Abstract
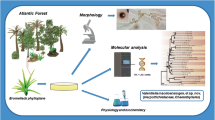
As part of a long-term study aiming to isolate and identify yeast species that inhabit the surface of leaves and fruits of native fine-aroma cacao in the department of Amazonas, Peru, we obtained multiple isolates of Hannaella species. Yeasts of the genus Hannaella are common inhabitants of the phyllosphere of natural and crop plants. On the basis of morphological, and physiological characteristics, and sequence analysis of the D1/D2 domains of the large subunit rRNA gene (LSU) and the internal transcribed spacer region (ITS), we identified five species of Hannaella from the phyllosphere of Peruvian cacao. Four have been previously described: H. phyllophila (isolates KLG-073, KLG-091), H. pagnoccae (KLG-076), H. sinensis (KLG-121), and H. taiwanensis (KLG-021). A fifth, represented by eight isolates (KLG-034, KLG-063, KLG-074, KLG-078, KLG-79, KLG-082, KLG-084, KLG-085), is not conspecific with any previously described Hannaella species, and forms the sister clade to H. surugaensis in the phylogenetic analysis. It has 2.6–3.9% (18–27 substitutions, 2–4 deletions, and 1–3 insertions in 610–938 bp-long alignments), and 9.8–10.0% nucleotide differences (37 substitutions and 14 insertions in 511–520 bp-long alignments) in the LSU and ITS regions, respectively, to H. surugaensis type strain, CBS 9426. Herein, the new species Hannaella theobromatis sp. nov. is described and characterised. The species epithet refers to its epiphytic ecology on its host Theobroma cacao.



Similar content being viewed by others
Data availability
All sequences generated in this study are publicly available under the NCBI Accession Numbers: OR086918-25, OR088043-50, OR088135-39, and OR088112-16 (For details, see Supplementary Table S1).
References
Aires A, Gonçalves C, Sampaio JP (2023) Hannaella floricola sp. nov., a novel basidiomycetous yeast species isolated from a flower of Lantana camara in Portugal. Int J Syst Evolut Microbiol 7:3. https://doi.org/10.1099/ijsem.0.005740
Albu S, Toome M, Aime MC (2015) Violaceomyces palustris gen. et sp. nov. and a new monotypic lineage, violaceomycetales ord. nov. in ustilaginomycetes. Mycologia 107:1193–1204. https://doi.org/10.3852/14-260
Belisle M, Mendenhall CD, Oviedo Brenes F, Fukami T (2014) Temporal variation in fungal communities associated with tropical hummingbirds and nectarivorous bats. Fungal Ecol 12:44–51. https://doi.org/10.1016/j.funeco.2014.02.007
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K et al (2009) BLAST+: architecture and applications. BMC Bioinform 10:421. https://doi.org/10.1186/1471-2105-10-421
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and high-performance computing. Nat Methods 9(8):772
Díaz-Valderrama JR, Leiva-Espinoza ST, Aime MC (2020) The history of cacao and its diseases in the Americas. Phytopathology 110:1604–1619. https://doi.org/10.1094/PHYTO-05-20-0178-RVW
Díaz-Valderrama JR, Zambrano R, Cedeño-Amador S, Córdova-Bermejo U, Casas GG, García-Zurita N et al (2022) Diversity in the invasive cacao pathogen Moniliophthora roreri is shaped by agriculture. Plant Pathol 71:1721–1734. https://doi.org/10.1111/ppa.13603
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. https://doi.org/10.1093/nar/gkh340
Fell JW, Boekhout T, Fonseca A, Scorzetti G, Statzell-Tallman A (2000) Biodiversity and systematics of basidiomycetous yeasts as determined by large-subunit rDNA D1/D2 domain sequence analysis. Int J Syst Evol Microbiol 50:1351–1371. https://doi.org/10.1099/00207713-50-3-1351
Glushakova AM, Yurkov AM, Chernov IY (2007) Massive isolation of anamorphous ascomycete yeasts Candida oleophila from plant phyllosphere. Microbiology 76:799–803. https://doi.org/10.1134/S0026261707060215
Gouy M, Guindon S, Gascuel O (2010) SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol Biol Evol 27:221–224. https://doi.org/10.1093/molbev/msp259
Han L, Li ZY, Guo XF, Tan JL, He SZ, Cui XL et al (2017) Hannaella dianchiensis sp. nov., a basidiomycetous yeast species isolated from lake water. Int J Syst Evol Microbiol 67:2014–2018. https://doi.org/10.1099/ijsem.0.001908
