Abstract
Gene flow affects the genetic diversity and structure of tree species and can be influenced by stress related to changing climatic conditions. The study of tree species planted in locations outside their natural range, such as arboreta or botanical gardens, allows us to analyse the effect of severe fragmentation on patterns and distances of gene flow. Paternity analysis based on microsatellite marker genotyping was used to analyse how fragmentation affects gene flow among individuals of Quercus rubra L. distributed in a small isolated group of trees (15 trees) planted in the arboretum on the North Campus of the University of Göttingen. For paternity analysis, 365 seedlings from four seed parents were selected and genotyped using 16 microsatellites. The analysis revealed the majority of pollen (84.89%) originated from trees within the site and identified three large full-sib families consisting of 145, 63 and 51 full-sibs. The average pollen dispersal distance for the four seed parents ranged from 17.3 to 103.6 meters. We observed substantial genetic differentiation among effective pollen clouds of the four seed parents (G’’ST = 0.407) as a result of cross pollination between neighboring trees. No self-fertilization was observed. Gene dispersal via pollen followed the expected distance-dependent pattern, and we observed a significant influx of external pollen (15.11%, ranging from 8.64 to 26.26% for individual seed parents) from a diverse set of donors (30). Long-distance pollen dispersal could explain the presence of significant genetic variation even in isolated natural Q. rubra populations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the face of changing climatic conditions, it is essential to anticipate how different species might cope with associated stresses. Since trees are unable to relocate quickly to more suitable regions, it is crucial to examine and comprehend their behaviour in the context of habitat fragmentation and how gene flow could be impacted in severely fragmented regions. Gene flow (pollen and seed dispersal) is a fundamental factor influencing the genetic diversity and structure of tree species (Ennos 1994). Gene flow by pollen in plant species has been extensively studied using molecular markers such as microsatellites and parentage analysis in open-pollinated seeds (Robledo-Arnuncio and Gil 2005; Bacles and Ennos 2008; Buschbom et al. 2011; Owusu et al. 2016). However, both pollen and seed dispersal mechanisms can be affected by fragmentation and land use. Populations under conditions of rapid change or stress can show reduced fitness causing a decrease in population size and density which directly affects gene flow patterns (Hoffmann and Hercus 2000). Isolation of populations or individuals as a result of land use and climate change can lead to a loss of genetic diversity and even to inbreeding depression (Young et al. 1996; Cole 2003). Although the severe impact of fragmentation on genetic connectivity is evident for some species and aligns with classical population genetic theory, it is not so evident for many other species (Kramer et al. 2008; Lowe et al. 2015). A large number of forest tree species are able to reduce the impact of habitat fragmentation through gene flow via pollen and/or seeds that travel long distances (Robledo-Arnuncio and Gil 2005; Bacles et al. 2006; García et al. 2007; Bacles and Ennos 2008; Piotti et al. 2012), which can maintain connectivity even in severely fragmented and degraded landscapes. The most severely fragmented habitats are often those generated by human activities, where population sizes can be greatly reduced and isolation is drastically increased (Young and Clarke 2000; McKinney 2006; Dubois and Cheptou 2017). The effect of fragmentation also strongly depends on specific life history traits, in particular on pollen and seed dispersal mechanisms. Pollen distribution patterns of insect pollinated woody species tend to be more restricted than those of wind pollinated species in habitats isolated by natural or artificial barriers (Su et al. 2003; Dick et al. 2008).
Molecular markers are a fundamental tool for seed and pollen dispersal studies as they enable the identification of parent trees within a group of mature trees. In numerous studies, molecular markers have enabled the estimation of pollen flow in fragmented habitats, where in several Quercus species a high rate of pollen influx from outside the stand and a considerable pollen dispersal distance have been observed (Dow et al. 1995; Nakanishi et al. 2004; Valbuena-Carabaña et al. 2005; Pluess et al. 2009; Curtu et al. 2009; Gerber et al. 2014; Zeng and Fischer 2020). Studies on pollen (and seed) movement are conducted using e.g. nuclear microsatellite DNA markers, which allow for the identification of parent trees of seeds or seedlings. When the seed parent is unknown, the use of uniparentally inherited markers, such as chloroplast microsatellites, in combination with biparentally inherited markers can enable the identification of both pollen donor and seed parent in the natural regeneration (Ashley 2010). Genotyped seeds assigned to a candidate pollen donor allow the identification of pollen dispersal patterns and distances (Robledo-Arnuncio and Gil 2005; Bacles and Ennos 2008; Moran and Clark 2012; Gerber et al. 2014; Khodwekar and Gailing 2017; Alexander and Woeste 2017; Oyama et al. 2017).
Trees planted outside their natural species range, e.g. in arboreta or botanical gardens, provide an ideal setting to study the effect of severe fragmentation on gene flow patterns and distances, when conspecific trees are absent or very rare (Ihara et al. 1986; Savolainen et al. 2007; Kremer et al. 2010; Sebbenn et al. 2011). Understanding these patterns in fragmented habitats allows us to predict the likely impacts and responses of populations in future situations (Wilson et al. 2016). Non-indigenous species often come from restricted geographical regions and may even originate from seed orchards. In addition, unlike many native species, introduced trees tend to have small populations in their new range (Pettenkofer et al. 2019; Pötzelsberger et al. 2020). Northern red oak (Quercus rubra L.) is a tree species native to the eastern United States and Canada with a very extensive natural range throughout eastern North America where it is often the dominant tree species (Sander 1990). It was introduced to Europe in the 17th century for ornamental purposes and has since become widely distributed throughout the continent. It is often planted as a roadside tree or in parks and gardens (Hickel 1932; Bauer 1953; Nicolescu et al. 2020), and has also been introduced in managed forests as a timber species due to its ability to thrive in a wide range of environments, rapid growth and high quality wood (Sander 1990; Nicolescu et al. 2020). It grows at an average temperature range of 4 to 16 °C and with an average annual rainfall of 760 to 2030 mm; however, it can survive in areas with only 550 –500 mm of average annual rainfall. This gives it a higher potential to adapt to projected climate change, in particular to drought, compared to native European oaks (Bauer 1953; Sander 1990; Major et al. 2013; Vor et al. 2015; Nicolescu et al. 2020).
To analyse the effect of fragmentation on gene flow patterns and distances in Northern red oak, parentage, pollen dispersal and differentiation among effective pollen clouds of seeds from single trees were estimated among isolated individuals of Q. rubra distributed in a small group of planted trees in an arboretum on the North campus of the University of Göttingen. For this purpose, acorns were collected and germinated and seedlings from four known seed parents were selected and genotyped using microsatellites. Once the parental relationships were identified, it was possible to calculate the pollen dispersal distances, directions and reproductive success of each pollen donor. In addition, full-sib families could be identified among the open-pollinated seedlings. In isolated populations, large full-sib families and spatial clustering could lead to increased biparental inbreeding. Such results would highlight potential early negative consequences associated with population fragmentation. We hypothesize, that (1) mating success is distance-dependent resulting in preferential matings between neighboring trees resulting in large full-sib families in this isolated planting and (2) that rare long-distance pollen influx can be observed despite its geographic isolation from conspecific plantings.
