Abstract
Objectives
To clone and characterize caffeic acid 3-O-methyltransferase (LcCOMT) from the rhizome of Ligusticum chuanxiong, a traditional medicinal herb having a high content of ferulic acid.
Results
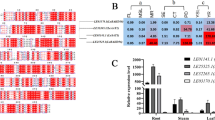
LcCOMT encoded an ORF of 362 amino acids with a calculated MW of 39,935 Da and pI of 5.94. Polygenetic tree indicated that LcCOMT was attributed to a new member of COMTs in plants. The recombinant LcCOMT was expressed in E. coli. HPLC and 1H NMR analyses of purified LcCOMT protein confirmed that it could catalyze caffeic acid to produce ferulic acid in vitro. The further site-mutagenesis proved that His268 was one key catalytic residue. In addition, the substantial changing expression level of LcCOMT under chilling treatment suggested that LcCOMT might play important role in the accumulation of ferulic acid under chilling treatment.
Conclusions
This is the first report of the isolation and characterization of a COMT clone from traditional medicine containing high contents of pharmaceutical ferulic acid.




Similar content being viewed by others
References
Bugos RC, Chiang VL, Campbell WH (1991) cDNA cloning, sequence analysis and seasonal expression of lignin-bispecific caffeic acid/5-hydroxyferulic acid O-methyltransferase of aspen. Plant Mol Biol 17:1203–1215
Gang DR, Lavid N, Zubieta C, Chen F, Beuerle T, Lewinsohn E, Noel JP, Pichersky E (2002) Characterization of phenylpropene O-methyltransferases from sweet basil: facile change of substrate specificity and convergent evolution within a plant O-methyltransferase family. Plant Cell 14:505–519
Hehmann M, Lukacin R, Ekiert H, Matern U (2004) Furanocoumarin biosynthesis in Ammi majus L. Cloning of bergaptol O-methyltransferase. Eur J Biochem 271:932–940
Li W, Tang Y, Chen Y, Duan JA (2012) Advances in the chemical analysis and biological activities of Chuanxiong. Molecules 17:10614–10651
Liu Z, Zhu Q, Li J, Zhang G, Jiamahate A, Zhou J, Liao H (2015a) Isolation, structure modeling and function characterization of a trypsin inhibitor from Cassia obtusifolia. Biotechnol Lett 37:863–869
Liu Z, Zhu Q, Li Y, Yu J, Wang W, Tan R, Zhou J, Liao H (2015b) Isolation and in silico characterization of a shikimate kinase from Cassia obtusifolia. Acta Physiol Plant 37:85
Morita Y, Saitoh M, Hoshino A, Nitasaka E, Iida S (2006) Isolation of cDNAs for R2R3-MYB, bHLH and WDR transcriptional regulators and identification of c and ca mutations conferring white flowers in the Japanese morning glory. Plant Cell Physiol 47:457–470
Song T, Liu ZB, Li JJ, Zhu QK, Tan R, Chen JS, Zhou JY, Liao H (2015) Comparative transcriptome of rhizome and leaf in Ligusticum chuanxiong. Plant Syst Evol. doi:10.1007/s00606-015-1211-4
Tao HM, Wang LS, Zhao DQ, Zhu QH, Liu YH (2011) Phenolic compounds from roots of Asparagus filicinus. Chin Tradit Herb Drugs 42:2181–2185 (in Chinese)
Trabucco GM, Matos DA, Lee SJ, Saathoff AJ, Priest HD, Mockler TC, Sarath G, Hazen SP (2013) Functional characterization of cinnamyl alcohol dehydrogenase and caffeic acid O-methyltransferase in Brachypodium distachyon. BMC Biotechnol 13:61
Tschaplinski TJ, Standaert RF, Engle NL, Martin MZ, Sangha AK, Parks JM, Smith JC, Samuel R, Jiang N, Pu Y, Ragauskas AJ, Hamilton CY, Fu C, Wang ZY, Davison BH, Dixon RA, Mielenz JR (2012) Down-regulation of the caffeic acid O-methyltransferase gene in switchgrass reveals a novel monolignol analog. Biotechnol Biofuels 5:71
Vanholme R, Morreel K, Darrah C, Oyarce P, Grabber JH, Ralph J, Boerjan W (2012) Metabolic engineering of novel lignin in biomass crops. N Phytol 196:978–1000
Vincent D, Lapierre C, Pollet B, Cornic G, Negroni L, Zivy M (2005) Water deficits affect caffeate O-methyltransferase, lignification, and related enzymes in maize leaves. Plant Physiol 137:949–960
Yoshihara N, Fukuchi-Mizutani M, Okuhara H, Tanaka Y, Yabuya T (2008) Molecular cloning and characterization of O-methyltransferases from the flower buds of Iris hollandica. J Plant Physiol 165:415–422
Zhou JM, Lee E, Kanapathy-Sinnaiaha F, Park Y, Kornblatt JA, Lim Y, Ibrahim RK (2010) Structure–function relationships of wheat flavone O-methyltransferase: homology modeling and site-directed mutagenesis. BMC Plant Biol 10:156
Zhu QK, Zhou JY, Zhang G, Liao H (2012) Homology modeling and molecular docking studies of Coptis chinensis (S)-scoulerine 9-O-methyltransferase. Chin J Chem 30:2533–2538
Zubieta C, Kota P, Ferrer JL, Dixon RA, Noel JP (2002) Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid 3/5-O methyltransferase. Plant Cell 14:1265–1277
Acknowledgments
This work was supported by the Grant (No. 31371232) from National Natural Science Foundation of China, Grant (2014ZX09304307001-019) from National Science and Technology Major Project and Grant (No. 2682014RC14) from Fundamental Research Funds for the Central Universities of China.
Supporting information
Supplementary Fig. 1—Sequence alignment of LcCOMT with other SAM-dependent OMT family members.
Supplementary Fig. 2—Phylogenetic tree of LcCOMT and other OMTs constructed by neighbor-joining algorithm.
Supplementary Fig. 3—1H NMR chemical shifts (ppm) for ferulic acid.
Supplementary Table 1—Comparison of the deduced amino acid sequences between LcCOMT and known SAM-dependent O-methyltransferases.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, JJ., Zhang, G., Yu, Jh. et al. Molecular cloning and characterization of caffeic acid 3-O-methyltransferase from the rhizome of Ligusticum chuanxiong . Biotechnol Lett 37, 2295–2302 (2015). https://doi.org/10.1007/s10529-015-1917-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1917-y