Abstract
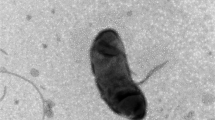
A novel bacterial strain designated CJ43T was isolated from fresh water located in Gangwon-do, South Korea, displaying multi-drug resistance. The isolate was Gram-stain-negative, aerobic, orange-pigmented, and rod-shaped. Strain CJ43T grew optimally at 30 °C and pH 7 on R2A agar in the absence of NaCl. Phylogenetic analyses based on 16S rRNA gene sequences revealed that strain CJ43T belonged to the genus Pedobacter in the family Sphingobacteriaceae and was most closely related to Pedobacter puniceum HX-22-1 T and P. glucosidilyticus 1-2 T (98.3 and 98.1% sequence similarity). The genome size of strain CJ43T was 3.9 Mb in a single contig with DNA G + C content of 34.9%. The genome included 3144 predicted protein-coding genes, as well as 55 tRNA, 9 rRNA and 3 ncRNA genes. The genome also contained 128 putative antibiotic resistance genes, reflecting its phenotypes. The average nucleotide identity values between strain CJ43T and two closely related strains P. puniceum HX-22-1 T and P. glucosidilyticus 1-2 T were 91.0 and 88.7%, respectively. In silico digital DNA-DNA hybridization results between strain CJ43T and the related strains were 42.8 and 38.6%, respectively. The major fatty acids of strain CJ43T were iso-C15:0, iso-C17:0 3-OH, and summed feature 3 (C16:1 ω6c and/or C16:1 ω7c). Strain CJ43T contained phosphatidylethanolamine as the major polar lipid and menaquinone-7 as the sole respiratory quinone. Based on the polyphasic taxonomy data, strain CJ43T represents a novel species of the genus Pedobacter, for which the name Pedobacter aquae sp. nov. is proposed with the type strain CJ43T (= KACC 21350 T = JCM 33709 T).


Similar content being viewed by others
Abbreviations
- ANI:
-
Average Nucleotide Identity
- dDDH:
-
Digital DNA-DNA hybridization
- ML:
-
Maximum-likelihood
- MP:
-
Maximum-parsimony
- NJ:
-
Neighbor-joining
- SMRT:
-
Single Molecule Real-Time
- UBCG:
-
Up-to-date bacterial core-gene
References
An DS, Kim SG, Ten LN, Cho CH (2009) Pedobacter daechungensis sp. nov., from fresh water lake sediment in South Korea. Int J Syst Evol Microbiol 59:69–72. https://doi.org/10.1099/ijs.0.001529-0
Baik KS, Park YD, Kim MS, Park SC, Moon EY et al (2007) Pedobacter koreensis sp. nov., isolated from fresh water. Int J Syst Evol Microbiol 57:2079–2083. https://doi.org/10.1099/ijs.0.65186-0
Baker G, Smith JJ, Cowan DA (2003) Review and re-analysis of domain-specific 16S primers. J Microbiol Methods 55:541–555. https://doi.org/10.1016/j.mimet.2003.08.009
Bayliss SC, Thorpe HA, Coyle NM, Sheppard SK, Feil EJ (2019) PIRATE: a fast and scalable pangenomics toolbox for clustering diverged orthologues in bacteria. GigaScience. https://doi.org/10.1093/gigascience/giz119
Bjerketorp J, Levenfors JJ, Nord C, Guss B, Öberg B, Broberg A (2021) Selective isolation of multidrug-resistant Pedobacter spp, producers of novel antibacterial peptides. Front Microbiol 12:642829. https://doi.org/10.3389/fmicb.2021.642829
Chun J, Kang JY, Jahng KY (2014) Pedobacter pituitosus sp. nov., isolated from a waterfall. Int J Syst Evol Microbiol 64:3838–3843. https://doi.org/10.1099/ijs.0.065235-0
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR et al (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466. https://doi.org/10.1099/ijsem.0.002516
Collins MD, Jones D (1981) Distribution of isoprenoid quinone structural types in bacteria and their taxonomic implications. Microbiol Rev 45:316–354. https://doi.org/10.1128/mr.45.2.316-354.1981
Costa MS, Albuquerque L, Nobre MF, Wait R (2011) The identification of polar lipids in prokaryotes. Methods Microbiol 38:165–181. https://doi.org/10.1016/B978-0-12-387730-7.00007-3
Cui MD, Wang X, Jiang WK, Hu G, Yang ZG et al (2018) Pedobacter agrisoli sp. nov., isolated from farmland soil. Int J Syst Evol Microbiol 68:886–891. https://doi.org/10.1099/ijsem.0.002604
Dusa A (2021) venn: draw venn diagrams. Available online: https://cran.r-project.org/web/packages/venn/venn.pdf (accessed March 15, 2021)
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791. https://doi.org/10.1111/j.1558-5646.1985.tb00420.x
Frank JA, Reich CI, Sharma S, Weisbaum JS, Wilson BA et al (2008) Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl Environ Microbiol 74:2461–2470. https://doi.org/10.1128/AEM.02272-07
Gallego V, García MT, Ventosa A (2006) Pedobacter aquatilis sp. nov., isolated from drinking water, and emended description of the genus Pedobacter. Int J Syst Evol Microbiol 56:1853–1858. https://doi.org/10.1099/ijs.0.64176-0
