Abstract
Objective
Rapid and convenient detection of protein–protein interactions (PPIs) is of great significance for understanding function of protein.
Results
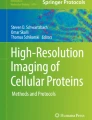
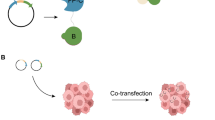
For efficiently detecting PPIs, we used the changes of proteins fluorescence localization to design a novel system, fluorescence translocation co-localization (FTCL), based on nuclear localization signal (NLS) in living cells. Depending on the original state of protein localization (both in the cytoplasm, both in the nucleus, one in the nucleus and another in the cytoplasm), two target proteins can be partitioned into the cytoplasm and nucleus by adding a NLS or mutating an existing NLS. Three independent results display that the changes of protein fluorescence co-localization were observed following co-expression of the two target proteins. At the same time, we verified the accuracy of fluorescence co-localization by co-immunoprecipitation.
Conclusions
There FTCL system provided a novel detection method for PPIs, regardless of protein localization in the nucleus or cytoplasm. More importantly, this study provides a new strategy for future protein interaction studies through organelle localization (such as mitochondria, Golgi and cytomembrane, etc.).





Similar content being viewed by others
References
Backes S et al (2018) Tom70 enhances mitochondrial preprotein import efficiency by binding to internal targeting sequences. J Cell Biol 217:1369–1382
Beaufour M, Godin F, Vallee B, Cadene M, Benedetti H (2012) Interaction proteomics suggests a new role for the Tfs1 protein in yeast. J Proteome Res 11:3211–3218
Braun P, Gingras AC (2012) History of protein-protein interactions: from egg-white to complex networks. Proteomics 12:1478–1498
Chen H, Puhl HL, Koushik SV, Vogel SS, Ikeda SR (2006) Measurement of FRET efficiency and ratio of donor to acceptor concentration in living cells. Biophys J 91:L39–L41
Clark NM et al. (2016) Tracking transcription factor mobility and interaction in Arabidopsis roots with fluorescence correlation spectroscopy. Elife 5
De Las RJ, Fontanillo C (2012) Protein-protein interaction networks: unraveling the wiring of molecular machines within the cell. Brief Funct Genom 11:489–496
Dinca AA, Chien WM, Chin MT (2018) Identification of novel mitochondrial localization signals in human Tafazzin, the cause of the inherited cardiomyopathic disorder Barth syndrome. J Mol Cell Cardiol 114:83–92
Dong ZQ et al. (2015) Oligomerization of baculovirus LEF-11 is involved in viral DNA replication. Plos One 10(12)
Fields S, Song OK (1989) A novel genetic system to detect protein protein interactions. Nature 340:245–246
Foltman M, Sanchez-Diaz A (2016) Studying protein-protein interactions in budding yeast using Co-immunoprecipitation methods. Mol Biol 1369:239–256
Freilich R, Arhar T, Abrams JL, Gestwicki JE (2018) Protein-protein interactions in the molecular chaperone network. Acc Chem Res 51:940–949
Giot L et al (2003) A protein interaction map of Drosophila melanogaster. Science 302:1727–1736
Guttman M, Betts GN, Barnes H, Ghassemian M, van der Geer P, Komives EA (2009) Interactions of the NPXY microdomains of the low density lipoprotein receptor-related protein 1. Proteomics 9:5016–5028
He JQ, Liu WD (2018) Identification of disrupted pathways associated with colon cancer based on combining protein-protein interactions and pathway data. J Cancer Res Therap 14:S998–S1003
Hu CD, Chinenov Y, Kerppola TK (2002) Visualization of interactions among bZip and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell 9:789–798
Iqbal H, Akins DR, Kenedy MR (2018) Co-immunoprecipitation for identifying protein-protein interactions in borrelia burgdorferi. Methods Mol Biol 1690:47–55
Li SM et al (2004) A map of the interactome network of the metazoan C-elegans. Science 303:540–543
Lin JS, Lai EM (2017) Protein-protein interactions: Co-immunoprecipitation methods. Mol Biol 1615:211–219
