Abstract
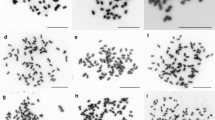
Polyploidy is a phenomenon that alters the genetic diversity of populations and has been reported as one of the most important evolutionary forces for plant diversification. The Psidium cattleyanum complex comprises a group of wild populations with several ploidy levels reported in the literature. The multiple cytotypes, associated with its wide distribution area, make this species a potential key model for understanding evolutionary processes related to polyploidization. In this study, we isolated and characterized nuclear microsatellite markers of P. cattleyanum and tested their transferability to other nine species of the genus. We performed a preliminary analysis of genetic diversity and population structure in three populations of P. cattleyanum. The three populations analyzed had different chromosome numbers, being polyploid cytotypes (2n = 6x = 66, 2n = 7x = 77 and 2n = 8x = 88). We designed 46 primer pairs and successfully amplified 37 markers, from which the 10 best were selected for analysis. Considering both the PIC and DP values, most of markers were highly informative. The new SSR markers were used to assess the levels of genetic diversity of the populations and detected one population with predominance of sexual reproduction. DAPC analysis pointed the formation of three groups, which corresponded to the populations analyzed. The markers were successfully amplified in related species, with some species presenting 80% transferability. By producing this panel of polymorphic microsatellites, we contribute to the understanding evolution in groups of natural polyploids for future studies.


Similar content being viewed by others
References
Atchison E (1947) Chromosome numbers in the Myrtaceae. Am J Bot. https://doi.org/10.1002/j.1537-2197.1947.tb12970.x
Baduel P, Bray S, Vallejo-Marin M et al (2018) The “polyploid hop”: shifting challenges and opportunities over the evolutionary lifespan of genome duplications. Front Ecol Evol. https://doi.org/10.3389/fevo.2018.00117
Baniaga AE, Marx HE, Arrigo N, Barker MS (2019) Polyploid plants have faster rates of multivariate niche differentiation than their diploid relatives. Ecol Lett. https://doi.org/10.1111/ele.13402
Billotte N, Lagoda PJL, Risterucci AM, Baurens FC (1999) Microsatellite-enriched libraries: applied methodology for the development of SSR markers in tropical crops. Fruits 54:277–288
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32(3):314
Chen ZJ, Ni Z (2006) Mechanisms of genomic rearrangements and gene expression changes in plant polyploids. BioEssays. https://doi.org/10.1002/bies.20374
Clark LV, Jasieniuk M (2011) POLYSAT: an R package for polyploid microsatellite analysis. Mol Ecol Resour. https://doi.org/10.1111/j.1755-0998.2011.02985.x
Coser SM, Ferreira MFS, Ferreira A, Mitre LK, Carvalho CR, Clarindo WR (2012) Assessment of genetic diversity in Psidium guajava L. using different approaches. Sci Hortic. https://doi.org/10.1016/j.scienta.2012.09.030
Costa IR (2009) Estudos evolutivos em Myrtaceae: aspectos citotaxonômicos e filogenéticos em Myrteae, enfatizando Psidium e gêneros relacionados. Dissertação de Mestrado, Universidade Estadual de Campinas
Costa IR, Forni-Martins ER (2006) Chromosome studies in Brazilian species of Campomanesia Ruiz & Pávon and Psidium L. (Myrtaceae Juss.). Caryologia. https://doi.org/10.1080/00087114.2006.10797891
Cruz RP, Federizzi LC, Milach SCK (1998) A apomixia no melhoramento de plantas. Cienc Rural. https://doi.org/10.1590/S0103-84781998000100028
De Oliveira BC, Tuler AC, Ferreira A, Carvalho MS, Nogueira AM, da Silva Ferreira MF (2018) Transferability of Psidium microsatellite loci in Myrteae (Myrtaceae): a phylogenetic signal. Euphytica. https://doi.org/10.1007/s10681-018-2195-2
De Souza ADG, Resende LV, De Lima IP, Martins LSS, Techio VH (2015) Chromosome number and nuclear DNA amount in Psidium spp. resistant and susceptible to Meloidogyne enterolobii and its relation with compatibility between rootstocks and commercial varieties of guava tree. Plant Syst Evol. https://doi.org/10.1007/s00606-014-1068-y
De Wet JMJ (1980) Origins of polyploids. In: Lewis WH (ed) Polyploidy: biological relevance. Springer, Boston
