Abstract
In recent years, iron sulfur (Fe–S) proteins have been identified as key players in mammalian metabolism, ranging from long-known roles in the respiratory complexes and the citric acid cycle, to more recently recognized roles in RNA and DNA metabolism. Fe–S cofactors have often been missed because of their intrinsic lability and oxygen sensitivity. More Fe–S proteins have now been identified owing to detection of their direct interactions with components of the Fe–S biogenesis machinery, and through use of informatics to detect a motif that binds the co-chaperone responsible for transferring nascent Fe–S clusters to domains of recipient proteins. Dissection of the molecular steps involved in Fe–S transfer to Fe–S proteins has revealed that direct and shielded transfer occurs through highly conserved pathways that operate in parallel in the mitochondrial matrix and in the cytosolic/nuclear compartments of eukaryotic cells. Because Fe–S clusters have the unusual ability to accept or donate single electrons in chemical reactions, their presence renders complex chemical reactions possible. In addition, Fe–S clusters may function as sensors that interconnect activity of metabolic pathways with cellular redox status. Presence in pathways that control growth and division may enable cells to regulate their growth according to sufficiency of energy stores represented by redox capacity, and oxidation of such proteins could diminish anabolic activities to give cells an opportunity to restore energy supplies. This review will discuss mechanisms of Fe–S biogenesis and delivery, and methods that will likely reveal important roles of Fe–S proteins in proteins not yet recognized as Fe–S proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Mammalian Fe–S proteins were largely overlooked until recent years
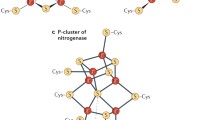
Fe–S proteins were not discovered and characterized until the late 1950s and early 1960s when methods were developed that detected their intrinsic magnetic properties, including electron paramagnetic imaging and Mossbauer spectroscopy methods (reviewed in Beinert et al. 1997). Fe–S clusters are composed of inorganic iron and sulfide atoms that assemble in different stoichiometries, including those composed of two iron and two sulfur atoms, and the common cubane cluster composed of four iron and four sulfide atoms; Fe–S clusters generally ligate to proteins through iron–cysteinyl bonds. Much more complex clusters are required for the functions of nitrogenase, which is key to nitrogen fixation, and hydrogenase, which releases energy upon oxidation of hydrogen. The fact that the characteristic absorbance of Fe–S clusters occurs in a portion of the uv visible spectrum where many other materials absorb permitted them to remain relatively unnoticed long after other key cofactors such as heme were identified and studied (Beinert 2000). Other factors that contributed to their obscurity include that they are readily disassembled by exposure to oxygen and other oxidants, and they are often degraded by the time a protein of interest has been isolated for further characterization. (Fig. 1)
Discovery of the Fe–S cluster of cytosolic aconitase and its role in regulation of cytosolic iron metabolism
Before 1990, very few Fe–S proteins had been identified and studied in mammalian cells, though many more had been discovered in bacteria (Johnson et al. 2005). They included two enzymes of the citric acid cycle, succinate dehydrogenase, and mitochondrial aconitase, which was extensively characterized by Beinert and colleagues in the 1980s (Kennedy et al. 1983; Beinert and Kennedy 1993). Complex I was also known to contain multiple deeply buried Fe–S clusters, but it was otherwise widely assumed that Fe–S proteins had gradually been phased out of metabolism during evolution of higher life forms because of their intrinsic lability.
During studies of mammalian regulation of iron metabolism genes, my colleagues and I discovered that multiple transcripts of iron proteins were regulated by a system known as the IRE-IRP system, in which transcripts that contain an RNA stem-loop known as an iron-responsive element (IRE) bind cytosolic iron-sensing proteins now known as iron regulatory proteins 1 and 2 (IRP1 and IRP2) in iron-depleted cells (reviewed in Klausner et al. 1993). Upon affinity purification of IRP1 using a column to which IREs were bound, the protein now known as IRP1 was purified and cloned (Rouault et al. 1990) and sequence resemblance to mitochondrial aconitase, particularly in the active site residues, suggested that IRP1 represented a functional cytosolic aconitase that interconverted citrate and isocitrate (Kaptain et al. 1991), which was confirmed by direct purification of beef liver cytosolic aconitase (Kennedy et al. 1992). Multiple studies demonstrated that alternation of IRP1 between functioning as an RNA binding protein or as a cytosolic aconitase depended on whether an intact [4Fe–4S] cluster was ligated in the active site cleft where a single cluster iron ligated the enzymatic substrate for aconitase activity, or IRP1 was present as an apoprotein that was devoid of an Fe–S cluster. The Fe–S cluster was present in a solvent-exposed cleft of the enzyme and was experimentally shown to be highly vulnerable to degradation (Haile et al. 1992). Structures of the cytosolic aconitase (Dupuy et al. 2006) and apo-IRP1 bound to the IRE (Walden et al. 2006) revealed how the IRE achieves high affinity binding to apo-IRP1 by binding to sites rendered accessible by conformational changes that occur upon loss of binding of the Fe–S cluster to the active site cleft (Fig. 2). These studies then led to intense focus on identifying the mechanisms by which Fe–S clusters are assembled and delivered to recipient proteins in mammalian cells.