Gordon NS, Valenzuela A, Adams SM, Ramsey PW, Pollock JL et al (2009) Pedobacter nyackensis sp. nov., Pedobacter alluvionis sp. nov. and Pedobacter borealis sp. nov., isolated from Montana flood-plain sediment and forest soil. Int J Syst Evol Microbiol 59:1720–1726. https://doi.org/10.1099/ijs.0.000158-0
Haft DH, DiCuccio M, Badretdin A, Brover V, Chetvernin V et al (2018) RefSeq: an update on prokaryotic genome annotation and curation. Nucleic Acids Res 46:D851–D860. https://doi.org/10.1093/nar/gkx1068
Hu SL, Huang JW, Yang MJ, Jia WB, Ke Z et al (2019) Pedobacter helvus sp. nov., isolated from farmland soil. Int J Syst Evol Microbiol 69:3806–3811. https://doi.org/10.1099/ijsem.0.003684
Hyatt D, Chen GL, LoCascio PF, Land ML, Larimer FW et al (2010) Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinformatics 11:119. https://doi.org/10.1186/1471-2105-11-119
Jia B, Raphenya AR, Alcock B, Waglechner N, Guo P et al (2017) CARD 2017: expansion and model-centric curation of the comprehensive antibiotic resistance database. Nucleic Acids Res 45:D566–D573. https://doi.org/10.1093/nar/gkw1004
Joung Y, Jang HJ, Park M, Song J, Cho JC (2018) Pedobacter aquicola sp. nov., isolated from fresh water. J Microbiol 56:478–484. https://doi.org/10.1007/s12275-018-7499-3
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic Press, New York, pp 21–132
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30. https://doi.org/10.1093/nar/28.1.27
Kang H, Kim H, Joug Y, Joh K (2014) Pedobacter rivuli sp. nov., isolated from a fresh water stream. Int J Syst Evol Microbiol 64:4073–4078. https://doi.org/10.1099/ijs.0.067579-0
Kim M, Oh HS, Park SC, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351. https://doi.org/10.1099/ijs.0.059774-0
Kim J, Na SI, Kim D, Chun J (2021) UBCG2: Up-to-date bacterial core genes and pipeline for phylogenomic analysis. J Microbiol 59:609–615. https://doi.org/10.1007/s12275-021-1231-4
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120. https://doi.org/10.1007/BF01731581
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Kwon SW, Kim BY, Lee KH, Jang KY, Seok SJ et al (2007) Pedobacter suwonensis sp. nov., isolated from the rhizosphere of Chinese cabbage (Brassica campestris). Int J Syst Evol Microbiol 57:480–484. https://doi.org/10.1099/ijs.0.64196-0
Kwon SW, Son JA, Kim SJ, Kim YS, Park IC et al (2011) Pedobacter rhizosphaerae sp. nov. and Pedobacter soli sp. nov., isolated from rhizosphere soil of Chinese cabbage (Brassica campestris). Int J Syst Evol Microbiol 61:2874–2879. https://doi.org/10.1099/ijs.0.026781-0
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Lee K, Kim DW, Lee DH, Kim YS, Bu JH et al (2020) Mobile resistome of human gut and pathogen drives anthropogenic bloom of antibiotic resistance. Microbiome 8:2. https://doi.org/10.1186/s40168-019-0774-7
Luo X, Wang Z, Dai J, Zhang L, Li J et al (2010) Pedobacter glucosidilyticus sp. nov., isolated from dry riverbed soil. Int J Syst Evol Microbiol 60:229–233. https://doi.org/10.1099/ijs.0.008060-0
Meier-Kolthoff JP, Klenk HP, Goker M (2014) Taxonomic use of DNA G+C content and DNA–DNA hybridization in the genomic age. Int J Syst Evol Microbiol 64:352–356. https://doi.org/10.1099/ijs.0.056994-0
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M et al (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241. https://doi.org/10.1016/0167-7012(84)90018-6
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, Oxford
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ et al (2014) The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:206–214. https://doi.org/10.1093/nar/gkt1226
Parte AC, Sardà Carbasse J, Meier-Kolthoff JP, Reimer LC, Göker M (2020) List of Prokaryotic names with standing in nomenclature (LPSN) moves to the DSMZ. Int J Syst Evol Microbiol 70:5607–5612. https://doi.org/10.1099/ijsem.0.004332
Price MN, Dehal PS, Arkin AP (2010) FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS ONE 5:e9490. https://doi.org/10.1371/journal.pone.0009490
Qiu X, Qu Z, Jiang F, Ren L, Chang X et al (2014) Pedobacter huanghensis sp. nov. and Pedobacter glacialis sp. nov., isolated from Arctic glacier foreland. Int J Syst Evol Microbiol 64:2431–2436. https://doi.org/10.1099/ijs.0.061648-0
Reichenbach H (2006) The order Cytophagales. In: Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt E (eds) The prokaryotes: a handbook on the biology of bacteria. Springer, New York, pp 549–590
Richter M, Rosselló-Móra R (2009) Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106:19126–19131. https://doi.org/10.1073/pnas.0906412106