Materials and methods
Study area
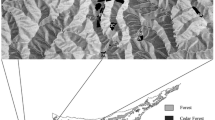
The study area was located in the arboretum at the North Campus of the University of Göttingen, Germany (51°33’26.9"N 9°57’26.1) and also included Q. rubra planted for ornamental purposes outside of the arboretum. A total of 14 Q. rubra (Qr) trees and one individual of the closely related species Quercus coccinea (Qc) were present in the study area. Both species are interfertile and hybridize in natural populations (Sullivan et al. 2016). The trees were planted between 1973 and 1975 and their origin is unknown, with the exception of one individual (Qr_04) which originated from the Morton Arboretum Illinois, USA. Nearby stands and provenance trials of the same species are located at a distance of least 20 km in the Bramwald (Krahl-Urban 1956, 1965). In addition, distributed throughout the city (at > 1 km distance) in gardens and parks we observed a few red oaks that could be potential pollen donors. Thus, a heterogeneous external pollen cloud would indicate significant long-distance pollen influx.
Plant material
Mature acorns of Q. rubra were collected from the ground directly underneath four putative seed parents in 2019, designated as Qr_N, Qr_S, Qr_W and Qr_P (Supplementary material Fig. S1). We performed an exhaustive sampling of the adult trees and collected seeds from four seed parents. We selected seed trees to represent a wide range of distances between seed and pollen parent (ca. 5 to 350 m). Acorns were tested for viability by floating them in water and those that were not viable were removed. The acorns were then kept at 6 °C in humid conditions until the root emerged. Once the radicle appeared, the acorns were transplanted into 24-cell seed trays with a capacity of 295–330 cc. All the acorns were planted and grown in mineral substrate (Kleeschulte Topfsubstrat mineralisch) in the greenhouse of the Department of Forest Genetics and Forest Tree Breeding at the University of Göttingen. The number of acorns sprouted from each mother, Qr_N, Qr_S, Qr_W and Qr_P, was 879, 491, 277 and 351, respectively. Leaf tissue from the 15 adult trees, including the mother trees, and from 1637 germinated seedlings (Supplementary material Table S1) was sampled and stored at -20°C. Male and female flowering was assessed on the lower branches for all trees from April 21 to the end of the flowering period on 28 June 2023. Male flowering was scored on a scale from 0 to 3 (0: no catkins visible, 1: catkins visible, 2: anthers fully developed and opening, 3: catkins withered microsporangia empty), female flowering was scored from 0 to 2 (0: no female flowers visible, 1: receptive female flowers, 2: female flowers no longer receptive).
DNA isolation
Leaf tissue from the 15 adult trees as well as from 365 randomly selected seedlings within each single tree progeny, 92 from Qr_N and 91 of each of the other three mothers (Qr_S, Qr_W and Qr_P) was selected for DNA isolation. DNA extraction was performed with the commercial DNeasy® 96 Plant Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions.
Microsatellite genotyping
To identify chloroplast haplotypes of the 15 adult trees, seven chloroplast simple sequence repeats (cpSSR) were selected from Götz and Gailing (2022) who used both newly developed markers and markers previously adapted for Q. rubra (Zhang et al. 2015; Pettenkofer et al. 2020; Götz et al. 2020). Two of the cpSSRs were universal markers developed for angiosperms (ccmp2 and ccmp4) (Weising and Gardner 1999), three were developed for Quercus species (ucd4, udt1 and udt4) (Deguilloux et al. 2003) and two were developed specifically for Q. rubra (QRcp01 and QRcp28) (Götz and Gailing 2022) (Supplementary material Table S2). From these seven markers, four (ucd4, udt4, QRcp01 and QRcp28) were sufficient to distinguish between haplotypes and were selected to assign seedlings to putative seed parents. Qr_S and Qr_W could not be differentiated as they shared the same chloroplast haplotype.
A total of 25 nuclear microsatellite markers was used to genotype adult trees and seedlings. Ten out of the 25 microsatellite markers were neutral nuclear simple sequence repeats (nSSRs) developed specifically for Q. rubra. Six of the microsatellite markers (quru-GA-0E09, quru-GA-1C06, quru-GA-1F07, quru-GA-0C11, quru-GA-2F05 and quru-GA-2M04) were developed by Aldrich et al. (2002) and the other four (1P10, 2P24, 3A05 and 3D15) by Sullivan et al. (2013). The remaining 15 microsatellite markers (FIR004, FIR013, FIR024, FIR028, FIR031, FIR035, FIR043, FIR104, GOT021, GOT037, GOT040, PIE040, PIE1125, VIT023 and VIT107) were expressed sequence tag-simple sequence repeats (EST-SSRs) developed for Q. robur (Durand et al. 2010) and successfully transferred to Q. rubra (Sullivan et al. 2013) (Supplementary material Table S3).
Polymerase chain reactions (PCR) were performed by combining the microsatellite markers in different multiplex reactions. For cpSSRs, five reactions were carried out. One of the reactions was carried out using a cost-effective tailed primer approach (Schuelke 2000; Kubisiak et al. 2013) (Supplementary file S2).
All PCR reactions were conducted in a Biometra Analytik T professional thermocycler (Jena, Germany) using a touchdown PCR program with the following conditions: initial denaturation at 95 °C for 15 min, followed by 10 cycles of denaturation at 94 °C for 1 min, annealing at 60 °C for 1 min (-1 °C per cycle to 50 °C) and elongation at 72 °C for 1 min. Subsequently, 25 cycles of denaturation at 94 °C for 1 min, annealing at 50 °C for 1 min and elongation at 72 °C for 1 min followed by a final extension at 72 °C for 20 min.
PCR products were separated on an ABI PRISM® 3130 Genetic Analyser and visualized using GeneMapper™ Software v4.1 (Applied Biosystems™). Of the 25 microsatellites, six SSRs (FIR004, FIR024, FIR043, GOT037, PIE040 and VIT107) were excluded from the analysis due to difficulties in amplification and/or scoring. Of the remaining 19, three markers (quru-GA-1C06, quru-GA-2M04 and quru-GA-2F05) showed high null allele frequencies of 0.2593, 0.1307 and 0.2783 and a high number of mismatches of 112, 93 and 79, respectively, based on the preliminary results of the parentage analysis in CERVUS (Supplementary material Tables S4 and S5), and were thus also excluded from further analyses.