Figure from Rouault (2006)
IRP1 alternates between function as a cytosolic aconitase, which contains a [Fe4–S4] cluster in the active-site cleft, to an apoprotein form that lacks the cluster and binds to IRE stem-loop structures present in several iron transcripts. Upon binding, IRP1 represses translation of multiple transcripts that contain IREs near the 5′-end of the transcript, including Ferritin H and L transcripts, HIF2 α, erythrocytic ALAS, and the iron export protein, ferroportin. Conversely, IRP binding to mRNAs that contain IREs at the 3′-UTR protects transcripts from endonucleolytic degradation: the most well-known such transcript is transferrin receptor 1. Apo-IRP1 undergoes a large conformational change that creates a complex IRE-binding pocket, in which the bulge C of the IRE binds to a pocket in domain 4, and three residues of the loop make finger-like binding projections into regions of domain 3 that become accessible after conformational change. The length of the upper base-paired stem of the IRE derived from NMR structural solution (Addess et al. 1997) optimizes the distance between the main IRP contact points, the unpaired C of the stem, and residues A15, G16 and U17 of the loop, resulting in high affinity binding
Overlapping and distinct roles of Irp1 and Irp2 were revealed by genetic studies
To dissect the physiological roles of the two IRPs, which represented a duplicated gene pair in which both are apparently expressed in differing ratios in all cell types, genetic knockouts of each IRP were generated in mice and analyzed. Deletion of both IRPs was embryonic lethal at the blastocyst stage of development (Smith et al. 2006), emphasizing the indispensable physiological need for Irp function at very early stages of development. Irp2−/− mice displayed neurodegeneration which most adversely affected motor neurons (Jeong et al. 2011) and caused axonopathy and degeneration in multiple other brain regions (LaVaute et al. 2001). In addition, animals developed a mild anemia and erythropoietic protoporphyria (Cooperman et al. 2005; Galy et al. 2005) attributable to functional iron deficiency from loss of TfR1 mediated iron uptake and overexpression of the multimeric cytosolic iron sequestration protein, ferritin. The clinical presentation of neurodegenerative disease was subtle, and mice often lived for 12 months or more. A dispute about the phenotype (Galy et al. 2006) was resolved by generation of a third knockout mouse (Zumbrennen-Bullough et al. 2014) which also manifested iron misregulation and impaired motor functions (summarized in Ghosh et al. 2015).
The clinical presentation of the Irp1−/− mice differed, perhaps because Irp1 was the predominant Irp in many non-CNS tissues including fat and kidneys (Meyron-Holtz et al. 2004a). Irp1−/− mice developed polycythemia (Ghosh et al. 2013; Anderson et al. 2013) and died suddenly on low iron diets, likely from hemorrhages related to occlusive clot formation caused by high hematocrits, which were even higher on the low iron diet (Ghosh et al. 2013). The polycythemia was driven by de-repression of HIF2 α, which is encoded by an IRE-containing transcript (Sanchez et al. 2007) that is primarily regulated by Irp1, likely because high amounts of Irp1 are expressed in erythropoietin-producing cells (Ghosh et al. 2013). HIF2 α drives expression of erythropoietin, which leads to excess red cell production, and also to pulmonary hypertension (Ghosh et al. 2013). Recent studies have shown that a stable nitroxide fed to animals, TEMPOL, can suppress the polycythemia of Irp1−/− mice, and also has a salutary effect on mice that have a mutation that interferes with function of the VHL degradation complex responsible for HIF degradation (Ghosh et al. 2018).
Experimental studies showed that IRPs faithfully reproduce their physiological activities when cells are cultured at low oxygen concentrations comparable to those that prevail in animal tissues (3–10%), rather than at room air concentrations of 21%, which constitutes hyperoxia and may adversely affect generation of meaningful experimental results in many settings (Meyron-Holtz et al. 2004b).
How are Fe–S clusters synthesized and ligated to recipient proteins in eukaryotic cells?
The discovery of bacterial operons that contained systems involved in synthesis of Fe–S clusters discovered in Azotobacter vinelandii and E. coli revolutionized studies of Fe–S biogenesis in cells. Bacterial operons generally contained the basic machinery for Fe–S cluster formation, consisting of a cysteine desulfurase that liberates sulfur for incorporation into nascent clusters (NifS or IscS), a scaffold protein upon which nascent clusters assemble (NifU or IscU), a ferredoxin which likely provides electrons for Fe–S cluster formation, and a chaperone-co-chaperone pair (HscA and B) that likely aid folding or cluster transfer (reviewed in Johnson et al. 2005). These genes are so highly conserved throughout evolution that counterparts were readily identified through homology searches in plants, fungi, and mammals (reviewed in Lill 2009; Rouault 2015a; Uzarska et al. 2018).
Based on studies performed in S. cerevisiae, it was proposed that all nascent Fe–S clusters were synthesized in the mitochondrial matrix, and a sulfur component was exported to cytosol by the ABC transporter, Atm1 (homologous to ABCB7 in mammals), whereupon specialized proteins in the cytosol, named the cytosolic iron sulfur assembly machinery or CIA machinery completed synthesis and delivery of Fe–S cofactors to numerous cytosolic and nuclear proteins (Lill et al. 2012), which include DNA metabolism proteins (Gari et al. 2012; Stehling et al. 2012), proteins involved in RNA metabolism (Barthelme et al. 2007), proteins involved in cytokinesis (Ben-Shimon et al. 2018) and in multiple other basic cellular processes.
