Abstract
Objectives
In this study, the gut microbiome of healthy adult honeybees, Apis cerana, was investigated by sequencing the V3 – V4 region in 16S rRNA gene using Illumina Miseq platform.
Results
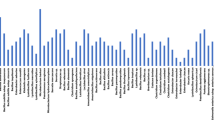
The total of 37,853 reads for 16S rRNA gene were obtained and 30,121 (79.6%) reads were valid with 25,291 (84.0%) reads that were classified into 116 species belonging to four major phyla. The relative abundances of the bacterial isolates in honeybee samples were phylum Proteobacteria (70.7%), Actinobacteria (10.7%), Firmicutes (10.3%), and Bacteroidetes (8.4%), respectively. Lactic acid bacteria comprised 18.95% with 10 groups including Bifidobacterium asteroides, B. indicum, Fructobacillus fructosus, Lactobacillus apinorum, L. apis, L. helsingborgensis, L. kimbladii, L. kullabergensis, and L. kunkeei.
Conclusions
The presence of beneficial bacteria in the gut highlighted their role in the honeybee and suggested that they can be promising candidates for the development of probiotics for health improvement, infection control and disease management of honeybees.



Similar content being viewed by others

Abbreviations
- AMPs:
-
Antimicrobial peptides
- DNA:
-
Deoxyribonucleic acid
- EDTA:
-
Ethylene diamine tetraacetic acid
- NGS:
-
Next generation sequencing
- OTU:
-
Operational taxonomic unit
- PCR:
-
Polymerase chain reaction
- QC:
-
Quality check
- rRNA:
-
Ribosomal ribonucleic acid
- SDS:
-
Sodium đoecyl sulfate
References
Ahn JH, Hong IP, Bok JI, Kim BY, Song J, Weon HY (2012) Pyrosequencing analysis of the bacterial communities in the guts of honey bees Apis cerana and Apis mellifera in Korea. J Microbiol 50:735–745
Alberoni D, Baffoni L, Gaggìa F, Ryan PM, Murphy K, Ross PR, Stanton C, Di Gioia D (2017) Impact of beneficial bacteria supplementation on the gut microbiota, colony development and productivity of Apis mellifera L. Benef Microbes. https://doi.org/10.3920/BM2017.0061
Al-Ghamdi A, Khan KA, Ansari MJ, Almasaudi SB, Al-Kahtani S (2018) Effect of gut bacterial isolates from Apis mellifera jemenitica on Paenibacillus larvae infected bee larvae. Saudi J Biol Sci 25:383–387
Al-Ghamdi A, Al-Abbadi AA, Khan KA, Ghramh HA, Ahmed AM, Ansari MJ (2020) In vitro antagonistic potential of gut bacteria isolated from indigenous honey bee race of Saudi Arabia against Paenibacillus larvae. J Apic Res. https://doi.org/10.1080/00218839.2019.1706912
Anderson KE, Sheehan TH, Eckholm BJ, Mott BM, DeGrandi-Hoffman G (2011) An emerging paradigm of colony health: microbial balance of the honey bee and hive (Apis mellifera). Insectes Soc 58:431–444
Anderson KE, Johansson A, Sheehan TH, Mott BM, Corby-Harris V, Johnstone L, Sprissler R, Fitz W (2013a) Draft genome sequences of two Bifidobacterium sp. from the honey bee (Apis mellifera). Gut Pathog 5:42
Anderson KE, Sheehan TH, Mott BM, Maes P, Snyder L, Schwan MR, Walton A, Jones BM, Corby-Harris V (2013b) Microbial ecology of the hive and pollination landscape: bacterial associates from floral nectar, the alimentary tract and stored food of honey bees (Apis mellifera). PLoS ONE 8(12):e83125
Anjum SI, Shah AH, Aurongzeb M, Kori J, Azim MK, Ansari MJ, Bin L (2018) Characterization of gut bacterial flora of Apis mellifera from north-west Pakistan. Saudi J Biol Sci 25:388–392