Parentage analysis
To differentiate between seed parents with cpSSRs and to validate the maternal origin of seeds collected underneath the seed trees, cpDNA haplotypes were characterized. Seedlings for which it was possible to validate the seed parent based on their haplotype were subsequently included in the paternity analyses using nSSR markers. The two neighboring seed parents, Qr_W and Qr_S shared the same cpDNA haplotype (see below). Thus, it was not possible to unambiguously assign seedlings to either Qr_W or Qr_S. For these seedlings, a parentage analysis with nSSRs was carried out. Seedlings whose seed parent could be identified following the parentage analysis were also included in the paternity analysis.
Parentage and paternity analyses were performed for nSSRs using the maximum likelihood approach implemented in CERVUS v3.0.7 (Marshall et al. 1998), and COLONY v2.0.6.8 (Jones and Wang 2010) was used to identify full-sib families. CERVUS software calculates the LOD scores (logarithm of the likelihood ratio) for each candidate parent-offspring pair over all given loci and calculates the critical delta value which is the difference between the LOD scores of the two most likely candidate parents and which produces a desired level of confidence in the assignments. COLONY uses the individual genotypes to assess sibship and parentage relationships jointly over the set of individuals rather than for pairs of individuals.
Null allele frequencies were estimated in CERVUS based on the iteration of the observed and expected frequencies of the different genotypes. Loci quru-GA-1C06, quru-GA-2M04, GA-2F05 with null allele frequencies equal to or greater than 0.05 were excluded from further analysis resulting in a final set of 16 markers. All the adult trees were considered as candidate pollen parents. A confidence level of 95% was used for the analyses and the critical delta value (> 0.00) was estimated by simulating 10,000 offspring individuals. The input parameters for CERVUS analysis included the proportion of candidate fathers sampled (90%), proportion of genotyped loci of 0.9904 for 16 loci and a basic error rate of 0.01. The maximum number of accepted mismatches was set to three.
COLONY was employed to identify full-sib families. Input parameters for identifying the full-sib families included a confidence level of 95%, female and male polygamy, with inbreeding, without clone, monoecious, diploid, length of run medium, full-likelihood method, high likelihood precision, run specification by default, sibship no prior. The error rate for each locus used in COLONY was estimated by CERVUS.
Pollen dispersal analysis
Allelic diversities (v = (Σi pi)−1) (Crow and Kimura 1970) for each pollen cloud were calculated based on the paternal (haploid) contribution from pollinator trees to all seedlings from each of the four seed parents. The genetic differentiation between the effective pollen contributions to each mother tree was calculated as G’’ST, Hedrick’s standardized GST (2005), further corrected for bias when the number of populations is small (Peakall and Smouse 2012) and as delta, the estimation of differentiation among effective pollen clouds based on the differentiation of each pollen cloud to its complement (Dj) (Gregorius and Roberds 1986).
The analyses were performed using the programs GSED (delta; Dj) (Gillet 2010) v3.0 and GenAlex v6.503 (G’’ST) (Peakall and Smouse 2012). The program input file requires knowledge about alleles coming from the seed and from the pollen parent. In most cases, as the seed parent was known from the chloroplast analysis and the potential pollen parents from the paternity analysis, it was possible to determine which allele came from which parent if the seed parent and pollen donor were not heterozygous for the same alleles. In cases where both parents were known but it was not possible to determine which tree was seed and pollen parent, the tree under which the seed had been collected was assigned as seed parent.
Relatedness of adult trees and seedlings was visualized in a PCoA based on individual-by-individual pairwise genetic distance matrix for codominant data in GenAlex (Peakall et al. 1995, Smouse and Peakall 1999).
Results
Haplotype diversity and seed parent assignment
Analysis with cpSSR markers revealed 5 different haplotypes A.1_2, A.2, A.17, B.4 and A.1_1/ A.5 (Supplementary material Table S6) for the 15 adult trees. A total of 226 out of the 365 seedlings could be assigned to their respective seed parents, 127 to Qr_N and 99 to Qr_P. For the remaining 139 seedlings, it could not be conclusively determined whether the seed tree was Qr_S or Qr_W, based on haplotype results.
nSSR frequency and statistics
The number of alleles per locus ranged from 2 to 19 with an average of 8.625 for the 16 loci. The polymorphic information content (PIC) was 0.528. The combined paternity exclusion probability, the probability of excluding a randomly selected unrelated parent, was 0.9970 for the remaining 16 nSSR loci (Supplementary material Tables S7 and S8).
Parentage analysis
To determine the possible seed parents of the 139 seedlings for which it was not possible to distinguish between putative seed parents Qr_S and Qr_W according to cpSSR markers, a parentage analysis for unknown parents was performed with nSSR markers using CERVUS. In this way, a total of 75 seedlings were assigned to Qr_W and 40 seedlings to Qr_S as seed parents. However, for a group of 24 seedlings it was not possible to distinguish between Qr_S and Qr_W as seed parent because both were seed tree and/or pollen donor of these siblings based on parentage analyses.
Paternity assignment
The 24 seedlings which could not unambiguously be assigned to a single seed parent based on parentage analysis, were assigned to a seed tree based on the location of the seeds when collected. One of these (N_1009) could not be assigned by its seed position under a seed tree, so it was eliminated from subsequent analysis. The remaining 364 seedlings assigned to a single seed parent (226 by cpSSR analysis, 115 by nSSRs parentage analysis and 23 by proximity to the seed tree) were subjected to paternity analysis in order to determine the pollen donor.
Of the 364 seedlings, 309 (84.89%) (Qr_N (111), Qr_P (73) Qr_S (51) and Qr_W (74)) were assigned to a pollen parent and 55 (15.11%) (Qr_N (16), Qr_P (26) Qr_S (6) and Qr_W (7)) were not genetically compatible with any of the genotyped adult trees. Focusing on each of the individual seed parents, 87.40% of the seedlings from Qr_N (111 seedlings) were fathered by the three neighbouring trees Qr_W, Qr_S and Qr_12, and 12.60% were not assigned representing pollen immigration (see Fig. 1).
For Qr_S seedlings, 59.65% (34 out of 57) received pollen from neighbouring Qr_N and 29.82% (17 seedlings) from Qr_W which is also located in the immediate vicinity (Fig. 2). The pollen influx from outside the study area was 10.53%. The offspring of Qr_W were sired by three of the 15 adult trees (Qr_N, Qr_S and Qr_12) with Qr_N being the main pollen donor for 82.72% of the seedlings (67 seedlings). The overall pollen influx from outside the study area was 8.64% (Figs. 1 and 2).