Huang C, Lee F, Tien C (2011) Cryptococcus taiwanensis sp. nov., a novel yeast in Taiwan. Fungal Sci 26:57–64
Inácio J, Portugal L, Spencer-Martins I, Fonseca Á (2005) Phylloplane yeasts from Portugal: seven novel anamorphic species in the Tremellales lineage of the Hymenomycetes (Basidiomycota) producing orange-coloured colonies. FEMS Yeast Res 5:1167–1183. https://doi.org/10.1016/j.femsyr.2005.05.007
Jaiboon K, Lertwattanasakul N, Limtong P, Limtong S (2016) Yeasts from peat in a tropical peat swamp forest in Thailand and their ability to produce ethanol, indole-3-acetic acid and extracellular enzymes. Mycol Prog 15:755–770. https://doi.org/10.1007/s11557-016-1205-9
da Junior ASE, Solís K, Pimenta A, Vera DI, Garzón I, Peñaherrera S et al (2022) Effect of antagonistic yeasts from cacao tissues on controlling growth and sporulation of Moniliophthora roreri. Biol Control 172:104956. https://doi.org/10.1016/j.biocontrol.2022.104956
Kaewwichian R, Jindamorakot S, Am-In S, Sipiczki M, Limtong S (2015) Hannaella siamensis sp. nov. and Hannaella phetchabunensis sp. nov., two new anamorphic basidiomycetous yeast species isolated from plants. Int J Syst Evol Microbiol 65:1297–1303. https://doi.org/10.1099/ijs.0.000101
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kurtzman CP, Fell JW, Boekhout T, Robert V (2011) Methods for isolation, phenotypic characterization and maintenance of yeasts. The Yeasts. Elsevier, Amsterdam, pp 87–110
Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek 98:331–371
Landell MF, Billodre R, Ramos JP, Leoncini O, Vainstein MH, Valente P (2010) Candida aechmeae sp. nov. and Candida vrieseae sp. nov., novel yeast species isolated from the phylloplane of bromeliads in Southern Brazil. Int J Syst Evol Microbiol 60:244–248. https://doi.org/10.1099/ijs.0.011577-0
Landell MF, Brandão LR, Barbosa AC, Ramos JP, Safar SVB, Gomes FCO et al (2014) Hannaella pagnoccae sp. nov., a tremellaceous yeast species isolated from plants and soil. Int J Syst Evol Microbiol 64:1970–1977. https://doi.org/10.1099/ijs.0.059345-0
Liu XZ, Wang QM, Göker M, Groenewald M, Kachalkin AV, Lumbsch HT et al (2015) Towards an integrated phylogenetic classification of the Tremellomycetes. Stud Mycol 81:85–147. https://doi.org/10.1016/j.simyco.2015.12.001
Lu H, Lu L, Zeng L, Fu D, Xiang H, Yu T et al (2014) Effect of chitin on the antagonistic activity of Rhodosporidium paludigenum against Penicillium expansum in apple fruit. Postharvest Biol Technol 92:9–15. https://doi.org/10.1016/j.postharvbio.2014.01.009
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, Von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37(5):1530–1534
Miller MA, Pfeiffer W, Schwartz T (2011) The CIPRES science gateway: a community resource for phylogenetic analyses. In: Proceedings of the 2011 TeraGrid conference: extreme digital discovery, pp 1–8
Molnár O, Prillinger H (2006) Cryptococcus zeae, a new yeast species associated with Zea mays. Microbiol Res 161:347–354. https://doi.org/10.1016/j.micres.2005.12.005
Molnárová J, Vadkertiová R, Stratilová E (2014) Extracellular enzymatic activities and physiological profiles of yeasts colonizing fruit trees. J Basic Microbiol 54:1–11. https://doi.org/10.1002/jobm.201300072
Nagahama T, Hamamoto M, Nakase T, Takaki Y, Horikoshi K (2003) Cryptococcus surugaensis sp. nov., a novel yeast species from sediment collected on the deep-sea floor of Suruga Bay. Int J Syst Evol Microbiol 53:2095–2098. https://doi.org/10.1099/ijs.0.02712-0
Neto AAP, Santos TR, Duarte EAA, de Oliveira TAS, de Andrade Silva EM, Uetanabaro APT et al (2021) Yeasts associated with aerial parts of Theobroma cacao L. in southern Bahia, Brazil, as prospective biocontrol agents against Moniliophthora perniciosa. Tropical Plant Pathol 46:109–128. https://doi.org/10.1007/s40858-020-00418-w