The key role of Atm1 (ABCB7 in mammals) was postulated because it was alleged to be crucial for Fe–S acquisition of enzymatic function in a critical Fe–S enzyme of the cytosolic leucine biosynthetic pathway of yeast (Kispal et al. 1999), though this conclusion appears to be incorrect because a defect in the experimental design led to increased transcription of the entire leucine biosynthetic operon in cells that contained functional Atm1, because selective inactivation of Atm1 was achieved by using a plasmid that re-introduced functional Leu2 into a Leu2-null strain (reviewed in Rouault 2015a), and the comparison of leu 1 enzymatic activity that was performed on cells that contained inactivated Atm1 had markedly lower transcript levels of leu1, which would be expected to markedly decrease leu 1 protein levels and enzymatic activity in the cell line in which Atm1 function was knocked out using a construct that restored leu2 gene expression to the leu2 deficient strain. This flawed experimental design would make it unlikely that than the previously reported comparison of leu 1 enzymatic activity (Kispal et al. 1999) was made between cells with equal leu1 proteins levels (see supplementary info in Hausmann et al. (2008) for transcriptional data), and therefore does not support that Atm1 is needed for leu1 to acquire a cytosolic Fe–S cluster. Other supporting evidence that initial Fe–S formation occurred solely in mitochondria was that mitochondrial Fe–S proteins were described as fully functional in the Atm1 knockout strains, whereas cytosolic Fe–S proteins were alleged to not function if a critical Fe–S biogenesis component was not delivered by Atm1 (Kispal et al. 1999). However, mitochondrial Fe–S proteins were later judged to be more lacking in normal Fe–S cofactors than cytosolic proteins in the same yeast strains when rigorous biophysical methodologies were used for analysis (Miao et al. 2009), contradicting the argument that Fe–S assembly in the mitochondrial matrix was normal. Further experimentation in mammalian cells supports that mitochondrial Fe–S assembly is significantly negatively impacted by loss of the mammalian Atm1 counterpart, ABCB7 (Kim et al. 2018), whereas cytosolic Fe–S proteins are minimally affected.
Dissecting the mechanism of Fe–S delivery to recipient proteins
An important breakthrough in general understanding occurred when the Fe–S protein, succinate dehydrogenase subunit B (SDHB) was discovered to bind directly to the cochaperone, HSC20, the mammalian orthologue of bacterial HscB in the Fe–S synthesis machinery. The yeast homologue of HSC20, Jac1, was already known to bind the yeast orthologues of the scaffold protein IscU, known as Isu1, and the orthologue of the mammalian chaperone HSPA9 (HscA in bacteria), Ssq1, from yeast studies (Andrew et al. 2006; Kampinga and Craig 2010). Dissection of the binding site of HSC20 to mammalian SDHB revealed that HSC20 contained a pocket-like domain that bound iterations of the motif, LYR, which was found at two different positions in the open reading frame of SDHB (Maio et al. 2014), in the SDHB assembly factor protein SDHAF1(Maio et al. 2016), in LYRM7, a protein involved in Fe–S acquisition of the single Fe–S cluster of respiratory complex III, and in numerous proteins of complex I. These discoveries led to the conclusion that a single adaptable cochaperone-scaffold complex delivers nascent iron-sulfur clusters to mammalian respiratory chain complexes I–III (Maio et al. 2017). The basic features of the LYR motif include an aliphatic residue, followed by a large aromatic (tyrosine or phenylalanine), followed by a positively charged arginine or lysine, which can be modeled to fit into a pocket of the solved structure of HSC20 (Maio and Rouault 2016; Bitto et al. 2008). Thus, the basic apparatus of the Fe–S biogenesis machinery has been defined for mitochondrial Fe–S biogenesis, though the roles of other involved proteins such as glutaredoxin 5 (GLRX5) (Ye et al. 2010), ISCA, a putative scaffold protein (Py et al. 2018) and multiple other proteins such as NFU (Tong et al. 2003) have not been specifically assigned, and may be involved as secondary scaffold carriers that deliver clusters to a subset of Fe–S proteins.
Parallel systems for Fe–S synthesis in mitochondrial matrix and cytosolic/nuclear compartments of mammalian cells
In mammalian cells, a full complement of Fe–S biogenesis proteins is synthesized and targeted to the mitochondrial matrix, but almost all of these proteins have also been observed in the cytosolic/nuclear compartments of cells. Though they are often transcribed from the same gene, cytosolic forms are generated by alternative splicing (Tong and Rouault 2006), a weak mitochondrial targeting signal (Maio et al. 2014), alternative utilization of initiator AUGs that retained or skipped mitochondrial targeting information (Land and Rouault 1998) and possibly other mechanisms. These proteins have not been found in the cytosol of the model system S. cerevisiae, although the human cysteine desulfurase is clearly present and functional (Marelja et al. 2008, 2013). Unlike in bacterial systems, the cysteine desulfurase must be bound to another protein, ISD11 (also known as LYRM4) for function (Adam et al. 2006; Wiedemann et al. 2006; Shi et al. 2009).