Distribution map of Quercus rubra (Qr) and Q. coccinea (Qc) in the study area on the North Campus of Göttingen University showing the seed parents and the potential pollen donors. Pollen flow between pollen donors and seed parents is indicated by arrows, the arrow size is proportional to the quantity of pollen flow estimated based on paternity analysis using 16 loci
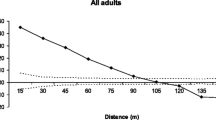
Of the seedlings originating from Qr_P, 73.74% (73 seedlings) were assigned to nine pollen parents (Qr_N, Qr_S, Qr_W, Qr_06, Qr_10, Qr_11, Qr_12, Qr_13 and Qr_14), and for 26.26% (26 seedlings) no pollen parent was identified among the genotyped adult trees. The main pollen donor was Qr_11 (51 seedlings) which is located at a distance of 79.3 m, siring about half of the offspring of Qr_P (51.52%) (Figs. 1 and 2). In 2023, receptiveness of female flowers in the four seed parents was overlapping with male flowering of putative pollen parents. The large pollen contribution of Qr_ 11 to Qr_P (51.2%) at a larger distance of ca. 79 m (Fig. 3) coincides with an overlap of female and male flowering of 20 days in 2023 (Supplementary material Fig. S2).
Principal Coordinates Analysis (PCoA) of Quercus rubra seedlings and mother trees based on genetic distance at 16 nSSRs. The four largest full-sib families and their chloroplast haplotypes are highlighted indicating reciprocal crosses for two families (Qr_N x Qr_W, Qr_N x Qr_S). Seed parents and seedlings have been assigned color-coded labels corresponding to their chloroplast haplotypes. Individuals with haplotype A.17 are represented in yellow, those with haplotype B4 in green, and those with haplotype A1.1 in blue
Relatedness of adult trees (Supplementary material Fig. S3) was not associated with mating patterns (i.e., preferential mating between related trees, for example due to similar phenology) (data not shown).
Family structure
Based on kinship analyses in COLONY and paternity analyses in CERVUS, four larger full-sib families were identified as visualized by the clustering of groups according to genetic distance, which clearly separated three of the four families (Fig. 4). One full-sib family of 145 individuals from the parent pair Qr_N and Qr_W, was identified with a probability of inclusion of 0.9999, consisting of 78 seedlings with Qr_N as the seed parent and for 67 seedlings with Qr_W as the seed parent. Another full-sib family consisted of 63 seedlings with a probability of inclusion of 0.9989 (Qr_N x Qr_S: 29 seedlings; Qr_S x Qr_N: 35 seedlings). A third family of 51 seedlings had Qr_P as seed and Qr_11 as pollen parent with a probability of inclusion of 0.9996. Finally, a fourth full-sib family consisted of 24 seedlings with an inclusion probability of 1, but it was not possible to determine which of the two parents (Qr_W, Qr_S) was seed or pollen parent. In addition, four smaller full-sib families were identified (Supplementary material Table S9).
COLONY can derive the genotype of the potential pollen parent even if it is not among the sampled potential candidates. Thirty unsampled parents were inferred by COLONY as potential external pollen parents, 17 of which sired more than one seedling, while 13 contributed pollen to one seedling (Supplementary material Fig. S4). Six of the external pollen donors fertilised more than one tree, but always with the pollen receptors Qr_N, Qr_W or Qr_S (Supplementary material Fig. S5). The unknown pollen donors for the neighboring trees Qr_N, Qr_W and Qr_S were different from the Qr_P donors. No donors were found to be common to all four seed trees (Supplementary material Fig. S5).
Variation within and among effective pollen clouds
Allelic diversities were calculated for effective pollen clouds of each seed tree, the pooled pollen clouds for all loci and on average (Supplementary material Table S10). Qr_P received pollen from 23 donors, Qr_N from 16, Qr_W from 10 and Qr_S only from six donors. According to the results of allelic diversity in each effective pollen cloud, Qr_P and Qr_N showed the highest diversity (2.864, 2.733), followed by Qr_S (2.370) and Qr_W (1.911).
G’’ST and delta values were calculated to determine the genetic differentiation among pollen contributions to each seed tree. For G’’ST a value of 0.407 was obtained, while a delta value of 0.382 was found (Table 1). The genetic differentiation of the four effective pollen clouds (Dj) to their complement ranged from 0.272 to 0.428. Qr_P received pollen from nine different known pollen donors and its genetic differentiation (Dj = 0.369) was slightly lower than that of Qr_N (Dj = 0.411) and Qr_W (Dj = 0.428). The effective pollen cloud of the seed tree Qr_S showed the least genetic differentiation (Dj = 0.272).
Distance dependent pollen dispersal
The percentage of seedlings fathered by each potential pollen parent plotted against the distance of the pollen donors to the seed parents (Fig. 3) decreased rapidly with distance. Most pollen was effective over small distances up to 40.8 m. Pollen migration distances ranged from 14.9 to 356.0 m within the study area (Supplementary material Table S11). Average pollen dispersal distance was different for each seed tree, with an average distance of 23.7 m for Qr_N, 20.5 m for Qr_W, 17.3 m for Qr_S and 103.6 m for Qr_P. However, the amount of pollen contributed by each pollen parent to seed parents was different in each case. Most of the pollen donors of Qr_N (84.25%) were located in an area within a radius of 14.9–50.0 m around the seed parent, 3.15% of the pollen came from a pollen donor located between 200 and 250 m and 12.60% of the pollen came from outside the study area (Supplementary material Table S11). Most of the pollen parents that contributed to the offspring of Qr_P (58.59%) were located within a radius between 50.0 and 100.0 m. However, trees that were nearer to Qr_P, like Qr 6, fathered only 6.06% of the seedlings. In the case of the seed parent Qr_S, pollen donors within the area were the nearest trees Qr_N (59.65%) and Qr_W (29.82%) at distances of 14.9 and 22.2 m. With respect of the seed parent Qr_W, the most successful pollen parent (Qr_N with 82.72%) was located in the direct vicinity of Qr_W (17.8 m). It also received pollen from Qr_S (7.41%) situated 22.2 m and even 1.23% of pollen from one tree located 187.1 m away (Qr_12).
Discussion
Gene flow and mating system
Knowledge of gene flow in severely fragmented landscapes is particularly important to estimate the level of “tolerable” fragmentation in the face of anthropogenic land use and the dramatic climate change that will be experienced in the coming years. However, only a few studies have estimated gene flow in Q. rubra in isolated habitats (Moran and Clark 2012; Alexander and Woeste 2017). In our study, we estimated patterns of gene flow outside the species’ native range within an arboretum with 15 interfertile trees isolated from conspecific plantations and calculated the frequency of pollen influx. Nearby stands and provenance trials of the same species were separated from the arboretum by at least 20 km, mostly located in the Bramwald (Krahl-Urban 1956, 1965). In addition, distributed throughout the city in gardens and parks we observed a few red oaks that could be potential pollen donors.