Oliva Cruz SM (2020) Caracterización socioeconómica de la diversidad biológica de cacao criollo fino de aroma en comunidades rurales de la región Amazonas. Universidad Nacional Toribio Rodríguez de Mendoza de Amazonas, Amazonas
Rambaut A (2006) FigTree: tree figure drawing tool. http://tree.bio.ed.ac.uk/
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci U S A 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Scorzetti G, Fell JW, Fonseca A, Statzell-Tallman A (2002) Systematics of basidiomycetous yeasts: a comparison of large subunit D1/D2 and internal transcribed spacer rDNA regions. FEMS Yeast Res 2:495–517. https://doi.org/10.1016/S1567-1356(02)00128-9
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Surussawadee J, Jindamorakot S, Nakase T, Lee CF, Limtong S (2015) Hannaella phyllophila sp. nov., a basidiomycetous yeast species associated with plants in Thailand and Taiwan. Int J Syst Evol Microbiol 65:2135–2140. https://doi.org/10.1099/ijs.0.000231
Tanimura A, Adachi H, Tanabe K, Ogawa J, Shima J (2023) Hannaella oleicumulans sp. nov. and Hannaella higashiohmiensis sp. nov., two novel oleaginous basidiomycetous yeast species. Int J Syst Evol Microbiol 73:1–8. https://doi.org/10.1099/ijsem.0.006027
Toome M, Aime CM, Roberson RW (2013) Meredithblackwellia eburnea gen. et sp. nov., kriegeriaceae fam. nov. and kriegeriales ord. nov.-toward resolving higher-level classification in microbotryomycetes. Mycologia 105:486–495. https://doi.org/10.3852/12-251
Wang QM, Bai FY (2008) Molecular phylogeny of basidiomycetous yeasts in the Cryptococcus luteolus lineage (Tremellales) based on nuclear rRNA and mitochondrial cytochrome b gene sequence analyses: Proposal of Derxomyces gen. nov. and Hannaella gen. nov., and description of eigh. FEMS Yeast Res 8:799–814. https://doi.org/10.1111/j.1567-1364.2008.00403.x
Wang QM, Bai FY, Lu HZ, Jia JH, Takashima M (2004) Bullera cylindrica sp. nov., Bullera hubeiensis sp. nov. and Bullera nakasei sp. nov., ballistoconidium-forming yeast species from plant leaves. Int J Syst Evol Microbiol 54:1877–1882. https://doi.org/10.1099/ijs.0.63092-0
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Zhang H, Wang L, Dong Y, Jiang S, Cao J, Meng R (2007) Postharvest biological control of gray mold decay of strawberry with Rhodotorula glutinis. Biol Control 40:287–292. https://doi.org/10.1016/j.biocontrol.2006.10.008
Acknowledgements
The authors would like to thank to all members of the Plant Health Laboratory of the Instituto de Investigación para el Desarrollo Sustentable de Ceja de Selva from the National University Toribio Rodríguez de Mendoza de Amazonas, for their help during the conduction of this study.
Funding
This study was funded by the Project “Creación e Implementación del Centro de Investigación e Innovación Tecnológica en Cacao de la Universidad Nacional Toribio Rodríguez de Mendoza de Amazonas” (CEINCACAO), CUI N° 2315081; and by PROCIENCIA-CONCYTEC, Peru, under contract N° 057-2021-FONDECYT.
Author information
Authors and Affiliations
Contributions
JRD-V and MCA contributed to the study conception and design. Material preparation, data collection was performed by KJL-G. Data analyses were performed by KJL-G and JRD-V. The first draft of the manuscript was written by KJL-G, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10482_2024_1936_MOESM1_ESM.docx
Supplementary file 1. Supplementary Table S1. Passport and sequence information of isolates collected in this study and used in the phylogenetic study
10482_2024_1936_MOESM2_ESM.xlsx
Supplementary file 2. Supplementary Table S2. Oxidation and assimilation tests of carbon-based compounds of the eigth strains of Hannaella theobromatis sp. nov.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Llanos-Gómez, K.J., Aime, M.C. & Díaz-Valderrama, J.R. The surface of leaves and fruits of Peruvian cacao is home for several Hannaella yeast species, including the new species Hannaella theobromatis sp. nov.. Antonie van Leeuwenhoek 117, 43 (2024). https://doi.org/10.1007/s10482-024-01936-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10482-024-01936-2