Long Y et al (2017) In vivo FRET-FLIM reveals cell-type-specific protein interactions in Arabidopsis roots. Nature 548:97–102
Mateus A, Kurzawa N, Becher I, Sridharan S, Helm D, Stein F, Typas A, Savitski MM (2020) Thermal proteome profiling for interrogating protein interactions. Mol Sys Biol 16(3):e9232
Miller JP, Lo RS, Ben-Hur A, Desmarais C, Stagljar I, Noble WS, Fields S (2005) Large-scale identification of yeast integral membrane protein interactions. Proc Natl Acad Sci USA 102:12123–12128
Murakami Y, Tripathi LP, Prathipati P, Mizuguchi K (2017) Network analysis and in silico prediction of protein-protein interactions with applications in drug discovery. Curr Opin Struct Biol 44:134–142
Pan MH, Cai XJ, Liu M, Lv J, Tang H, Tan J, Lu C (2010) Establishment and characterization of an ovarian cell line of the silkworm, Bombyx mori. Tissue Cell 42:42–46
Rual JF et al (2005) Towards a proteome-scale map of the human protein-protein interaction network. Nature 437:1173–1178
Sekar RB, Periasamy A (2003) Fluorescence resonance energy transfer (FRET) microscopy imaging of live cell protein localizations. J Cell Biol 160:629–633
Smith DB, Johnson KS (1988) Single-step purification of polypeptides expressed in escherichia-coli as fusions with glutathione S-transferase. Gene 67:31–40
Stelzl U et al (2005) A human protein-protein interaction network: a resource for annotating the proteome. Cell 122:957–968
Turriziani B, von Kriegsheim A, Pennington SR (2016) Protein-protein interaction detection via mass spectrometry-based proteomics modern proteomics—sample preparation. Anal Pract Appl 919:383–396
von Mering C, Krause R, Snel B, Cornell M, Oliver SG, Fields S, Bork P (2002) Comparative assessment of large-scale data sets of protein-protein interactions. Nature 417:399–403
Walhout AJ, Vidal M (2001a) High-throughput yeast two-hybrid assays for large-scale protein interaction mapping. Methods 24:297–306
Walhout AJM, Vidal M (2001b) Protein interaction maps for model organisms. Nat Rev Mol Cell Biol 2:55–62
Walker C, Bottger S, Low B (2006) Mortalin-based cytoplasmic sequestration of p53 in a nonmammalian cancer model. Am J Pathol 168:1526–1530
Wei Hu, Yuan Yi, Wang C-H, Tian H-T et al (2019) Genetically encoded residue-selective photo-crosslinker to capture protein-protein interactions in living cells. Chem 5:2955–2968
Yuekun Lang,Zhong Li,Hongmin Li (2019) Analysis of protein‐protein interactions by split luciferase complementation assay. Current Protocols in Toxicology, 82(1)
Zhang J et al. (2014) Identification of a novel nuclear localization signal of baculovirus late expression factor 11 Virus Research 184:111–119
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31872428 and 31902214), Natural Science Foundation of Chongqing (cstc2019jcyj-msxm2371) and the China Agriculture Research System (CARS-18).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Authors declare that they have no potential conflicts of interests.
Ethical approval
This article does not contain any experiments with human participants or animals (except invertebrate’s cell lines, which are exempt from ethical concerns) performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
10529_2020_2934_MOESM1_ESM.doc
Figure S1: The functional domain of LEF-11. Large rectangle shows the amino acid sequence of LEF-11 where gray areas represent functional motifs and black rectangles represent different LEF-11 forms. The nuclear localization signal is located at the C-terminal, and mutation of the 100th amino acid residue K influences LEF-11 nuclear localization. (DOC 75 kb)
Rights and permissions
About this article
Cite this article
Hu, N., Dong, ZQ., Chen, TT. et al. A novel system to rapidly detect protein–protein interactions (PPIs) based on fluorescence co-localization. Biotechnol Lett 42, 2111–2122 (2020). https://doi.org/10.1007/s10529-020-02934-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-020-02934-w