Low population density affects the mating system, increasing reproduction between neighbours and affects gene flow, which may be limited due to a decrease in the number of pollen donors (Young et al. 1996; Finkeldey 2002; Lowe et al. 2005). The application of paternity analysis in this isolated group of Q. rubra trees revealed gene flow patterns within the plot and pollen influx from outside. It showed a significant relationship between pollination frequency and the distance of the seed parent to the pollen donors, showing that the closest neighbours provided the majority of the effective pollen, which has been frequently observed in wind-pollinated trees, such as oaks (Streiff et al. 1999; Lind and Gailing 2013; Zhang et al. 2015; Alexander and Woeste 2017). However, pollen immigration (15.11%) was comparatively high in this isolated small plantation outside the species’ native range. Only nine out of the 15 adult trees acted as pollen donors, but they contributed between 73.74% and 91.36% of pollen to each of the four seed parents. Preferential mating between neighboring trees and even lower pollen immigration of 7% was observed by Alexander and Woeste (2017) within a Q. rubra seed orchard within the natural distribution range of the species. Most of the pollen contribution observed in our study occurred within a radius ranging from 17.3 m to 103.6 m, with a mean distance of 40.8 m. This result is consistent with the mean distance of pollination in Q. rubra observed in other studies, where the mean distance ranged, for example, from 40 to 60 m in a study conducted in a Q. rubra seed orchard (Alexander and Woeste 2017). Mean pollen flow distances of 57 m and 89 m were observed in two mixed red oak forests with a low tree density due to anthropogenic interventions (Moran and Clark 2012). Studies on other Quercus species showed similar results in terms of pollination distances regardless of whether the trees were distributed in isolated forest patches or in continuous forests. For example, a study on 62 adult Q. macrocarpa trees distributed on an old farm estimated that the mean distance of pollen flow ranged from 64 to 80 m (Dow and Ashley 1998). Other studies in Q. robur and Q. petraea in a continuous forest showed a similar mean pollen dispersal distance for both species ranging from 22 to 65 m depending on the family (Streiff et al. 1999). Another example is found in Q. salicina (Nakanishi et al. 2004) where studies conducted in a mixed forest showed a mean pollination distance of 67 m and the study conducted in Quercus lobata in a mixed oak savannah showed a range of pollination distances between 69 and 151 m depending on the seed tree (Pluess et al. 2009). Albeit pollination distance was different depending on the seed parent from which the seeds originated, the mean pollination distance observed in the study area was comparable to distances recorded in other studies, regardless of whether those studies were carried out in continuous forests or in forest patches, indicating that, independent of habitat type, the range of distances over which pollination occurs tends to be consistent for Quercus spp.
The considerable amount of immigrant pollen (15.11%; from 8.64 to 26.26% for individual seed parents) from an estimated number of 30 external pollen donors, showed that there is significant long-distance gene flow with the nearest plantation being more than 20 km apart. In contrast, the percentage of effective immigrant pollen was much higher in the species’ native range with comparatively high conspecific tree density, e.g. in managed natural Q. rubra populations ranging from 59 to 64% (Moran and Clark 2012). As expected, our estimates of pollen influx were also lower than in other studies conducted in natural populations on different Quercus species. For example, external pollen constituted 57% in an isolated forest patch of Q. macrocarpa, where the nearest tree from outside the study area was at least 100 m away (Dow and Ashley 1998). Pollen flow from outside the study area in Q. petrea and Q. pyrenaica represented 38% and 34%, respectively, in a heterogeneous stand, where both species were scattered in low to medium density patches (Valbuena-Carabaña et al. 2005) or between 18 and 24% for Q. frainetto and 31% for Q. pubescens in a continuous mixed oak forest (Curtu et al. 2009). Thus overall in oaks although the level of pollen immigration can be quite low (2.6% for Q. robur in Moracho et al. 2016), pollen immigration levels can also be surprisingly high (50% or more in cases reviewed by Ashley 2021). Furthermore, reported dispersal distances for pollen of oak species have also varied greatly, ranging from 16 m to app. 5400 m (Gerber et al. 2014) for four European white oak species, to between 1000 and 8500 m for pedunculate oak (Q. robur) refugial populations (Moracho et al. 2016), and even over 80 km for a small isolated Q. robur stand (Buschbom et al. 2011).
In seed orchards or in very isolated populations where many of the trees shed pollen at the same time, a preferential contribution of pollen from nearby trees is likely to occur and the contribution of external pollen will be limited. However, studies conducted in different seed orchards, have demonstrated that the extent of external pollen contamination varies across studies. For instance, two seed orchards of Eucalyptus species reported external contamination levels of approximately 4.5% and 14.7%, respectively (Grosser et al. 2010; Gonzaga et al. 2016), while in a separate study involving pine seed orchards, the external pollen contamination reached was up to 52% (Torimaru et al. 2009). The notable amount of external pollen flow observed in our study into the strongly isolated Q. rubra planting could account for the considerable genetic diversity observed even in peripheral populations of Q. rubra (Lind and Gailing 2013; Götz et al. 2020). In addition, the fertilization by external pollen was different between trees, as was the number of pollen donors to each seed parent. Of the 30 external pollen donors, we observed 14 that were exclusively pollen donors to Qr_P. Therefore, Qr_P received a higher and more diverse amount of external pollen than any of the other seed parents. Immigrant pollen pools also varied significantly among seed trees in a mixed Quercus robur and Q. petraea stand in northern Poland (Chybicki and Burczyk 2013). The observed variations in pollen donor contributions may be attributed, among other factors, to the different flowering times and intensity of one particular tree compared to the rest of the trees in the arboretum. This difference in flowering time probably resulted in a scenario in which the Qr_P tree was receptive precisely at the time when there was a greater amount of external pollen in the surrounding environment. Thus, reproductive success will be influenced by the flowering time and intensity of both the pollen donor and the seed parent and this timing also may be modified from year to year depending on environmental factors (Alexander and Woeste 2016, 2017). However, based on our observations, bud burst as proxy for flowering time (Chesnoiu et al. 2009) occurred approximately at the same time in the year of seed collection. Male and female flowering is more difficult to assess, and due to the two-year seed maturation in Q. rubra, it would had needed to be assessed one year preceding the seed collection (in 2018). While phenological ranks in oaks are often stable from year to year (e.g., Gailing et al. 2003), our assessment of the onset and duration of male and female flowering in 2023 did not explain potential mating preferences among groups of neighboring trees or the large amount of different external pollen parent contributing to the progeny of Qr_P (see above). Also, other factors such as the distribution of individual trees within the study area, for example, the isolation of some trees from others due to physical barriers, could affect gene flow patterns. In addition, the landscape included artificial structures, such as buildings, which can further hinder the pollination process and consequently increase the observed distances over which pollination occurs. This, in turn, can have a significant impact on the amount of pollen accessible to each tree (Su et al. 2003; Piana et al. 2019).