Complexes involved in synthesis of Fe–S clusters have been defined by native gels and mass spectrometry in both mammalian mitochondria and in the cytosolic/nuclear compartment
To better understand the process of Fe–S assembly, native gels and mass spectrometry have been helpful in defining stages in the assembly process. The validity of the approach was established by discovering that SDHB acquired its Fe–S clusters in stages, beginning with binding to HSC20 and ISCU, and progressing to ejection of the HSC20 transfer complex, transfer of the cluster to SDHB, binding of holo-SDHB to SDHA in the multiprotein SDHB complex, and finally to the membrane bound subunits, SDHC and SDHD (Maio et al. 2014). Unlike proposals for plants and yeast, the HSC20 transfer complex was shown to be required for Fe–S acquisition in complex I, which contains eight Fe–S clusters, and also in complex III, which contains a single [2Fe–2S] cluster (Maio et al. 2017). The universality of function suggested that each recipient protein possessed the requisite information that would allow the transfer apparatus to bind and transfer its cargo directly to the recipient proteins; these findings supported that the general underlying principles of Fe–S acquisition by recipient protein were shared. In some proteins such as lipoic acid synthase, which requires two Fe–S clusters, it is not yet clear exactly how other potential scaffolds participate in cluster transfer. They may mediate transfer as secondary scaffolds that contain targeting information for a specific subset and type of recipient proteins, such as those that contain Fe–S proteins that are consumed during incorporation of sulfur into octanoic acid to generate lipoic acid (6,8-dithiooctanoic acid), which functions in the swinging arm of multi- subunit dehydrogenases such as α-keto acid dehydrogenase, pyruvate dehydrogenase, branched chain ketoacid dehydrogenase and the glycine cleavage system (McCarthy and Booker 2017) (reviewed in Solmonson and DeBerardinis 2018) for which the exact molecular donor of the cluster is under study (McCarthy et al. 2018). It is likely that multiple secondary scaffolds may acquire their Fe–S cofactors from the canonical Fe–S biogenesis machinery, and may then distribute their bound Fe–S to specific subsets of proteins based on specific protein recognition sites in recipient proteins that promote complex formation and enshrouded transfer of Fe–S clusters from the secondary scaffold. An important role of NFU1 in lipoyl synthase cluster delivery was revealed in studies of human patients with Multiple Mitochondrial Dysfunctions Syndrome 1 (Navarro-Sastre et al. 2011; Cameron et al. 2011). The Cameron et al. study also revealed a critical role for BOLA3, a mitochondrial protein for which the role of Fe–S assembly has not yet been clarified.
Native gels and mass spectrometry also greatly enhanced studies of Fe–S biogenesis in the cytosol, where individual assembly steps have been defined (Kim et al. 2018). Differences between Fe–S biogenesis in the mitochondrial matrix include that the biogenesis machinery of the cytosol associates with Glutaredoxin 3 and BolA2, and instead of associating with a ferredoxin, the nascent complex may obtain necessary electrons through the thioredoxin system (thioredoxin reductase was identified in the complex), and/or from CBR1, an NADPH-Dependent Carbonyl Reductase that can bind glutathione (Malátková et al. 2010).
Studies showed that a branch point was reached in Fe–S cytosolic delivery to Fe–S proteins. In some examples, such as NUBP1, no other steps were required to facilitate transfer from the nascent apparatus, composed of the cysteine desulfurase, primary scaffold, and a chaperone co-chaperone pair. Interestingly, frataxin, which plays a critical function in the mitochondrial Fe–S biogenesis complex (Rouault 2015a), was clearly identified in the cytosol through mass spectrometry, though endogenous frataxin had not previously been found in mammalian cytosol. Cytosolic HSC20 was shown to be required for acquisition of radiolabeled iron in the DNA metabolism protein, POLD1, and for activation of Dihydropyrimidine Dehydrogenase in thymine metabolism (Kim et al. 2018) in experiments in which either HSC20 or ISCU were rendered dysfunctional. In some experiments, HSC20 expression was knocked out, and in others, the cytosolic isoform of the main scaffold protein ISCU was mutagenized and overexpressed to generate a dominant negative protein that was unable to release its nascent radiolabeled Fe–S cluster to cytosolic targets. Notably, these findings are inconsistent with the notion that these cytosolic Fe–S proteins derive their Fe–S co-factors solely from a precursor in the mitochondrial matrix (Lill 2009), since an exclusively cytosolic form of IscU was used in these experiments. In some other cytosolic and nuclear Fe–S proteins, the nascent complex associated with the large protein, CIAO1 (Srinivasan et al. 2007), by binding through HSC20 to an LYR sequence in CIAO1 (Kim et al. 2018). Since HSC20 dimerizes, it could readily bind to CIAO1 through one monomer, and bind to other binding partners through the LYR of the second half of the dimer in a cytosolic Fe–S transfer complex composed of MMS19 and FAM96B (Stehling et al. 2012; Gari et al. 2012). The CIA complex also contains other sites that may bind recipient proteins (Odermatt and Gari 2017) and it is particularly important for Fe–S acquisition by DNA metabolism proteins. Its role in Fe–S acquisition of other cytosolic proteins is a subject of ongoing work. However, recent work strongly indicates the CIA pathway previously outlined (Lill 2009) that originates solely in mitochondria is not the sole source of Fe–S to cytosolic recipients (Fig. 3).
