Abstract
Objectives
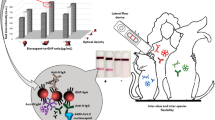
A Neissaria bacterial pilus sugar, bacillosamine, was synthesized and, for the first time, used as a probe to screen a single-chain variable fragment (scFv).
Results
Four Neisseria, Neisseria gonorrhoeae, Neisseria meningitidis, Neisseria sicca and Neisseria subflava, and two negative controls, Streptococcus pneumoniae and Escherichia coli, were tested through ELISA, immunostaining and gold nanoparticle immunological assay. All results indicated that the selected scFv is feasible for the specific detection of Neisseria species via the recognition of bacillosamine.
Conclusions
The recombinant scFv could detect Neisseria strains at 106 CFU/ml.




Similar content being viewed by others
References
Ahmad ZA, Yeap SK, Ali AM, Ho WY, Alitheen NB, Hamid M (2012) scFv antibody: principles and clinical application. Clin Dev Immunol 2012:980250
Amin MN, Ishiwata A, Ito Y (2006) Synthesis of asparagine-linked bacillosamine. Carbohydr Res 341:1922–1929
Aral SO, Mosher WD, Cates W (1991) Self-reported pelvic inflammatory disease in the United States, 1988. J Am Med Assoc 266:2570–2573
Carbonnelle E, Hill DJ, Morand P, Griffiths NJ, Bourdoulous S, Murillo I, Nassif X, Virji M (2009) Meningococcal interactions with the host. Vaccine 27:B78–B89
Hartley MD, Morrison MJ, Aas FE, Borud B, Koomey M, Imperiali B (2011) Biochemical characterization of the O-linked glycosylation pathway in Neisseria gonorrhoeae responsible for biosynthesis of protein glycans containing N,N′-diacetylbacillosamine. Biochemistry 50:4936–4948
Huttunen S, Toivanen M, Liu C, Tikkanen-Kaukanen C (2016) Novel anti-infective potential of salvianolic acid B against human serious pathogen Neisseria meningitidis. BMC Res Notes 9:25–30
Power PM, Ku SC, Rutter K, Warren MJ, Limnios EA, Tapsall JW, Jennings MP (2007) The phase-variable Allele of the pilus glycosylation Gene pglA is not strongly associated with strains of Neisseria gonorrhoeae isolated from patients with disseminated gonococcal infection. Infect Immun 75:3202–3204
Raag R, Whitlow M (1995) Single-chain Fvs. FASEB J 9:73–80
Sockolosky JT, Szoka FC (2013) Periplasmic production via the pET expression system of soluble, bioactive human growth hormone. Prot Expr Purif 87:129–135
Storhoff JJ, Elghanian R, Mucic RC, Mirkin CA, Letsinger RL (1998) One-pot colorimetric differentiation of polynucleotides with single base imperfections using gold nanoparticle probes. J Am Chem Soc 120:1959–1964
Acknowledgements
This work was financially supported by Grants from the Ministry of Science and Technology, Taiwan (MOST 105-2113-M-009-005), and the Center for interdisciplinary science, National Chiao Tung University.
Supporting information
Supplementary Methods—Compound 1 synthesis and BSA modification. Screening of compound 1-specific clones by monoclonal phage ELISA. Bacterial strain and culture conditions. Isolation of Neisseria pili. Characterization of purified recombinant anti-compound 1 scFv antibody. Immunofluorescence staining and confocal laser scanning microscopy. Nano-gold particle immunological assay. Supplementary Scheme 1—Reagents and conditions of compound 1 synthesis. Supplementary Scheme 2—Installation of compound 1 to BSA. Supplementary Fig. 1—Procedure used to select compound 1-specific recombinant scFv antibodies. Supplementary Fig. 2—Approach used to determine scFv binding activity to compound 1. Supplementary Fig. 3—Immunofluorescence of Neisseria species. (Incubation with fluorescein were considered as the background.)
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, CY., Weng, CC., Lin, CH. et al. Development of a novel engineered antibody targeting Neisseria species. Biotechnol Lett 39, 407–413 (2017). https://doi.org/10.1007/s10529-016-2258-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-016-2258-1