Genetic differentiation among effective pollen clouds
Our study area represents a highly fragmented habitat with a small number of individuals, where we observed high genetic differentiation, among the four effective pollen clouds of Qr_N, Qr_S, Qr_W and Qr_P (G’’ST = 0.407, δ = 0.382 and Dj ranged from 0.272 to 0.428). Genetic differentiation among the four effective pollen clouds was considerably higher than the genetic differentiation among pollen clouds frequently observed in oak species in natural populations, GST= 0.01 among Q. robur and Q. petrea, ΦFT = 0.034 in Q. salicina, FST= 0.035–0.054 in Q. robur, FST= 0.149 and ΦFT = 0.381 in isolated Q. castanea (Streiff et al. 1999; Nakanishi et al. 2004; Buschbom et al. 2011) as well as in other temperate forest tree species that are wind-pollinated such as Picea (GST= 0.001), Fraxinus (Φ = 0.08) or Pinus (FST ≤ 0.001) species (Finkeldey 1995; Bacles and Ennos 2008; Sousa et al. 2010; Jiménez-Ramírez et al. 2021). As a consequence of the decrease in the effective number of pollen donors, the differentiation among effective pollen clouds of each single seed parent increases (Finkeldey and Hattemer 2007). The high differentiation among pollen clouds observed in the present study is related to the selection of neighboring intermating individuals as seed trees in a small and isolated planting. A high genetic differentiation among effective pollen clouds is also frequently observed in insect pollinated tropical tree species that occur in low density (Ahmed et al. 2009; Sebbenn et al. 2011; Owusu et al. 2016; Cristóbal-Pérez et al. 2021), but has not been observed in highly fragmented natural populations of temperate wind-pollinated trees species (Finkeldey 1995; Sousa et al. 2010; Gonzaga et al. 2016). However, unlike in tropical forests where the rate of selfing in low density populations is significant (Aldrich and Hamrick 1998; Finkeldey and Hattemer 2007; Dick et al. 2008) no selfing was observed in any of the seed parents which indicates that Q. rubra is an exclusively outcrossing species even when the habitat is severely fragmented (Schwarzmann and Gerhold 1991; Ducousso et al. 1993). While the genetic differentiation among effective pollen clouds was high, genetic differentiation in the offspring was reduced as result of predominant reciprocal gene flow between two neighboring seed parents. For example, Qr_N and Qr_W were both seed trees and the main pollen donors to each other. Consequently, the genetic differentiation between pairwise effective pollen clouds was high (PG’’ST = 0.603), the offspring exhibited low genetic differentiation (OPG’’ST = 0.048) and among the 208 analysed seedlings (81 from Qr_W and 127 from Qr_N) were 145 full-siblings (see Fig. 4). The significant differentiation observed between effective pollen clouds and the abundance of full-sibs in the pedigree reflects that matings preferentially occur between neighbouring individuals. In subsequent generations, this could result in mating between closely related trees, which, if gene influx is restricted, will lead to biparental inbreeding (Uyenoyama 1986) and possibly to strong inbreeding depression (Young et al. 1996; Stacy 2001; Finkeldey and Hattemer 2007; Zhang et al. 2015; Cristóbal-Pérez et al. 2021). Similar effects are expected in strongly isolated marginal natural populations at the rear and leading edge (Kremer et al. 2014; Vakkari et al. 2020; Götz et al. 2022). These populations can potentially face a limitation of pollen flow, which, in turn, could increase the probability of inbreeding within the populations. The presence of a significant number of full-sibs and the subsequent occurrence of biparental inbreeding could serve as early indicators of the negative consequences associated with population fragmentation.
Finally, oaks, being long-lived trees, have a long maturation period of about 25 years before they start flowering and producing seeds. This long period is a challenge when attempting controlled crosses in large forest trees with irregular flowering patterns and inconsistent seed production. Isolated plantings outside the native range with a limited number of interbreeding individuals offer the possibility to identify full-sib families for quantitative genetic and QTL (Quantitative Trait Locus) mapping studies.
References
Ahmed S, Compton SG, Butlin RK, Gilmartin PM (2009) Wind-borne insects mediate directional pollen transfer between desert fig trees 160 kilometers apart. Proc Natl Acad Sci 106:20342–20347. https://doi.org/10.1073/pnas.0902213106
Aldrich PR, Hamrick JL (1998) Reproductive dominance of pasture trees in a fragmented tropical forest mosaic. Science 281:103–105. https://doi.org/10.1126/science.281.5373.103
Aldrich PR, Michler CH, Sun W, Romero-Severson J (2002) Microsatellite markers for northern red oak (Fagaceae: Quercus rubra). Mol Ecol Notes 2:472–474. https://doi.org/10.1046/j.1471-8286.2002.00282.x
Alexander L, Woeste K (2016) Phenology, dichogamy, and floral synchronization in a northern red oak (Quercus rubra) seed orchard. Can J Res 46:629–636. https://doi.org/10.1139/cjfr-2015-0312
Alexander L, Woeste K (2017) Pollen gene flow, male reproductive success, and genetic correlations among offspring in a northern red oak (Quercus rubra L.) seed orchard. PLoS ONE 12:e0171598. https://doi.org/10.1371/journal.pone.0171598
Ashley MV (2010) Plant parentage, pollination, and dispersal: how DNA microsatellites have altered the landscape. Crit Rev Plant Sci 29:148–161. https://doi.org/10.1080/07352689.2010.481167
Ashley MV (2021) Answers blowing in the wind: a quarter century of genetic studies of pollination in oaks. Forests 12(5):575. https://doi.org/10.3390/f12050575
Bacles CFE, Ennos RA (2008) Paternity analysis of pollen-mediated gene flow for Fraxinus excelsior L. in a chronically fragmented landscape. Heredity 101:368–380. https://doi.org/10.1038/hdy.2008.66
Bacles CFE, Lowe AJ, Ennos RA (2006) Effective seed dispersal across a fragmented landscape. Science 311:628–628. https://doi.org/10.1126/science.1121543
Bauer F (1953) Northern red oak. Sauerländer, Frankfurt am Main
Buschbom J, Yanbaev Y, Degen B (2011) Efficient long-distance gene flow into an isolated relict oak stand. J Hered 102:464–472. https://doi.org/10.1093/jhered/esr023
Chesnoiu EN, Șofletea N, Curtu AL et al (2009) Bud burst and flowering phenology in a mixed oak forest from Eastern Romania. Annals for Res 52:199–206. https://doi.org/10.15287/afr.2009.136