Franzon RC, Campos LDO, Proença CEB, Sousa-Silva JC (2009) Araçás do gênero Psidium: principais espécies, ocorrência, descrição e usos. Embrapa, Planaltina https://www.infoteca.cnptia.embrapa.br/infoteca/bitstream/doc/697560/1/doc266.pdf
Global Invasive Species Database (2020) Species profile: Psidium cattleianum. https://www.iucngisd.org/gisd/speciesname/Psidium+cattleianum. Accessed 28 April 2020
Guerra M (2002) Como observar cromossomos: um guia de técnicas em citgenética vegetal, animal e humana. Funpec, Ribeirão Preto
Hirano RT, Nakazone HY (1969) Chromosome numbers of ten species and clones in the genus Psidium. J Am Soc Hortic Sci 94:83–86
Hodel RG, Segovia-Salcedo MC, Landis JB et al (2016) The report of my death was an exaggeration: a review for researchers using microsatellites in the 21st century. Appl Plant Sci. https://doi.org/10.3732/apps.1600025
Jombart T, Devillard S, Balloux F (2010) Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC genet. https://doi.org/10.1186/1471-2156-11-94
Kamvar ZN, Tabima JF, Grünwald NJ (2014) Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. Peer J. https://doi.org/10.7717/peerj.281
Kherwar D, Usha K, Mithra SA, Singh B (2018) Microsatellite (SSR) marker assisted assessment of population structure and genetic diversity for morpho-physiological traits in guava (Psidium guajava L.). J Plant Biochem Biot. https://doi.org/10.1007/s13562-017-0438-2
Kolár F, Čertner M, Suda J, Schönswetter P, Husband BC (2017) Mixed-ploidy species: progress and opportunities in polyploid research. Trends Plant Sci. https://doi.org/10.1016/j.tplants.2017.09.011
Machado RM (2016) Distribuição geográfica e análise cariotípica de citótipos de Psidium cattleianum Sabine (Myrtaceae). Dissertação de Mestrado, Universidade Estadual de Campinas
Mani A, Mishra R, Thomas G (2011). Elucidation of diversity among Psidium species using morphological and SPAR methods. J Phytol. https://updatepublishing.com/journal/index.php/jp/article/view/2305
Marques A, Tuler AC, Carvalho CR et al (2016) Refinement of the karyological aspects of Psidium guineense (Swartz,1788): a comparison with Psidium guajava (Linnaeus, 1753). Comp Cytogenet. https://doi.org/10.3897/CompCytogen.v10i1.6462
Medina SNV (2014) Psidium cattleyanum Sabine y Acca sellowiana (Berg.) Burret (Myrtaceae): caracterización cromossómica y cariotípica em poblaciones silvestre y genótipos selecionados em programas nacionales de mejoramiento. Universidade de la República, Monografia
Meirmans PG, Liu S, Van Tienderen PH (2018) The analysis of polyploid genetic data. J Hered. https://doi.org/10.1093/jhered/esy006
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number individuals. Genetics 89:583–590
Noia LR, Tuler AC, Ferreira A, Ferreira MFS (2017) Relationship between Psidium species (Myrtaceae) by resistance gene analog markers: focus on nematode resistance. Genet Mol Res. https://doi.org/10.4238/gmr16019441
Normand F, Habib R (2001) Phenology of strawberry guava (Psidium cattleianum) in ReÂunion Island. J Hortic Sci Biotech. https://doi.org/10.1080/14620316.2001.11511407
Oliveira FA, Cidade FW, Fávero AP, Vigna BB, Souza AP (2016) First microsatellite markers for Paspalum plicatulum (Poaceae) characterization and cross-amplification in different Paspalum species of the Plicatula group. BMC Res Notes. https://doi.org/10.1186/s13104-016-2312-z
Otto SP, Whitton J (2000) Polyploidy: incidence and evolution. Annu Rev Genet. https://doi.org/10.1146/annurev.genet.34.1.401
Queirós J, Godinho R, Lopes S et al (2015) Effect of microsatellite selection on individual and population genetic inferences: an empirical study using cross-specific and species-specific amplifications. Mol Ecol Resour. https://doi.org/10.1111/1755-0998.12349
Raseira MCB, Raseira A (1996) Contribuição ao estudo do araçazeiro. Embrapa, Pelotas
Schifino-Wittmann MT (2004) Poliploidia e seu impacto na origem e evolução das plantas silvestres e cultivadas. Agric Sci Technol. https://doi.org/10.18539/cast.v10i.2.904
Schuelke M (2000) An economic method for the fluorescent labeling of PCR fragments. Nat Biotechnol. https://doi.org/10.1038/72708
