Abstract
Background
Delftia acidovorans is distributed widely in the environment and has the potential to promote the growth of plants and degrade organic pollutants. However, it is also an opportunistic pathogen for human and many reports demonstrated that D. acidovorans has strong resistance to aminoglycosides and polymyxins.
Objective
The aim of this work was to reveal the antibiotic resistance genes and pathogenic genes in a novel conditional pathogenic strain—D. acidovorans B804, which was isolated from the radiation-polluted soil from Xinjiang Uyghur Autonomous Region, China.
Methods
The antibiotic resistance test was performed according to the Kirby–Bauer disk diffusion method and evaluated by the standards of the Clinical and Laboratory Standards Institute guidelines. The genome of D. acidovorans B804 was sequenced by a PacBio RS II and Illumina HiSeq 4000 platform in Shanghai Majorbio Biopharm Technology Co., Ltd. (Shanghai, China).
Results
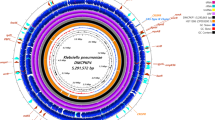
The multidrug resistance phenotypes of D. acidovorans B804 was experimentally confirmed and its genome was sequenced. The total size of D. acidovorans B804 genome was 6,661,314 bp with a GC content of 66.73%. 403 genes associated with antibiotic resistances were predicted. Meanwhile, 89 pathogenic genes were also predicted and 17 of these genes might be capable of causing diseases to human, such as infections and salmonellosis.
Conclusions
This genomic information can be used as a reference sequence for comparative genomic studies. The results provided more insights regarding the pathogenesis and drug resistance mechanism of D. acidovorans, which will be meaningful for developing more effective therapies toward D. acidovorans-related diseases.







Similar content being viewed by others
Data Availability
The whole-genome sequence of D. acidovorans B804 can be downloaded from GenBank with the accession number: CP058970.
References
Ayangbenro AS, Babalola OO (2017) A new strategy for heavy metal polluted environments a review of microbial biosorbents. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph14010094 (PubMed PMID: WOS:000392578200092)
Bilgin H, Sarmis A, Tigen E, Soyletir G, Mulazimoglu L (2016) Delftia acidovorans: a rare pathogen in immunocompetent and immunocompromised patients. Can J Infect Dis Med Microbiol 26(5):277–279
Chotikanatis K, Baecker M, Rosas-Garcia G, Hammerschlag MR (2011) Recurrent intravascular-catheter-related bacteremia caused by Delftia acidovorans in a hemodialysis patient. J Clin Microbiol 49(9):3418–3421. https://doi.org/10.1128/jcm.00625-11 (PubMedPMID:WOS:000294416000063)
Clinical and Laboratory Standards Institute (CLSI) (2022) Performance Standards for Antimicrobial Susceptibility Testing. USA.
Hagiya H, Murase T, Sugiyama J, Kuroe Y, Nojima H, Naito H et al (2013) Delftia acidovorans bacteremia caused by bacterial translocation after organophosphorus poisoning in an immunocompetent adult patient. J Infect Chemother 19(2):338–341. https://doi.org/10.1007/s10156-012-0472-x (PubMedPMID:WOS:000317429200023)
Hojgaard SMM, Rezahosseini O, Knudsen JD, Fuglebjerg NJU, Skov M, Nielsen SD et al (2022) Characteristics and outcomes of patients with Delftia acidovorans infections: a retrospective cohort study. Microbiol Spectr. https://doi.org/10.1128/spectrum.00326-22 (PubMedPMID:WOS:000826300100003)
Jian Z, Zeng L, Xu T, Sun S, Yan S, Yang L et al (2021) Antibiotic resistance genes in bacteria: occurrence, spread, and control. J. Basic Microbiol 61(12):1049–1070. https://doi.org/10.1002/jobm.202100201 (PubMedPMID: WOS:000707252300001)