Chybicki IJ, Burczyk J (2013) Seeing the forest through the trees: comprehensive inference on individual mating patterns in a mixed stand of Quercus robur and Q. petraea. Ann Botany 112:561–574. https://doi.org/10.1093/aob/mct131
Cole CT (2003) Genetic variation in rare and common plants. Annu Rev Ecol Evol Syst 34:213–237. https://doi.org/10.1146/annurev.ecolsys.34.030102.151717
Cristóbal-Pérez EJ, Fuchs EJ, Martén-Rodríguez S, Quesada M (2021) Habitat fragmentation negatively affects effective gene flow via pollen, and male and female fitness in the dioecious tree, Spondias purpurea (Anacardiaceae). Biol Conserv 256:109007. https://doi.org/10.1016/j.biocon.2021.109007
Crow JF, Kimura M (1970) An introduction to Population Genetics Theory. Harper and Row, New York, Evanston, London
Curtu AL, Gailing O, Finkeldey R (2009) Patterns of contemporary hybridization inferred from paternity analysis in a four-oak-species forest. BMC Evol Biol 9:284. https://doi.org/10.1186/1471-2148-9-284
de Sousa SN, Gailing O, Finkeldey R et al (2010) Reproduction system of Norway spruce (Picea abies [L.] Karst.) In pure and mixed stands with different density. Forstarchiv 81:218–227
Deguilloux MF, Dumolin-Lapègue S, Gielly L et al (2003) A set of primers for the amplification of chloroplast microsatellites in Quercus. Mol Ecol Notes 3:24–27. https://doi.org/10.1046/j.1471-8286.2003.00339.x
Dick CW, Hardy OJ, Jones FA, Petit RJ (2008) Spatial scales of pollen and seed-mediated gene flow in tropical rain forest trees. Trop Plant Biol 1:20–33. https://doi.org/10.1007/s12042-007-9006-6
Dow B, Ashley M (1998) High levels of gene flow in bur oak revealed by paternity analysis using microsatellites. J Hered 89:62–70. https://doi.org/10.1093/jhered/89.1.62
Dow BD, Ashley MV, Howe HF (1995) Characterization of highly variable (GA/CT) microsatellites in the bur oak, Quercus macrocarpa. Theor Appl Genet 91:137–141. https://doi.org/10.1007/BF00220870
Dubois J, Cheptou P-O (2017) Effects of fragmentation on plant adaptation to urban environments. Philos Trans R Soc B Biol Sci 372:20160038. https://doi.org/10.1098/rstb.2016.0038
Ducousso A, Michaud H, Lumaret R (1993) Reproduction and gene flow in the genus Quercus L. Ann Sci for 50:91s–106s. https://doi.org/10.1051/forest:19930708
Durand J, Bodénès C, Chancerel E et al (2010) A fast and cost-effective approach to develop and map EST-SSR markers: oak as a case study. BMC Genomics 11:570. https://doi.org/10.1186/1471-2164-11-570
Ennos RA (1994) Estimating the relative rates of pollen and seed migration among plant populations. Heredity 72:250–259. https://doi.org/10.1038/hdy.1994.35
Finkeldey R (1995) Homogeneity of pollen allele frequencies of single seed trees in Picea abies (L.) Karst. Plantations. Heredity 74:451–463. https://doi.org/10.1038/hdy.1995.66
Finkeldey R (2002) Reproduction in continuous cover forests — the geneticist’s perspective. In: von Gadow K, Nagel J, Saborowski J (eds) Continuous cover forestry. Managing Forest ecosystems. Springer, Springer, Dordrecht. https://doi.org/10.1007/978-94-015-9886-6_6
Finkeldey R, Hattemer HH (2007) Tropical forest genetics. Springer Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-37398-8
Gailing O, Wachter H, Leinemann L, Hosius B et al (2003) Charakterisierung Verschiedener Ursprungsgebiete Der Späteiche (Quercus robur L.) mit Hilfe Von DNS-Markern aus dem Chloroplastengenom. Allgemeine Forst- und Jagdzeitung 174:227–231
García C, Jordano P, Godoy JA (2007) Contemporary pollen and seed dispersal in a Prunus mahaleb population: patterns in distance and direction. Mol Ecol 16:1947–1955. https://doi.org/10.1111/j.1365-294X.2006.03126.x
Gerber S, Chadœuf J, Gugerli F et al (2014) High rates of gene flow by pollen and seed in oak populations across Europe. PLoS ONE 9:e85130. https://doi.org/10.1371/journal.pone.0085130
Gillet EM (2010) GSED Version 3.0. https://www.uni-goettingen.de/de/95607.html
Gonzaga JMS, Manoel RO, Sousa ACB et al (2016) Pollen contamination and non-random mating in a Dehnh seedling seed orchard. Silvae Genet 65:1–11. https://doi.org/10.1515/sg-2016-0001
Götz J, Gailing O (2022) A novel set of polymorphic chloroplast microsatellite markers for northern red oak (Q. rubra L). Plant Genet Resour Charact Util 20:174–177. https://doi.org/10.1017/S1479262122000156
Götz J, Krutovsky KV, Leinemann L et al (2020) Chloroplast haplotypes of northern red oak (Quercus rubra L.) stands in Germany suggest their origin from northeastern Canada. Forests 11:1025. https://doi.org/10.3390/f11091025
Götz J, Rajora OP, Gailing O (2022) Genetic structure of natural northern range-margin mainland, peninsular, and island populations of northern red oak (Quercus rubra L). Front Ecol Evol 10:907414. https://doi.org/10.3389/fevo.2022.907414
Gregorius H-R, Roberds JH (1986) Measurement of genetical differentiation among subpopulations. Theor Appl Genet 71:826–834. https://doi.org/10.1007/BF00276425
Grosser C, Potts BM, Vaillancourt RE (2010) Microsatellite based paternity analysis in a clonal Eucalyptus nitens seed orchard. Silvae Genet 59:57–62. https://doi.org/10.1515/sg-2010-0007
Hedrick PW (2005) A standardized genetic differentiation measure. Evolution 59:1633–1638. https://doi.org/10.1111/j.0014-3820.2005.tb01814.x
Hickel R (1932) Dendrologie forestière. Publ P Lechevalier. Paris
Hoffmann AA, Hercus MJ (2000) Environmental stress as an evolutionary force. Bioscience 50:217–226. https://doi.org/10.1641/0006-3568(2000)050[0217:ESAAEF]2.3.CO;2
Ihara M, Gadrinab LU, Siregar UJ, Iyama S (1986) Genetic control of alcohol dehydrogenase and estimation of some population parameters in Hopea odorata Roxb. (Dipterocarpaceae) 61:127–136. https://doi.org/10.1266/jjg.61.127
Jiménez-Ramírez A, Grivet D, Robledo-Arnuncio JJ (2021) Measuring recent effective gene flow among large populations in Pinus sylvestris: local pollen shedding does not preclude substantial long-distance pollen immigration. PLoS ONE 16:e0255776. https://doi.org/10.1371/journal.pone.0255776
Jones OR, Wang J (2010) COLONY: a program for parentage and sibship inference from multilocus genotype data. Mol Ecol Resour 10:551–555. https://doi.org/10.1111/j.1755-0998.2009.02787.x
Khodwekar S, Gailing O (2017) Evidence for environment-dependent introgression of adaptive genes between two red oak species with different drought adaptations. Am J Bot 104:1088–1098. https://doi.org/10.3732/ajb.1700060