Babendreier D, Joller D, Romeis J, Bigler F, Widmer F (2007) Bacterial community structures in honeybee intestines and their response to two insecticidal proteins. FEMS Microbiol Ecol 59:600–610
Billiet A, Meeus I, Cnockaert M, Vandamme P, Van Oystaeyen A, Wackers F, Smagghe G (2016) Effect of oral administration of lactic acid bacteria on colony performance and gut microbiota in indoor-reared bumblebees (Bombus terrestris). Apidologie 48(1):41–50
Billiet A, Meeus I, Van Nieuwerburgh F, Deforce D, Wäckers F, Smagghe G (2017) Colony contact contributes to the diversity of gut bacteria in bumblebees (Bombus terrestris). Insect Sci 24:270–277
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Brodschneider R, Crailsheim K (2010) Nutrition and health in honey bees. Apidol 41:278–294
Butler E, Oien RF, Lindholm C, Olofsson TC, Nilson B, Vásquez A (2014) A pilot study investigating lactic acid bacterial symbionts from the honey bee in inhibiting human chronic wound pathogens. Int Wound J. https://doi.org/10.1111/iwj.12360
Cariveau DP, Powell JE, Koch H, Winfree R, Moran NA (2014) Variation in gut microbial communities and its association with pathogen infection in wild bumble bees (Bombus). ISME J 8(12):2369–2379
Casteels P, Ampe C, Jacobs F, Vaek M, Tempst P (1989) Apidaecins: antimicrobial peptides from honeybees. EMBO J 8:2387–2391
Casteels P, Ampe C, Riviere L, Damme JV, Elicone C, Fleming M, Jacobs F, Tempst P (1990) Isolation and characterization of abaecin, a major antimicrobial peptide in the honeybee (Apis mellifera). Eur J Biochem 187:381–386
Cox-Foster DL, Conlan S, Holmes EC, Palacios G, Evans JD, Moran NA, Quan PL, Briese T, Hornig M, Geiser DM, Martinson V, vanEngelsdorp D, Kalkstein AL, Drysdale A, Hui J, Zhai J, Cui L, Hutchison SK, Simons JF, Egholm M, Pettis JS, Lipkin WI (2007) A metagenomic survey of microbes in honey bee colony collapse disorder. Science 318:283–287
Disayathanoowat T, Young JP, Helgason T, Chantawannakul P (2011) T-RFLP analysis of bacterial communities in the midguts of Apis mellifera and Apis cerana honey bees in Thailand. FEMS Microbiol Ecol 79(2):273–281
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol 7:e1002195
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Engel P, Martinson VG, Moran NA (2012) Functional diversity within the simple gut microbiota of the honey bee. PNAS 109(27):11002–11007
Engel P, Kwong WK, McFrederick Q, Anderson KE, Barribeau SM, Chandler JA, Cornman RS, Dainat J, de Miranda JR, Doublet V, Emery O, Evans JD, Farinelli L, Flenniken ML, Granberg F, Grasis JA, Gauthier L, Hayer J, Koch H, Kocher S, Martinson VG, Moran N, Munoz-Torres M, Newton I, Paxton RJ, Powell E, Sadd BM, Schmid-Hempel P, Schmid-Hempel R, Song SJ, Schwarz RS, vanEngelsdorp D, Dainat B (2016) The bee microbiome: impact on bee health and model for evolution and ecology of host-microbe interactions. mBio 7(2):e02164–e02115
Evans JD, Schwarz RS (2011) Bees brought to their knees: microbes affecting honey bee health. Trends Microbiol 19(2):614–620
Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28:3150–3152
Gündüz EA, Douglas AE (2009) Symbiotic bacteria enable insect to use a nutritionally inadequate diet. Proce Roy Soc London Biol Sci 7:987–991
Jiang WY, Geng LL, Dai PL, Lang ZH, Shu CL, Lin Y, Zhou T, Song FP, Zhang J (2013) The influence of Bt-transgenic maize pollen on the bacterial diversity in the midgut of Chinese honeybees. Apis Cerana Cerana JIA 12:474–482