A summary of pathways of cytosolic Fe–S biogenesis. The nascent cluster assembles on the ISCU scaffold using cytosolic isoforms of biogenesis proteins, and interaction with the chaperone/co-chaperone apparatus permits these clusters to transfer directly to NARFL and the NUBP1/NUBP2 complex. A subset of Fe–S proteins involved in DNA metabolism and ribosome biogenesis, including DNA repair, RNA metabolism, and numerous other cytosolic proteins obtain their Fe–S cluster cofactors after interacting with the CIA complex, depicted as composed of CIAO1, MMS19, and FAM96B. Figure modified from Kim et al. (2018)
Implications for discovery of new Fe–S proteins
Discovery of the important role for LYR motifs opens the possibility that informatics can be used to identify prospective Fe–S proteins, and feasible follow-up steps including analysis by ICP-MS followed by scale up for analysis by definitive techniques that require large amounts of protein such as EPR and Mossbauer could revolutionize Fe–S protein discovery and lead to revision of many metabolic pathways (Rouault 2015a, b). The LYR motif has also been implicated as important in binding the acyl carrier protein (ACP) (Van Vranken et al. 2018) which may aid in coordinating respiration with fatty acid synthesis, though mutagenesis studies have not been performed to test the specific role of the LYR motifs in the larger binding site of ACP. The fact that numerous mutations of the LYR motif have been shown in experimental systems to prevent binding of the chaperone HSC20 (Maio et al. 2014, 2017; Maio and Rouault 2016) indicates that LYR motifs are involved in binding HSC20, but they may also bind ACP. These events need not be mutually exclusive, as the role of HSC20 binding to LYR could be crucial to Fe–S delivery, and after the Fe–S containing protein separates from the nascent Fe–S delivery machinery, it could bind a new binding partner such as ACP that possibly contributes to regulation of respiration. ACP was found in newer crystal structures of the nascent Fe–S complex (Cory et al. 2017; Boniecki et al. 2017), and its cytosolic ACP counterpart, fatty acid synthase (FASN), was identified by mass spectrometry in a cytosolic complex purified using antibodies to NFS1 in cytosolic fractions FASN (Kim et al. 2018).
Role of mutations of Fe–S biogenesis machinery in multiple rare diseases
In a fundamental process such as Fe–S biogenesis, it is not surprising that mutations to the proteins involved in the process cause human diseases, particularly in cases when loss of function is not complete. The most well-known of these diseases is Friedreich’s ataxia, which causes ataxia and cerebellar dysfunction due to a loss of expression of frataxin, which is caused by by expansion of a trinucleotide repeat in the first intron that represses expression by an unknown mechanism. These diseases may offer insights into the mechanisms that account for tissue-specific pathology in disease. The subject has been extensively reviewed (Rouault and Tong 2008; Rouault 2012; Stehling et al. 2014), and many insights and potential therapeutic strategies may arise from better understanding the pathogenesis of these diseases.
Future directions
It is likely that the role of Fe–S proteins will become increasingly recognized as key to metabolism and regulation of cellular responses as more key Fe–S proteins are identified and their roles are revealed. It is quite possible that Fe–S proteins represent a repository of energy stored in the form of reducing power, and that cells rely on this energy to maintain homeostasis. One could imagine that many pathways involved in cellular growth and regulation could be disrupted if a Fe–S cofactor becomes oxidized, leading to its dysfunction through changed conformation and/or charge interactions.
Iron sulfur cofactors evolved early in evolution, and the iron sulfur origin of life theory is a viable and attractive hypothesis (Boyd et al. 2017; Wächtershäuser 2007). By discovering direct protein- protein interactions between proteins in the Fe–S biogenesis pathways, it is likely that an elaborate distribution system for Fe–S cofactors will emerge that simultaneously ensures safe transfer and delivery to target proteins that need Fe–S cofactors for function, structure, or sensing. By following leads provided by direct interactions with proteins such as HSC20, NFU, ISCA, glutaredoxins and other proteins implicated in the process, the process will ultimately be sorted out, revealing exciting surprises and insights into growth of mammalian cells and tissues, and regulation of their functions.
References
Adam AC, Bornhovd C, Prokisch H, Neupert W, Hell K (2006) The Nfs1 interacting protein Isd11 has an essential role in Fe/S cluster biogenesis in mitochondria. EMBO J 25:174–183
Addess KJ, Basilion JP, Klausner RD, Rouault TA, Pardi A (1997) Structure and dynamics of the iron responsive element RNA: implications for binding of the RNA by iron regulatory binding proteins. J Mol Biol 274(1):72–83
