Abstract
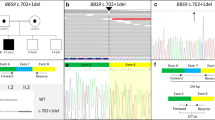
Bardet–Biedl syndrome (BBS) is a rare inherited ciliopathy disorder characterized by a broad spectrum of clinical symptoms such as retinal dystrophy, obesity, polydactyly, genitourinary and kidney anomalies, learning disability, and hypogonadism. The understanding of the variants involved in BBS-causing genes remains incomplete, highlighting the need for further research to develop a molecular diagnostic strategy for this syndrome. Singleton whole-exome sequencing (WES) was performed on sixteen patients. Our study revealed (1) nine patients carried eight homozygous pathogenic variants with four of them being novel (2) Specifically, a synonymous splicing variant (c.471G > A) in BBS2 gene in six patients with Baloch ethnicity. The identification of runs of homozygosity (ROH) calling was performed using the BCFtools/RoH software on WES data of patients harboring c.471G > A variant. The presence of shared homozygous regions containing the identified variant was confirmed in these patients. In-silico analysis predicted the effect of the c.471G > A variants on BBS2 mRNA splicing. This variant results in disrupted wild-type donor site and intron retention in the mature mRNA. (3) And a deletion of exons 14 to 17 in the BBS1 gene was identified in one patient by Copy-Number Variation (CNV) analysis using the ExomeDepth pipeline. Our results identified the founder variant c.471G > A in the BBS2 gene in the Baloch ethnicity of the Iranian population. This finding can guide the diagnostic approach of this syndrome in future studies.






Similar content being viewed by others
Notes
G-protein-coupled receptors.
Alstrom Syndrome Protein 1.
Nephrocystin 4.
Human Splicing Finder.
Abbreviations
- BBS:
-
Bardet–Biedl syndrome
- WES:
-
Whole exome sequencing
- ROH:
-
Runs of homozygosity
- CNV:
-
Copy-Number Variation
- NGS:
-
Next-generation sequencing
References
Beales PL, Elcioglu N, Woolf AS, Parker D, Flinter FA (1999) New criteria for improved diagnosis of Bardet-Biedl syndrome: results of a population survey. J Med Genet 36(6):437–446
Bhanuprakash V, Chhotaray S, Pruthviraj DR, Rawat C, Karthikeyan A, Panigrahi M (2018) Copy number variation in livestock: a mini review. Vet World 11(4):535–541
Bravo-Gil N, Méndez-Vidal C, Romero-Pérez L, González-del Pozo M, Rodríguez-de la Rúa E, Dopazo J et al (2016) Improving the management of Inherited Retinal Dystrophies by targeted sequencing of a population-specific gene panel. Sci Rep 6:23910
Carr IM, Bhaskar S, O’Sullivan J, Aldahmesh MA, Shamseldin HE, Markham AF et al (2013) Autozygosity mapping with exome sequence data. Hum Mutat 34(1):50–56
Dallali H, Kheriji N, Kammoun W, Mrad M, Soltani M, Trabelsi H et al (2021) Multiallelic rare variants in BBS genes support an oligogenic ciliopathy in a non-obese juvenile-onset syndromic diabetic patient: a case report. Front Genet 12:664963
Esposito G, Testa F, Zacchia M, Crispo AA, Di Iorio V, Capolongo G et al (2017) Genetic characterization of Italian patients with Bardet-Biedl syndrome and correlation to ocular, renal and audio-vestibular phenotype: identification of eleven novel pathogenic sequence variants. BMC Med Genet 18(1):10
Fattahi Z, Rostami P, Najmabadi A, Mohseni M, Kahrizi K, Akbari MR et al (2014) Mutation profile of BBS genes in Iranian patients with Bardet-Biedl syndrome: genetic characterization and report of nine novel mutations in five BBS genes. J Hum Genet 59(7):368–375
Forsythe E, Beales PL (2013) Bardet-Biedl syndrome. Eur J Hum Genet 21(1):8–13
Forsythe E, Kenny J, Bacchelli C, Beales PL (2018) Managing Bardet-Biedl syndrome-now and in the future. Front Pediatr 6:23
Grochowsky A, Gunay-Aygun M (2019) Clinical characteristics of individual organ system disease in non-motile ciliopathies. Transl Sci Rare Dis 4(1–2):1–23
https://grch37.ensembl.org/Homo_sapiens/Variation/Population [
Janssen S, Ramaswami G, Davis EE, Hurd T, Airik R, Kasanuki JM et al (2011) Mutation analysis in Bardet-Biedl syndrome by DNA pooling and massively parallel resequencing in 105 individuals. Hum Genet 129(1):79–90
Komlósi K, Gyenesei A, Bene J (2022) Editorial: copy number variation in rare disorders. Front Genet 13:898059
