cell Oncol (Dordr) 34:71–88
Michailidou K, Hall P, Gonzalez-Neira A, Ghoussaini M, Dennis J et al (2013) Large-scale genotyping identifies 41 new loci associated with breast cancer risk. Nat Genet 45:353–361
Michailidou K, Beesley J, Lindstrom S, Canisius S, Dennis J et al (2015) Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat Genet 47:373–380
Michailidou K, Lindstrom S, Dennis J, Beesley J, Hui S et al (2017) Association analysis identifies 65 new breast cancer risk loci. Nature 551:92–94
Easton DF, Pharoah PD, Antoniou AC, Tischkowitz M, Tavtigian SV et al (2015) Gene-panel sequencing and the prediction of breast-cancer risk. N Engl J Med 372:2243–2257
Mavaddat N, Michailidou K, Dennis J, Lush M, Fachal L et al (2019) Polygenic risk scores for prediction of breast cancer and breast cancer subtypes. Am J Hum Genet 104:21–34
Byng JW, Yaffe MJ, Jong RA, Shumak RS, Lockwood GA et al (1998) Analysis of mammographic density and breast cancer risk from digitized mammograms. Radiographics 18:1587–1598
Highnam RBM, Yaffe MJ et al (2010) Robust breast composition measurement-Volpara™. In: Marti JOA, Freixnet J, Marti R (eds) Digital mammography IWDM 2010 lecture notes in computer science. Springer, Berlin, pp 342–349
Sak MA, Littrup PJ, Duric N, Mullooly M, Sherman ME et al (2015) Current and future methods for measuring breast density: a brief comparative review. Breast Cancer Manag 4:209–221
Boyd NF, Dite GS, Stone J, Gunasekara A, English DR et al (2002) Heritability of mammographic density, a risk factor for breast cancer. N Engl J Med 347:886–894
Hopper JL, Dite GS, MacInnis RJ, Liao Y, Zeinomar N et al (2018) Age-specific breast cancer risk by body mass index and familial risk: prospective family study cohort (ProF-SC). Breast Cancer Res 20:132
Rosner B, Tamimi RM Kraft P, Gao C, Mu Y et al (2021) Simplified breast risk tool integrating questionnaire risk factors, mammographic density, and polygenic risk score: development and validation. Cancer Epidemiol Biomarkers Prev 30:600–607
Gail MH, Brinton LA, Byar DP, Corle DK, Green SB et al (1989) Projecting individualized probabilities of developing breast cancer for white females who are being examined annually. J Natl Cancer Inst 81:1879–1886
Mealiffe ME, Stokowski RP, Rhees BK, Prentice RL, Pettinger M et al (2010) Assessment of clinical validity of a breast cancer risk model combining genetic and clinical information. J Natl Cancer Inst 102:1618–1627
Dite GS, Jenkins MA, Southey MC, Hocking JS, Giles GG et al (2003) Familial risks, early-onset breast cancer, and BRCA1 and BRCA2 germline mutations. J Natl Cancer Inst 95:448–457
Odefrey F, Stone J, Gurrin LC, Byrnes GB, Apicella C et al (2010) Common genetic variants associated with breast cancer and mammographic density measures that predict disease. Cancer Res 70:1449–1458
Colditz GA, Hankinson SE (2005) The Nurses’ health study: lifestyle and health among women. Nat Rev Cancer 5:388–396
Hanley JA, McNeil BJ (1982) The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143:29–36
DeLong ER, DeLong DM, Clarke-Pearson DL (1988) Comparing the areas under two or more correlated receiver operating characteristic curves: a nonparametric approach. Biometrics 44:837–845
Pencina MJ, Steyerberg EW (2017) Net reclassification index at event rate: properties and relationships. Stat Med 36:4455–4467
Kerr KF, Wang Z, Janes H, McClelland RL, Psaty BM et al (2014) Net reclassification indices for evaluating risk prediction instruments: a critical review. Epidemiology 25:114–121
Gaudet MM, Gierach GL, Carter BD, Luo J, Milne RL (2018) Pooled analysis of nine cohorts reveals breast cancer risk factors by tumor molecular subtype. Cancer Res 78:6011–6021
Rojas K, Stuckey A (2016) Breast cancer epidemiology and risk factors. Clin Obstet Gynecol 59:651–672
Liu Y, Colditz GA (2018) Comparison of performance between a short categorized lifestyle exposure-based colon cancer risk prediction tool and a model using continuous measures. Cancer Prev Res 11:841–848
Rifas-Shiman SL, Willett WC, Lobb R, Kotch J, Dart C et al (2001) PrimeScreen, a brief dietary screening tool: reproducibility and comparability with both a longer food frequency questionnaire and biomarkers. Public Health Nutr 4:249–254