Anderson SA, Nizzi CP, Chang YI, Deck KM, Schmidt PJ, Galy B, Damnernsawad A, Broman AT, Kendziorski C, Hentze MW, Fleming MD, Zhang J, Eisenstein RS (2013) The IRP1-HIF-2α axis coordinates iron and oxygen sensing with erythropoiesis and iron absorption. Cell Metab 17:282–290
Andrew AJ, Dutkiewicz R, Knieszner H, Craig EA, Marszalek J (2006) Characterization of the interaction between the J-protein Jac1p and the scaffold for Fe–S cluster biogenesis, Isu1p. J Biol Chem 281:14580–14587
Barthelme D, Scheele U, Dinkelaker S, Janoschka A, Macmillan F, Albers SV, Driessen AJ, Stagni MS, Bill E, Meyer-Klaucke W, Schunemann V, Tampe R (2007) Structural organization of essential iron-sulfur clusters in the evolutionarily highly conserved ATP-binding cassette protein ABCE1. J Biol Chem 282:14598–14607
Beinert H (2000) Iron-sulfur proteins: ancient structures, still full of surprises. J Biol Inorg Chem 5:2–15
Beinert H, Kennedy MC (1993) Aconitase, a two-faced protein: enzyme and iron regulatory factor. FASEB J 7:1442–1449
Beinert H, Holm RH, Munck E (1997) Iron-sulfur clusters: nature’s modular, multipurpose structures. Science 277:653–659
Ben-Shimon L, Paul VD, David-Kadoch G, Volpe M, Stümpfig M, Bill E, Mühlenhoff U, Lill R, Ben-Aroya S (2018) Fe–S cluster coordination of the chromokinesin KIF4A alters its subcellular localization during mitosis. J Cell Sci 131 (12):jcs211432
Bitto E, Bingman CA, Bittova L, Kondrashov DA, Bannen RM, Fox BG, Markley JL, Phillips GNJ (2008) Structure of human J-type co-chaperone HscB reveals a tetracysteine metal-binding domain. J Biol Chem 283:30184–30192
Boniecki MT, Freibert SA, Mühlenhoff U, Lill R, Cygler M (2017) Structure and functional dynamics of the mitochondrial Fe/S cluster synthesis complex. Nat Commun 8:1287
Boyd ES, Schut GJ, Shepard EM, Broderick JB, Adams MWW, Peters JW (2017) Origin and evolution of Fe–S proteins and enzymes. In: Rouault TA (ed) Iron sulfur clusters in chemistry and biology. deGruyter, Berlin, pp 445–461
Cameron JM, Janer A, Levandovskiy V, Mackay N, Rouault TA, Tong WH, Ogilvie I, Shoubridge EA, Robinson BH (2011) Mutations in iron-sulfur cluster scaffold genes NFU1 and BOLA3 cause a fatal deficiency of multiple respiratory chain and 2-oxoacid dehydrogenase enzymes. Am J Hum Genet 89:486–495
Cooperman SS, Meyron-Holtz EG, Olivierre-Wilson H, Ghosh MC, McConnell JP, Rouault TA (2005) Microcytic anemia, erythropoietic protoporphyria, and neurodegeneration in mice with targeted deletion of iron-regulatory protein 2. Blood 106:1084–1091
Cory SA, Van Vranken JG, Brignole EJ, Patra S, Winge DR, Drennan CL, Rutter J, Barondeau DP (2017) Structure of human Fe–S assembly subcomplex reveals unexpected cysteine desulfurase architecture and acyl-ACP-ISD11 interactions. Proc Natl Acad Sci USA 114:E5325–E5334
Dupuy J, Volbeda A, Carpentier P, Darnault C, Moulis JM, Fontecilla-Camps JC (2006) Crystal structure of human iron regulatory protein 1 as cytosolic aconitase. Structure 14:129–139
Galy B, Ferring D, Minana B, Bell O, Janser HG, Muckenthaler M, Schumann K, Hentze MW (2005) Altered body iron distribution and microcytosis in mice deficient in iron regulatory protein 2 (IRP2). Blood 106:2580–2589
Galy B, Hölter SM, Klopstock T, Ferring D, Becker L, Kaden S, Wurst W, Gröne HJ, Hentze MW (2006) Iron homeostasis in the brain: complete iron regulatory protein 2 deficiency without symptomatic neurodegeneration in the mouse. Nat Genet 38:967–969
Gari K, Leon Ortiz AM, Borel V, Flynn H, Skehel JM, Boulton SJ (2012) MMS19 links cytoplasmic iron-sulfur cluster assembly to DNA metabolism. Science 337:243–245
Ghosh MC, Zhang DL, Jeong SY, Kovtunovych G, Ollivierre-Wilson H, Noguchi A, Tu T, Senecal T, Robinson G, Crooks DR, Tong WH, Ramaswamy K, Singh A, Graham BB, Tuder RM, Yu ZX, Eckhaus M, Lee J, Springer DA, Rouault TA (2013) Deletion of iron regulatory protein 1 causes polycythemia and pulmonary hypertension in mice through translational derepression of HIF2 α. Cell Metab 17:271–281
Ghosh MC, Zhang DL, Rouault TA (2015) Iron misregulation and neurodegenerative disease in mouse models that lack iron regulatory proteins. Neurobiol Dis 81:66–75
Ghosh MC, Zhang DL, Ollivierre H, Eckhaus MA, Rouault TA (2018) Translational repression of HIF2α expression in mice with Chuvash polycythemia reverses polycythemia. J Clin Invest 128:1317–1325
Haile DJ, Rouault TA, Harford JB, Kennedy MC, Blondin GA, Beinert H, Klausner RD (1992) Cellular regulation of the iron-responsive element binding protein: disassembly of the cubane iron-sulfur cluster results in high-affinity RNA binding. Proc Natl Acad Sci USA 89:11735–11739
Hausmann A, Samans B, Lill R, Mühlenhoff U (2008) Cellular and mitochondrial remodeling upon defects in iron-sulfur protein biogenesis. J Biol Chem 283:8318–8330
Jeong SY, Crooks DR, Wilson-Ollivierre H, Ghosh MC, Sougrat R, Lee J, Cooperman S, Mitchell JB, Beaumont C, Rouault TA (2011) Iron insufficiency compromises motor neurons and their mitochondrial function in Irp2-null mice. PLoS ONE 6:e25404