Rosselló-Móra R, Amann R (2015) Past and future species definitions for bacteria and archaea. Syst Appl Microbiol 38:209–216. https://doi.org/10.1016/j.syapm.2015.02.001
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Sasser M (1990) Identification of bacteria by gas chromatography of cellular fatty acids. MIDI Inc Technical Notes 101:1
Shivaji S, Chaturvedi P, Reddy GS, Suresh K (2005) Pedobacter himalayensis sp. nov., from the Hamta glacier located in the Himalayan mountain ranges of India. Int J Syst Evol Microbiol 55:1083–1088. https://doi.org/10.1099/ijs.0.63532-0
Singh H, Du J, Ngo HTT, Hwa K, Ki W, Kim Y (2015) Pedobacter edaphicus sp nov isolated from forest soil in South Korea. Arch Microbiol 197:781e787. https://doi.org/10.1007/s00203-015-1114-3
Stackebrandt E, Ebers J (2006) Taxonomic parameters revisited: tarnished gold standards. Microbiol Today 33:152–155
Steyn PL, Segers P, Vancanneyt M, Sandra P, Kersters K et al (1998) Classification of heparinolytic bacteria into a new genus, Pedobacter, comprising four species: Pedobacter heparinus comb. nov., Pedobacter piscium comb. nov., Pedobacter africanus sp. nov. and Pedobacter saltans sp. nov. proposal of the family Sphingobacteriaceae fam. nov. Int J Syst Evol Microbiol 48:156–177. https://doi.org/10.1099/00207713-48-1-165
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33–36. https://doi.org/10.1093/nar/28.1.33
Tatusova T, DiCuccio M, Badretdin A, Chetvernin V, Nawrocki EP et al (2016) NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res 44:6614–6624. https://doi.org/10.1093/nar/gkw569
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680. https://doi.org/10.1093/nar/22.22.4673
Ullmann IF, Nygaard AB, Tunsjø HS, Charnock C (2020) Whole genome sequencing and antibiotic diffusion assays, provide new insight on drug resistance in the genus Pedobacter. FEMS Microbiol Ecol. https://doi.org/10.1093/femsec/fiaa088
Urios L, Intertaglia L, Magot M (2013) Pedobacter tournemirensis sp. nov., isolated from a fault water sample of a deep Toarcian argillite layer. Int J Syst Evol Microbiol 63:303–308. https://doi.org/10.1099/ijs.0.038968-0
Viana AT, Caetano T, Covas C, Santos T, Mendo S (2018) Environmental superbugs: the case study of Pedobacter spp. Environ Pollut 241:1048–1055. https://doi.org/10.1016/j.envpol.2018.06.047
Yang DJ, Hong JK (2017) Pedobacter solisilvae sp. nov., isolated from forest soil. Int J Syst Evol Microbiol 67:4814–4819. https://doi.org/10.1099/ijsem.0.002383
Yang Z, Xu J, Sheng M, Qiu J, Zhu J, Zhang J, He J (2020) Pedobacter puniceum sp. nov. Isolated Sludge Curr Microbiol 77:4186–4191. https://doi.org/10.1007/s00284-020-02235-5
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y et al (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Zhang B, Liu ZQ, Zheng YG (2018) Pedobacter quisquiliarum sp. nov., isolated from activated sludge. Int J Syst Evol Microbiol 68:438–442. https://doi.org/10.1099/ijsem.0.002531
Zhou LY, Chen XY, Du ZJ, Mu DS (2019) Pedobacter chinensis sp. nov., a cellulose-decomposing bacterium from Arctic tundra soil. Int J Syst Evol Microbiol 69:1926–1933. https://doi.org/10.1099/ijsem.0.003403
Acknowledgements
Authors thank Professor Aharon Oren for assistance with nomenclature.
Funding
This work was supported by a grant from the National Institute of Biological Resources (NIBR) funded by the Ministry of Environment (MOE) of the Republic of Korea (NIBR201801204). This research was also supported by the Chung-Ang University Young Scientist Scholarship in 2019.
Author information
Authors and Affiliations
Contributions
Chang-Jun Cha designed the study. Material preparation, data collection and analysis were performed by Le Tran Tien Chau and Yong-Seok Kim. The first draft of the manuscript was written by Le Tran Tien Chau and all authors revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare that they do not have any conflicts of interest.
Data availability
The 16S rRNA gene and the whole genome sequences of strain CJ43T that support the findings of this study have been deposited in the GenBank/EMBL/DDBJ database with the accession numbers MK129424 and CP043329, respectively.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chau, L.T.T., Kim, YS. & Cha, CJ. Pedobacter aquae sp. nov., a multi-drug resistant bacterium isolated from fresh water. Antonie van Leeuwenhoek 115, 445–457 (2022). https://doi.org/10.1007/s10482-022-01708-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-022-01708-w