Usher-Smith JA, Walter FM, Emery JD, Win AK, Griffin SJ (2016) Risk prediction models for colorectal cancer: a systematic review. Cancer Prev Res (Phila) 9:13–26
Smith T, Muller DC, Moons KGM, Cross AJ, Johansson M et al (2019) Comparison of prognostic models to predict the occurrence of colorectal cancer in asymptomatic individuals: a systematic literature review and external validation in the EPIC and UK Biobank prospective cohort studies. Gut 68:672–683
Shieh Y, Hu D, Ma L, Huntsman S, Gard CC et al (2017) Joint relative risks for estrogen receptor-positive breast cancer from a clinical model, polygenic risk score, and sex hormones. Breast Cancer Res Treat 166:603–612
Vachon CM, Pankratz VS, Scott CG, Haeberle L, Ziv E et al (2015) The contributions of breast density and common genetic variation to breast cancer risk. J Natl Cancer Inst. https://doi.org/10.1093/jnci/dju397
Vachon CM, Scott CG, Tamimi RM, Thompson DJ, Fasching PA et al (2019) Joint association of mammographic density adjusted for age and body mass index and polygenic risk score with breast cancer risk. Breast Cancer Res 21:68
Zhang X, Rice M, Tworoger SS (2018) Addition of a polygenic risk score, mammographic density, and endogenous hormones to existing breast cancer risk prediction models: a nested case-control study. PLoS Med 15:e1002644
van Veen EM, Brentnall AR, Byers H, Harkness EF, Astley SM et al (2018) Use of single-nucleotide polymorphisms and mammographic density plus classic risk factors for breast cancer risk prediction. JAMA Oncol 4:476–482
Fisher B, Costantino JP, Wickerham DL, Cecchini RS, Cronin WM et al (2005) Tamoxifen for the prevention of breast cancer: current status of the national surgical adjuvant breast and bowel project P-1 study. J Natl Cancer Inst 97:1652–1662
Fisher B, Costantino JP, Wickerham DL, Redmond CK, Kavanah M et al (1998) Tamoxifen for prevention of breast cancer: report of the national surgical adjuvant breast and bowel project P-1 study. J Natl Cancer Inst 90:1371–1388
Vogel VG, Costantino JP, Wickerham DL, Cronin WM, Cecchini RS et al (2006) Effects of tamoxifen vs raloxifene on the risk of developing invasive breast cancer and other disease outcomes: the NSABP study of Tamoxifen and Raloxifene (STAR) P-2 trial. JAMA 295:2727–2741
Goss PE, Ingle JN, Alés-Martínez JE, Cheung AM, Chlebowski RT et al (2011) Exemestane for breast-cancer prevention in postmenopausal women. N Engl J Med 364:2381–2391
Cuzick J, Sestak I, Forbes JF, Dowsett M, Knox J et al (2014) Anastrozole for prevention of breast cancer in high-risk postmenopausal women (IBIS-II): an international, double-blind, randomised placebo-controlled trial. Lancet 383:1041–1048
Ball S, Arevalo M, Juarez E, Payne JD, Jones C (2019) Breast cancer chemoprevention: an update on current practice and opportunities for primary care physicians. Prev Med 129:105834
Acknowledgements
We wish to thank Mr Lawrence Whiting for his invaluable expertise in the management of the large data files from the UK Biobank.
Funding
Erika Spaeth is an employee of Phenogen Sciences Inc (a subsidiary of Genetic Technologies Limited). This study was fully funded by Genetic Technologies Limited, which had no role in the conceptualisation, design, data analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by GD, YM, and BR. The first draft of the manuscript was written by RA and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
Gillian Dite and Richard Allman are employees of Genetic Technologies Limited. Erika Spaeth is an employee of Phenogen Sciences Inc (a subsidiary of Genetic Technologies Limited). Aspects of this manuscript are covered by Australian Provisional Patent Application No: 2021903955, Breast Cancer Risk Assessment.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Allman, R., Mu, Y., Dite, G.S. et al. Validation of a breast cancer risk prediction model based on the key risk factors: family history, mammographic density and polygenic risk. Breast Cancer Res Treat 198, 335–347 (2023). https://doi.org/10.1007/s10549-022-06834-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-022-06834-7