Abstract
Objectives
Genetic modifications to bacterial chromosomes are important for research; recently we reported a two-plasmid system for single locus modification in Escherichia coli and an improved method for simultaneous multiple-loci modification is needed.
Results
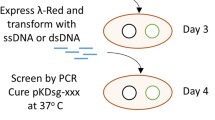
An intermediate bacterial strain was generated with different resistance marker genes flanked by I-SceI recognition sites at multiple target loci. Then a donor plasmid carrying several alleles with desired modifications was transformed into the intermediate strain together with a bifunctional helper plasmid encoding λ-Red recombinase and I-SceI endonuclease. I-SceI would induce double-strand breaks (DSBs) in the chromosome and λ-Red would induce recombination between chromosome DSBs and allele fragments from the donor plasmid, resulting in genomic modifications.
Conclusions
This method has been used to successfully perform three different loci modifications simultaneously.


Similar content being viewed by others
References
Blank K, Hensel M, Gerlach RG (2011) Rapid and highly efficient method for scarless mutagenesis within the Salmonella enterica chromosome. PLoS One 6:e15763
Datsenko KA, Wanner BL (2000) One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97:6640–6645
Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA 3rd, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6:343–345
Gust B, Challis GL, Fowler K, Kieser T, Chater KF (2003) PCR-targeted Streptomyces gene replacement identifies a protein domain needed for biosynthesis of the sesquiterpene soil odor geosmin. Proc Natl Acad Sci USA 100:1541–1546
Herring CD, Glasner JD, Blattner FR (2003) Gene replacement without selection: regulated suppression of amber mutations in Escherichia coli. Gene 311:153–163
Jiang Y, Chen B, Duan C, Sun B, Yang J, Yang S, Kelly RM (2015) Multigene editing in the Escherichia coli Genome via the CRISPR-Cas9 System. Appl Environ Microbiol 81:2506–2514
Kuhlman TE, Cox EC (2010) Site-specific chromosomal integration of large synthetic constructs. Nucleic Acids Res 38:e92
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25:1754–1760
Li H et al (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Lu D, Zhou X, Sun H, Song X, Chen H (2012) Efficient directional seamless DNA (deoxyribonucleic acid) segment connecting method CN patent. Patent 201210284031.
Marx CJ, Lidstrom ME (2002) Broad-host-range cre-lox system for antibiotic marker recycling in gram-negative bacteria. Biotechniques 33:1062–1067
Milne I, Bayer M, Cardle L, Shaw P, Stephen G, Wright F, Marshall D (2010) Tablet–next generation sequence assembly visualization. Bioinformatics 26:401–402
Sawitzke JA, Thomason LC, Costantino N, Bubunenko M, Datta S, Court DL (2007) Recombineering: in vivo genetic engineering in E. coli, S. enterica, and beyond. Methods Enzymol 421:171–199
Shao Z, Zhao H (2009) DNA assembler, an in vivo genetic method for rapid construction of biochemical pathways. Nucleic Acids Res 37:e16
Stoddard BL (2005) Homing endonuclease structure and function. Q Rev Biophys 38:49–95
Tischer BK, von Einem J, Kaufer B, Osterrieder N (2006) Two-step red-mediated recombination for versatile high-efficiency markerless DNA manipulation in Escherichia coli. Biotechniques 40:191–197
Tsuge K, Matsui K, Itaya M (2003) One step assembly of multiple DNA fragments with a designed order and orientation in Bacillus subtilis plasmid. Nucleic Acids Res 31:e133
Yang J et al (2014) High-efficiency scarless genetic modification in Escherichia coli using lambda-red recombination and I-Scei cleavage. Appl Environ Microbiol 80:3826–3834
Yu D, Ellis HM, Lee EC, Jenkins NA, Copeland NG, Court DL (2000) An efficient recombination system for chromosome engineering in Escherichia coli. Proc Natl Acad Sci USA 97:5978–5983
Yu BJ, Kang KH, Lee JH, Sung BH, Kim MS, Kim SC (2008) Rapid and efficient construction of markerless deletions in the Escherichia coli genome. Nucleic Acids Res 36:e84
Acknowledgments
We thank Dr. Jian Li for thoughtful discussions and reading of the manuscript. We thank Benjamin S. Glick (University of Chicago) for providing SnapGene software for producing plasmid maps. We thank Angel Yeast Co. Ltd for providing yeast extract for experiments.
This work was supported by the National Basic Research Program (973 Program) of China [2014CB745100, 2011CBA00800].
Conflict of interest
The authors have declared that no conflict of interests exist.
Supporting information
Supplementary Fig. S1: PCR analysis for cadA deletion, thrBC deletion and gdhA integration.
Supplementary Fig. S2: Lysine production in shake flask.
Supplementary Table S1: Oligonucleotide primers used in this study.
Supplementary Table S2: Plasmids used in this study.
Supplementary Methods and Protocols.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yang, J., Sun, B., Huang, H. et al. Multiple-site genetic modifications in Escherichia coli using lambda-Red recombination and I-SceI cleavage. Biotechnol Lett 37, 2011–2018 (2015). https://doi.org/10.1007/s10529-015-1878-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1878-1