Shannon CE (1948) A mathematical theory of communication. Mob Comput Commun. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Sharma AS, Sehrawat SK, Singhrot RS, Boora KS (2005) Assessment of genetic diversity and relationship among Psidium spp. through RAPD analysis. International Guava Symposium. Acta Hort. https://doi.org/10.17660/ActaHortic.2007.735.7
Simpson EH (1949) Measurement of diversity. Nature. https://doi.org/10.1038/163688a0
Singhal VK, Gill BS, Bir SS (1984) Cytology of cultivated woody species (Polypetalae). Proc Indian Sci Congr Assoc 71:143–144
Soltis DE, Albert VA, Leebens-Mack J et al (2009) Polyploidy and angiosperm diversification. Am J Bot. https://doi.org/10.3732/ajb.0800079
Soltis DE, Buggs RJA, Doyle JJ, Soltis PS (2010) What we still don’t know about polyploidy. Taxon 59:1387–1403
Soltis DE, Segovia-Salcedo MC, Jordon-Thaden I et al (2014) Are polyploids really evolutionary dead-ends (again)? A critical reappraisal of Mayrose et al. (2011). New Phytol. https://doi.org/10.1111/nph.12756
Soltis PS, Soltis DE (2000) The role of genetic and genomic attributes in the success of polyploids. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.97.13.7051
Souza-Pérez M, Speroni G (2017) New apomictic pathway in Myrtaceae inferred from Psidium cattleyanum female gametophyte ontogeny. Flora. https://doi.org/10.1016/j.flora.2017.06.010
te Beest M, Le Roux JJ, Richardson DM et al (2011) The more the better? The role of polyploidy in facilitating plant invasions. Ann Bot. https://doi.org/10.1093/aob/mcr277
Tel-Zur N, Abbo S, Myslabodski D et al (1999) Modified CTAB procedure for DNA isolation from epiphytic cacti of the genera Hylocereus and Selenicereus (Cactaceae). Plant Mol Biol Rep. https://doi.org/10.1023/A:1007656315275
Tessier C, David J, This P et al (1999) Optimization of the choice of molecular markers for varietal identification in Vitis vinifera L. Theor Appl Genet. https://doi.org/10.1007/s001220051054
Tuler AC, Carrijo TT, Nóia LR et al (2015) SSR markers: a tool for species identification in Psidium (Myrtaceae). Mol Biol Rep. https://doi.org/10.1007/s11033-015-3927-1
Tuler AC, Carrijo TT, Peixoto AL et al (2019) Diversification and geographical distribution of Psidium (Myrtaceae) species with distinct ploidy levels. Trees. https://doi.org/10.1007/s00468-019-01845-2
Van Drunen WE, Husband BC (2019) Evolutionary associations between polyploidy, clonal reproduction, and perenniality in the angiosperms. New Phytol. https://doi.org/10.1111/nph.15999
Wikler C (2007) Problemática do araçazeiro como planta invasora. In: Pedrosa-Macedo JH, DalMolin A, Smith CW (eds) O araçazeiro: ecologia e controle biológico. Fupef, Curitiba
Acknowledgements
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES)—Finance Code 001. We thank the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and FAEPEX/UNICAMP for the financial support. FAO received a Post-Doctoral fellowship from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP 2018/18527-9). We thank Prof. Marcia Dr. Flores da Silva Ferreira, of Universidade Federal do Espírito Santo, for sending the different Psidium species plant material, and Dr. André Vito Scatigna for his support in fieldwork and collecting plant material from the individual grown in Uruguaiana (RS). We also thank the staff of Laboratório de Análises Genéticas e Moleculares (LAGM/UNICAMP).
Funding
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior—Brasil (CAPES)—Finance Code 001, Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) and FAEPEX/UNICAMP. FAO received a Post-Doctoral fellowship from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP 2018/18527-9).
Author information
Authors and Affiliations
Contributions
All authors contributed equally to this manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they have no conflicts of interest.
Ethics Approval
This article does not include studies about humans or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Machado, R.M., de Oliveira, F.A., de Matos Alves, F. et al. Population Genetics of Polyploid Complex Psidium cattleyanum Sabine (Myrtaceae): Preliminary Analyses Based on New Species-Specific Microsatellite Loci and Extension to Other Species of the Genus. Biochem Genet 59, 219–234 (2021). https://doi.org/10.1007/s10528-020-10002-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10528-020-10002-1