Lindstrand A, Frangakis S, Carvalho CM, Richardson EB, McFadden KA, Willer JR et al (2016) Copy-number variation contributes to the mutational load of Bardet-Biedl syndrome. Am J Hum Genet 99(2):318–336
Mary L, Chennen K, Stoetzel C, Antin M, Leuvrey A, Nourisson E et al (2019) Bardet-Biedl syndrome: antenatal presentation of forty-five fetuses with biallelic pathogenic variants in known Bardet-Biedl syndrome genes. Clin Genet 95(3):384–397
M’Hamdi O, Ouertani I, Chaabouni-Bouhamed H (2014) Update on the genetics of bardet-biedl syndrome. Mol Syndromol 5(2):51–56
Moore SJ, Green JS, Fan Y, Bhogal AK, Dicks E, Fernandez BA et al (2005) Clinical and genetic epidemiology of Bardet-Biedl syndrome in Newfoundland: a 22-year prospective, population-based, cohort study. Am J Med Genet A 132A(4):352–360
Nakayama K, Katoh Y (2020) Architecture of the IFT ciliary trafficking machinery and interplay between its components. Crit Rev Biochem Mol Biol 55(2):179–196
Narasimhan V, Danecek P, Scally A, Xue Y, Tyler-Smith C, Durbin R (2016) BCFtools/RoH: a hidden Markov model approach for detecting autozygosity from next-generation sequencing data. Bioinformatics 32(11):1749–1751
Perea-Romero I, Solarat C, Blanco-Kelly F, Sanchez-Navarro I, Bea-Mascato B, Martin-Salazar E et al (2022) Allelic overload and its clinical modifier effect in Bardet-Biedl syndrome. NPJ Genom Med 7(1):41
Sanchez-Navarro I, Silva LRJD, Blanco-Kelly F, Zurita O, Sanchez-Bolivar N, Villaverde C et al (2018) Combining targeted panel-based resequencing and copy-number variation analysis for the diagnosis of inherited syndromic retinopathies and associated ciliopathies. Sci Rep 8(1):5285
Simons DL, Boye SL, Hauswirth WW, Wu SM (2011) Gene therapy prevents photoreceptor death and preserves retinal function in a Bardet-Biedl syndrome mouse model. Proc Natl Acad Sci USA 108(15):6276–6281
Suspitsin EN, Imyanitov EN (2016) Bardet-Biedl syndrome. Mol Syndromol 7(2):62–71
Tian X, Zhao H, Zhou J (2023) Organization, functions, and mechanisms of the BBSome in development, ciliopathies, and beyond. Elife. https://doi.org/10.7554/eLife.87623
Tobin JL, Beales PL (2008) Restoration of renal function in zebrafish models of ciliopathies. Pediatr Nephrol 23(11):2095–2099
Wang F, Wang H, Tuan HF, Nguyen DH, Sun V, Keser V et al (2014) Next generation sequencing-based molecular diagnosis of retinitis pigmentosa: identification of a novel genotype-phenotype correlation and clinical refinements. Hum Genet 133(3):331–345
Waters AM, Beales PL (2011) Ciliopathies: an expanding disease spectrum. Pediatr Nephrol 26(7):1039–1056
Weisschuh N, Mayer AK, Strom TM, Kohl S, Glöckle N, Schubach M et al (2016) Mutation detection in patients with retinal dystrophies using targeted next generation sequencing. PLoS ONE 11(1):e0145951
Zhuang Z, Gusev A, Cho J, Pe’er I (2012) Detecting identity by descent and homozygosity mapping in whole-exome sequencing data. PLoS ONE 7(10):e47618
Acknowledgements
We express our gratitude to our patients and their families for their collaboration in this research, as well as to the colleagues who assisted us in this study.
Funding
This research was supported by Mashhad University of Medical Sciences and University of Social Welfare and Rehabilitation Sciences.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by MHF, MA, AE, EE, MYVM, MS, ZF, HRKK, MM. The first draft of the manuscript was written by SF and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Ethical Approval
This study was performed in line with the principles of the Declaration of Helsinki. Approval was granted by the Research Ethics Committee of University of Social Welfare and Rehabilitation Science, Tehran, Iran (Date: 2021-12-27/No: IR.USWR.REC.1400.235).
Consent to Participate
Informed consent to participate in this research was obtained from all the patient’s parents.
Consent to Publish
Informed consent for publication of the results was obtained from all the patient’s parents.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Feizabadi, M.H., Alerasool, M., Eslahi, A. et al. Characterizing Homozygous Variants in Bardet-Biedl Syndrome-Associated Genes Within Iranian Families: Unveiling a Founder Variant in BBS2, c.471G>A. Biochem Genet (2024). https://doi.org/10.1007/s10528-023-10637-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10528-023-10637-w