Kam SK, Lee WS, Ou TY, Teng SO, Chen FL (2012) Delftia acidovorans bacteremia associated with ascending urinary tract infections proved by molecular method. J Exp Clini Med 4(3):180–182
Liu X, Tang WH, Zhao XM, Chen L, Simon R (2010) A network approach to predict pathogenic genes for fusarium graminearum. PLoS ONE 5(10):13021–13022
Patel D, Iqbal AM, Mubarik A, Vassa N, Godil R, Saad M et al (2019) Delftia acidovorans: a rare cause of septic pulmonary embolism from catheter-related infection: case report and literature review. Respir Med Case Rep. https://doi.org/10.1016/j.rmcr.2019.100835 (PubMed PMID: WOS:000477769000012)
Perry BJ, Bergsveinson J, Tambalo DD, Yost CK, Khan NH, Whiting M (2017) Complete genome sequence of Delftia acidovorans RAY209 a plant growth-promoting rhizobacterium for canola and soybean. Genom Announc. https://doi.org/10.1128/genomeA.01224-17 (PubMed PMID: WOS:000460739800018)
Perumal PG, Appalaraju B, Thiruvengadam P (2022) Delftia acidovorans: a rare pathogen isolated from pleural fluid in an immunocompromised host. J Clin Diagn Res. https://doi.org/10.7860/jcdr/2022/58627.16914 (PubMed PMID: WOS:000876477000012)
Rema T, Medihala P, Lawrence JR, Vidovic S, Leppard GG, Reid M et al (2016) Proteomic analyses of chlorhexidine tolerance mechanisms in Delftia acidovorans biofilms. Msphere. https://doi.org/10.1128/mSphere.00017-15 (PubMed PMID: WOS:000392583700011)
Sahay P, Goel S, Nagpal R, Maharana PK, Sinha R, Agarwal T et al (2020) Infectious keratitis caused by rare and emerging micro-organisms. Curr Eye Res 45(7):761–773. https://doi.org/10.1080/02713683.2019.1708407 (PubMedPMID:WOS:000506266900001)
Shetty AR, de Gannes V, Obi CC, Lucas S, Lapidus A, Cheng J-F et al (2015) Complete genome sequence of the phenanthrene-degrading soil bacterium Delftia acidovorans Cs 1–4. Stand Genom Sci. https://doi.org/10.1186/s40793-015-0041-x (PubMed PMID: WOS:000367992200001)
Smits WJ, Feucht H-H, Oellig F, Zoellner B (2020) Infection of the orbita due to Delftia acidovorans after a cat scratch. Dtsch Med Wochenschr 145(21):1559–1561. https://doi.org/10.1055/a-1173-4598 (PubMedPMID:WOS:000581078900013)
Wen A, Fegan M, Hayward C, Chakraborty S, Sly LI (1999) Phylogenetic relationships among members of the comamonadaceae, and description of Delftia acidovorans (den Dooren de Jong 1926 and Tamaoka et al. 1987) gen. nov., comb. nov. Int j Syst Bacteriol. https://doi.org/10.1099/00207713-49-2-567 (PubMed PMID: MEDLINE:10319477)
Yablokov AV, Nesterenko VB, Nesterenko AV (2009) Chapter III consequences of the chernobyl catastrophe for the environment. Ann NY Acad Sci. https://doi.org/10.1111/j.1749-6632.2009.04830.x (PubMed PMID: MEDLINE:20002049)
Funding
This work was financially supported by China Postdoctoral Science Foundation (2021M690081).
Author information
Authors and Affiliations
Contributions
K.L. and X.L. wrote the main manuscript text and W.L.Z. and N.W. prepared figures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical Approval
Not required (This research did not involve humans or animals).
Consent to Publication
All authors have read and approved this version of the article and due care has been taken to ensure the integrity of the work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, K., Liu, X., Zuo, W. et al. Whole-Genome Sequencing of a Multidrug-Resistant Strain: Delftia acidovorans B408. Biochem Genet 61, 1086–1096 (2023). https://doi.org/10.1007/s10528-022-10306-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-022-10306-4