Jeyaprakash A, Hoy MA, Allsopp MH (2003) Bacterial diversity in worker adults of Apis mellifera capensis and Apis mellifera scutellata (Insecta: Hymenoptera) assessed using 16S rRNA sequences. J Invertebr Pathol 84:96–103
Kakumanu ML, Reeves AM, Anderson TD, Rodrigues RR, Williams MA (2016) Honeybee gut microbiome is altered by in-hive pesticide exposures. Front Microbiol 7:1255. https://doi.org/10.3389/fmicb.2016.01255
Kamada N, Chen GY, Inohara N, Núñez G (2013) Control of pathogens and pathobionts by the gut microbiota. Nat Immunol 14(7):685–690
Khan KA, Ansari MJ, Al-Ghamdi A, Nuru A, Harakeh S, Iqbal J (2017) Investigation of gut microbial communities associated with indigenous honey bee (Apis mellifera jemenitica) from two different eco-regions of Saudi Arabia. Saudi J Biol Sci 24:1061–1068
Khan KA, Al-Ghamdi A, Ghramh HA, Ansari MJ, Ali H, Alamri SA, Al-Kahtani SN, Adgaba N, Qasim M, Hafeez M (2020) Structural diversity and functional variability of gut microbial communities associated with honey bees. Microb Pathog 138(2020):103793
Klaudiny J, Albert Š, Bachanová K, Kopernický J, Šimúth J (2005) Two structurally different defensin genes, one of them encoding a novel defensin isoform, are expressed in honeybee Apis mellifera. Insect Biochem Mol Biol 35:11–22
Koch H, Schmid-Hempel P (2011) Bacterial communities in central European bumblebees: low diversity and high specificity. Microb Ecol 62:121–133
Kwong WK, Engel P, Koch H, Moran NA (2014) Genomics and host specialization of honey bee and bumble bee gut symbionts. PNAS 111(31):11509–11514
Kwong WK, Mancenido AL, Moran NA (2017) Immune system stimulation by the native gut microbiota of honey bees. R Soc Open Sci 4:170003. https://doi.org/10.1098/rsos.170003
Lalhmangaihi R, Ghatak S, Laha R, Gurusubramanian G, Kumar NS (2014) Protocol for optimal quality and quantity pollen DNA isolation from honey samples. J Biomol Tech 25(4):92–95
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systermatics. Wiley, New York, pp 115–175
Lee B, Moon T, Yoon S, Weissman T (2017) DUDE-Seq: fast, flexible, and robust denoising for targeted amplicon sequencing. PLoS ONE 12(7):e0181463
Li J, Qin H, Wu J, Sadd BM, Wang X, Evans JD, Peng W, Chen Y (2012) The prevalence of parasites and pathogens in Asian honeybees Apis cerana in China. PLoS ONE 7:e47955
Lim HC, Chu CC, Seufferheld MJ, Cameron SA (2015) Deep sequencing and ecological characterization of gut microbial communities of diverse bumble bee species. PLoS ONE 10(3):e0118566
Ludvigsen J, Rangberg A, Avershina E, Sekelja M, Kreibich C, Amdam G, Rudi K (2015) Shifts in the midgut/pyloric microbiota composition within a honey bee apiary throughout a season. Microbes Environ 30(3):235–244
Martinson VG, Danforth BN, Minckley RL, Rueppell O, Tingek S, Moran NA (2011) A simple and distinctive microbiota associated with honey bees and bumble bees. Mol Ecol 20:619–628
Masella AP, Bartram AK, Truszkowski JM, Brown DG, Neufeld JD (2012) PANDAseq: paired-end assembler for illumina sequences. BMC Bioinform 13:31. https://doi.org/10.1186/1471-2105-13-31
Mattila HR, Rios D, Walker-Sperling VE, Roeselers G, Newton ILG (2012) Characterization of the active microbiotas associated with honey bees reveals healthier and broader communities when colonies are genetically diverse. PLoS One 7(3):e32962