Johnson DC, Dean DR, Smith AD, Johnson MK (2005) Structure, function, and formation of biological iron-sulfur clusters. Annu Rev Biochem 74:247–281
Kampinga HH, Craig EA (2010) The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat Rev Mol Cell Biol 11:579–592
Kaptain S, Downey WE, Tang C, Philpott C, Haile D, Orloff DG, Harford JB, Rouault TA, Klausner RD (1991) A regulated RNA binding protein also possesses aconitase activity. Proc Natl Acad Sci USA 88:10109–10113
Kennedy MC, Emptage MH, Dreyer JL, Beinert H (1983) The role of iron in the activation-inactivation of aconitase. J Biol Chem 258:11098–11105
Kennedy MC, Mende-Mueller L, Blondin GA, Beinert H (1992) Purification and characterization of cytosolic aconitase from beef liver and its relationship to the iron-responsive element binding protein. Proc Natl Acad Sci USA 89:11730–11734
Kim KS, Maio N, Singh A, Rouault TA (2018) Cytosolic HSC20 integrates de novo iron-sulfur cluster biogenesis with the CIAO1-mediated transfer to recipients. Hum Mol Genet 27:837–852
Kispal G, Csere P, Prohl C, Lill R (1999) The mitochondrial proteins Atm1p and Nfs1p are essential for biogenesis of cytosolic Fe/S proteins. EMBO J 18:3981–3989
Klausner RD, Rouault TA, Harford JB (1993) Regulating the fate of mRNA: the control of cellular iron metabolism. Cell 72:19–28
Land T, Rouault TA (1998) Targeting of a human iron-sulfur cluster assembly enzyme, nifs, to different subcellular compartments is regulated through alternative AUG utilization. Mol Cell 2:807–815
LaVaute T, Smith S, Cooperman S, Iwai K, Land W, Meyron-Holtz E, Drake SK, Miller G, Abu-Asab M, Tsokos M, Switzer R, Grinberg A, Love P, Tresser N, Rouault TA (2001) Targeted deletion of the gene encoding iron regulatory protein-2 causes misregulation of iron metabolism and neurodegenerative disease in mice. Nat Genet 27:209–214
Lill R (2009) Function and biogenesis of iron-sulphur proteins. Nature 460:831–838
Lill R, Hoffmann B, Molik S, Pierik AJ, Rietzschel N, Stehling O, Uzarska MA, Webert H, Wilbrecht C, Muhlenhoff U (2012) The role of mitochondria in cellular iron-sulfur protein biogenesis and iron metabolism. Biochim Biophys Acta 1823:1491–1508
Maio N, Rouault TA (2016) Mammalian Fe–S proteins: definition of a consensus motif recognized by the co-chaperone HSC20. Metallomics 8:1032–1046
Maio N, Singh A, Uhrigshardt H, Saxena N, Tong WH, Rouault TA (2014) Cochaperone binding to LYR motifs confers specificity of iron sulfur cluster delivery. Cell Metab 19:445–457
Maio N, Ghezzi D, Verrigni D, Rizza T, Bertini E, Martinelli D, Zeviani M, Singh A, Carrozzo R, Rouault TA (2016) Disease-causing SDHAF1 mutations impair transfer of Fe–S clusters to SDHB. Cell Metab 23:292–302
Maio N, Kim KS, Singh A, Rouault TA (2017) A single adaptable cochaperone-scaffold complex delivers nascent iron-sulfur clusters to mammalian respiratory chain complexes I–III. Cell Metab 25:945–953.e6
Malátková P, Maser E, Wsól V (2010) Human carbonyl reductases. Curr Drug Metab 11:639–658
Marelja Z, Stocklein W, Nimtz M, Leimkuhler S (2008) A novel role for human Nfs1 in the cytoplasm: Nfs1 acts as a sulfur donor for MOCS3, a protein involved in molybdenum cofactor biosynthesis. J Biol Chem 283:25178–25185
Marelja Z, Mullick Chowdhury M, Dosche C, Hille C, Baumann O, Lohmannsroben HG, Leimkuhler S (2013) The l-cysteine desulfurase NFS1 is localized in the cytosol where it provides the sulfur for molybdenum cofactor biosynthesis in humans. PLoS ONE 8:e60869
McCarthy EL, Booker SJ (2017) Destruction and reformation of an iron-sulfur cluster during catalysis by lipoyl synthase. Science 358:373–377
McCarthy EL, Rankin AN, Dill ZR, Booker SJ (2018) The A-type domain in Escherichia coli NfuA is required for regenerating the auxiliary [4Fe–4S] cluster in Escherichia coli lipoyl synthase. J Biol Chem 294:1609–1617
Meyron-Holtz EG, Ghosh MC, Iwai K, LaVaute T, Brazzolotto X, Berger UV, Land W, Ollivierre-Wilson H, Grinberg A, Love P, Rouault TA (2004a) Genetic ablations of iron regulatory proteins 1 and 2 reveal why iron regulatory protein 2 dominates iron homeostasis. EMBO J 23:386–395
Meyron-Holtz EG, Ghosh MC, Rouault TA (2004b) Mammalian tissue oxygen levels modulate iron-regulatory protein activities in vivo. Science 306:2087–2090
Miao R, Kim H, Koppolu UM, Ellis EA, Scott RA, Lindahl PA (2009) Biophysical characterization of the iron in mitochondria from Atm1p-depleted Saccharomyces cerevisiae. Biochemistry 48:9556–9568
Navarro-Sastre A, Tort F, Stehling O, Uzarska MA, Arranz JA, Del Toro M, Labayru MT, Landa J, Font A, Garcia-Villoria J, Merinero B, Ugarte M, Gutierrez-Solana LG, Campistol J, Garcia-Cazorla A, Vaquerizo J, Riudor E, Briones P, Elpeleg O, Ribes A, Lill R (2011) A fatal mitochondrial disease is associated with defective NFU1 function in the maturation of a subset of mitochondrial Fe–S proteins. Am J Hum Genet 89:656–667
Odermatt DC, Gari K (2017) The CIA targeting complex is highly regulated and provides two distinct binding sites for client iron-sulfur proteins. Cell Rep 18:1434–1443