Mohr KI, Tebbe CC (2006) Diversity and phylotype consistency of bacteria in the guts of three bee species (Apoidea) at an oilseed rape field. Environ Microbiol 8:258–272
Moran NA (2015) Genomics of the honeybee microbiome. Curr Opin Insect Sci 10:22–28
Moran NA, Hansen AK, Powell JE, Sabree ZL (2012) Distinctive gut microbiota of honey bees assessed using deep sampling from individual worker bees. PLoS One 7(4):e36393
Myers EW, Miller W (1988) Optimal alignments in linear space. CABIOS 4:11–17
Nonthapa P, Chanchao C (2015) Pathogen detection and gut bacteria identification in Apis cerana indica in Thailand. Afr J Biotechnol 14(49):3235–3247
Olofsson TC, Vásquez A (2008) Detection and identification of a novel lactic acid bacterial flora within the honey stomach of the honeybee Apis mellifera. Curr Microbiol 57:356–363
Piccart K, Vásquez A, Piepers S, De Vliegher S, Olofsson TC (2016) Short communication: lactic acid bacteria from the honeybee inhibit the in vitro growth of mastitis pathogens. J Dairy Sci 99:1–5
Raymann K, Moran NA (2018) The role of the gut microbiome in health and disease of adult honey bee workers. Curr Opin Insect Sci 26:97–104
Tarpy DR, Mattila HR, Newton ILG (2015) Development of the honey bee gut microbiome throughout the queen-rearing process. Appl Environ Microbiol 81:3182–3191
Vásquez A, Forsgren E, Fries I, Paxton RJ, Flaberg E, Szekely L, Olofsson TC (2012) Symbionts as major modulators of insect health: lactic acid bacteria and honeybees. PLoS ONE 7:e33188
Wu M, Sugimura Y, Taylor DM, Yoshiyama M (2013a) Honeybee gastrointestinal bacteria for novel and sustainable disease control strategies. JDSA 8:85–90
Wu M, Sugimura Y, Takaya N, Takamatsu D, Kobayashi M, Taylor D, Yoshiyama M (2013b) Characterization of bifidobacteria in the digestive tract of the Japanese honeybee, Apis cerana japonica. J Invertebr Pathol 112:88–93
Wu M, Sugimura Y, Iwata K, Takaya N, Takamatsu D, Kobayashi M, Taylor D, Kimura K, Yoshiyama M (2014) Inhibitory effect of gut bacteria from the Japanese honey bee, Apis cerana japonica, against Melissococcus plutonius, the causal agent of European foulbrood disease. J Insect Sci 14:129. https://doi.org/10.1093/jis/14.1.129
Yanez O, Gauthier L, Chantawannakul P, Neumann P (2016) Endosymbiotic bacteria in honey bees: Arsenophonus spp. are not transmitted transovarially. FEMS Microbiol Lett 363:fnw147
Yoshiyama M, Kimura K (2009) Bacteria in the gut of Japanese honeybee, Apis cerana japonica, and their antagonistic effect against Paenibacillus larvae, the causal agent of American foulbrood. J Invertebr Pathol 102:91–96
Acknowledgements
This research was supported by Vietnam National Foundation for Science and Technology Development (NAFOSTED) under grant number 106.04-2019.24, and by an international research fund (Project Code No. I-1543081-2017-18-0102) by Animal and Plant Quarantine Agency, South Korea. The authors would like to thank to Dr. Byung-Yong Kim for his invaluable scientific support.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Duong, B.T.T., Lien, N.T.K., Thu, H.T. et al. Investigation of the gut microbiome of Apis cerana honeybees from Vietnam. Biotechnol Lett 42, 2309–2317 (2020). https://doi.org/10.1007/s10529-020-02948-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02948-4