Py B, Gerez C, Huguenot A, Vidaud C, Fontecave M, Ollagnier de Choudens S, Barras F (2018) The ErpA/NfuA complex builds an oxidative resistant Fe–S cluster delivery pathway. J Biol Chem 293:7689–7702
Rouault TA (2006) Biochemistry. If the RNA fits, use it. Science 314:1886–1887
Rouault TA (2012) Biogenesis of iron-sulfur clusters in mammalian cells: new insights and relevance to human disease. Dis Model Mech 5:155–164
Rouault TA (2015a) Mammalian iron-sulphur proteins: novel insights into biogenesis and function. Nat Rev Mol Cell Biol 16:45–55
Rouault TA (2015b) Iron-sulfur proteins hiding in plain sight. Nat Chem Biol 11:442–445
Rouault TA, Tong WH (2008) Iron-sulfur cluster biogenesis and human disease. Trends Genet 24:398–407
Rouault TA, Tang CK, Kaptain S, Burgess WH, Haile DJ, Samaniego F, McBride OW, Harford JB, Klausner RD (1990) Cloning of the cDNA encoding an RNA regulatory protein––the human iron-responsive element-binding protein. Proc Natl Acad Sci USA 87:7958–7962
Sanchez M, Galy B, Muckenthaler MU, Hentze MW (2007) Iron-regulatory proteins limit hypoxia-inducible factor-2α expression in iron deficiency. Nat Struct Mol Biol 14:420–426
Shi Y, Ghosh MC, Tong WH, Rouault TA (2009) Human ISD11 is essential for both iron-sulfur cluster assembly and maintenance of normal cellular iron homeostasis. Hum Mol Genet 18:3014–3025
Smith SR, Ghosh MC, Ollivierre-Wilson H, Hang Tong W, Rouault TA (2006) Complete loss of iron regulatory proteins 1 and 2 prevents viability of murine zygotes beyond the blastocyst stage of embryonic development. Blood Cells Mol Dis 36:283–287
Solmonson A, DeBerardinis RJ (2018) Lipoic acid metabolism and mitochondrial redox regulation. J Biol Chem 293:7522–7530
Srinivasan V, Netz DJ, Webert H, Mascarenhas J, Pierik AJ, Michel H, Lill R (2007) Structure of the yeast WD40 domain protein Cia1, a component acting late in iron-sulfur protein biogenesis. Structure 15:1246–1257
Stehling O, Vashisht AA, Mascarenhas J, Jonsson ZO, Sharma T, Netz DJ, Pierik AJ, Wohlschlegel JA, Lill R (2012) MMS19 assembles iron-sulfur proteins required for DNA metabolism and genomic integrity. Science 337:195–199
Stehling O, Wilbrecht C, Lill R (2014) Mitochondrial iron-sulfur protein biogenesis and human disease. Biochimie 100:61–77
Tong WH, Rouault TA (2006) Functions of mitochondrial ISCU and cytosolic ISCU in mammalian iron-sulfur cluster biogenesis and iron homeostasis. Cell Metab 3:199–210
Tong WH, Jameson GN, Huynh BH, Rouault TA (2003) Subcellular compartmentalization of human Nfu, an iron-sulfur cluster scaffold protein, and its ability to assemble a [4Fe-4S] cluster. Proc Natl Acad Sci USA 100:9762–9767
Uzarska MA, Przybyla-Toscano J, Spantgar F, Zannini F, Lill R, Mühlenhoff U, Rouhier N (2018) Conserved functions of Arabidopsis mitochondrial late-acting maturation factors in the trafficking of iron-sulfur clusters. Biochim Biophys Acta Mol Cell Res 1865:1250–1259
Van Vranken JG, Nowinski SM, Clowers KJ, Jeong MY, Ouyang Y, Berg JA, Gygi JP, Gygi SP, Winge DR, Rutter J (2018) ACP acylation is an acetyl-coA-dependent modification required for electron transport chain assembly. Mol Cell 71:567–580.e4
Wächtershäuser G (2007) On the chemistry and evolution of the pioneer organism. Chem Biodivers 4:584–602
Walden WE, Selezneva AI, Dupuy J, Volbeda A, Fontecilla-Camps JC, Theil EC, Volz K (2006) Structure of dual function iron regulatory protein 1 complexed with ferritin IRE-RNA. Science 314:1903–1908
Wiedemann N, Urzica E, Guiard B, Muller H, Lohaus C, Meyer HE, Ryan MT, Meisinger C, Muhlenhoff U, Lill R, Pfanner N (2006) Essential role of Isd11 in mitochondrial iron-sulfur cluster synthesis on Isu scaffold proteins. EMBO J 25:184–195
Ye H, Jeong SY, Ghosh MC, Kovtunovych G, Silvestri L, Ortillo D, Uchida N, Tisdale J, Camaschella C, Rouault TA (2010) Glutaredoxin 5 deficiency causes sideroblastic anemia by specifically impairing heme biosynthesis and depleting cytosolic iron in human erythroblasts. J Clin Invest 120:1749–1761
Zumbrennen-Bullough KB, Becker L, Garrett L, Hölter SM, Calzada-Wack J, Mossbrugger I, Quintanilla-Fend L, Racz I, Rathkolb B, Klopstock T, Wurst W, Zimmer A, Wolf E, Fuchs H, Gailus-Durner V, de Angelis MH, Romney SJ, Leibold EA (2014) Abnormal brain iron metabolism in Irp2 deficient mice is associated with mild neurological and behavioral impairments. PLoS ONE 9:e98072
Acknowledgements
I wish to thank members of the Rouault lab for their important contributions to insights on mammalian Fe–S biogenesis, and Nunziata Maio for critically reading this manuscript and making helpful suggestions. I gratefully acknowledge that this work was supported by the intramural program of the Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, USA.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Rouault, T.A. The indispensable role of mammalian iron sulfur proteins in function and regulation of multiple diverse metabolic pathways. Biometals 32, 343–353 (2019). https://doi.org/10.1007/s10534-019-00191-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10534-019-00191-7