Krahl-Urban J (1956) Über Eichen Provenienzversuche (Erster Bericht über anlage und vorläufige Ergebnisse Meiner Versuchsflächen). Lehrforstamt Bramwald Hemelni Hann Münden
Krahl-Urban J (1965) North American deciduous species; purpose and results of study trip. Forstarchiv 36(5):93–97
Kramer AT, Ison JL, Ashley MV, Howe HF (2008) The paradox of forest fragmentation genetics. Conserv Biol 22:878–885. https://doi.org/10.1111/j.1523-1739.2008.00944.x
Kremer A, Le Corre V, Petit RJ, Ducousso A (2010) Historical and contemporary dynamics of adaptive differentiation in European oaks. Molecular approaches in natural resource conservation. Cambridge University Press, Cambridge. https://doi.org/10.1017/CBO9780511777592.006
Kremer A, Potts BM, Delzon S (2014) Genetic divergence in forest trees: understanding the consequences of climate change. Funct Ecol 28:22–36. https://doi.org/10.1111/1365-2435.12169
Kubisiak TL, Nelson CD, Staton ME et al (2013) A transcriptome-based genetic map of Chinese chestnut (Castanea mollissima) and identification of regions of segmental homology with peach (Prunus persica). Tree Genet Genomes 9:557–571. https://doi.org/10.1007/s11295-012-0579-3
Lind JF, Gailing O (2013) Genetic structure of Quercus rubra L. and Quercus ellipsoidalis E. J. Hill populations at gene-based EST-SSR and nuclear SSR markers. Tree Genet Genomes 9:707–722. https://doi.org/10.1007/s11295-012-0586-4
Lowe AJ, Boshier D, Ward M et al (2005) Genetic resource impacts of habitat loss and degradation; reconciling empirical evidence and predicted theory for neotropical trees. Heredity 95:255–273. https://doi.org/10.1038/sj.hdy.6800725
Lowe AJ, Cavers S, Boshier D et al (2015) The resilience of forest fragmentation genetics no longer a paradox. We were just looking in the wrong place. Heredity 115:97–99. https://doi.org/10.1038/hdy.2015.40
Major KC, Nosko P, Kuehne C et al (2013) Regeneration dynamics of non-native northern red oak (Quercus rubra L.) populations as influenced by environmental factors: a case study in managed hardwood forests of southwestern Germany. For Ecol Manag 291:144–153. https://doi.org/10.1016/j.foreco.2012.12.006
Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Mol Ecol 7:639–655. https://doi.org/10.1046/j.1365-294x.1998.00374.x
McKinney ML (2006) Urbanization as a major cause of biotic homogenization. Biol Conserv 127:247–260. https://doi.org/10.1016/j.biocon.2005.09.005
Moracho E, Moreno G, Jordano P, Hampe A (2016) Unusually limited pollen dispersal and connectivity of pedunculate oak (Quercus robur) refugial populations at the species’ southern range margin. Mol Ecol 25(14):3319–3333. https://doi.org/10.1111/mec.13692
Moran EV, Clark JS (2012) Between-site differences in the scale of dispersal and gene flow in red oak. PLoS ONE 7:e36492. https://doi.org/10.1371/journal.pone.0036492
Nakanishi A, Tomaru N, Yoshimaru H et al (2004) Patterns of pollen flow and genetic differentiation among pollen pools in Quercus salicina in a warm temperate old–growth evergreen broad-leaved forest. Silvae Genet 53:258–264. https://doi.org/10.1515/sg-2004-0048
Nicolescu V-N, Vor T, Mason WL et al (2020) Ecology and management of northern red oak (Quercus rubra L. syn. Q. borealis F. Michx.) In Europe: a review. Int J Res 93:481–494. https://doi.org/10.1093/forestry/cpy032
Owusu SA, Schlarbaum SE, Carlson JE, Gailing O (2016) Pollen gene flow and molecular identification of full-sib families in small and isolated population fragments of Gleditsia triacanthos L. Botany 94:523–532. https://doi.org/10.1139/cjb-2015-0244
Oyama K, Herrera-Arroyo ML, Rocha-Ramírez V et al (2017) Gene flow interruption in a recently human-modified landscape: the value of isolated trees for the maintenance of genetic diversity in a Mexican endemic red oak. For Ecol Manag 390:27–35. https://doi.org/10.1016/j.foreco.2017.01.018
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Peakall R et al (1995) Evolutionary implications of allozyme and RAPD Variation in diploid populations of dioecious buffalograss Buchloe dactyloides. Molecular Ecology 4: 135–147. https://doi.org/10.1111/j.1365-294X.1995.tb00203.x
Pettenkofer T, Burkardt K, Ammer C (2019) Genetic diversity and differentiation of introduced red oak (Quercus rubra) in Germany in comparison with reference native North American populations. Eur J For Res 138:275–285. https://doi.org/10.1007/s10342-019-01167-5
Pettenkofer T, Finkeldey R, Müller M et al (2020) Genetic variation of introduced red oak (Quercus rubra) stands in Germany compared to north American populations. Eur J Res 139:321–331. https://doi.org/10.1007/s10342-019-01256-5
Piana MR, Aronson MF, Pickett ST, Handel SN (2019) Plants in the city: understanding recruitment dynamics in urban landscapes. Front Ecol Environ 17:455–463. https://doi.org/10.1002/fee.2098
Piotti A, Leonardi S, Buiteveld J et al (2012) Comparison of pollen gene flow among four European beech (Fagus sylvatica L.) populations characterized by different management regimes. Heredity 108:322–331. https://doi.org/10.1038/hdy.2011.77
Pluess A, Sork V, Dolan B et al (2009) Short distance pollen movement in a wind-pollinated tree, Quercus lobata (Fagaceae). For Ecol Manag 288:735–744. https://doi.org/10.1016/j.foreco.2009.05.014
Pötzelsberger E, Spiecker H, Neophytou C et al (2020) Growing non-native trees in European forests brings benefits and opportunities but also has its risks and limits. Curr Rep 6:339–353. https://doi.org/10.1007/s40725-020-00129-0
Robledo-Arnuncio JJ, Gil L (2005) Patterns of pollen dispersal in a small population of Pinus sylvestris L. revealed by total exclusion paternity analysis. Heredity 94:13–22. https://doi.org/10.1038/sj.hdy.6800542
Sander IL (1990) Northern red oak (Quercus rubra L). In: Burns RM, Honkala BH (eds) Silvics of North America. Forest Service, Agriculture Handbook, USA
Savolainen O, Pyhäjärvi T, Knürr T (2007) Gene flow and local adaptation in trees. Annu Rev Ecol Evol Syst 38:595–619. https://doi.org/10.1146/annurev.ecolsys.38.091206.095646
Schuelke M (2000) An economic method for the fluorescent labelling of PCR fragments. Nat Biotechnol 18:233–234. https://doi.org/10.1038/72708
Schwarzmann JF, Gerhold HD (1991) Genetic structure and mating system of northern red oak (Quercus rubra L.) in Pennsylvania. Sci 37:1376–1389. https://doi.org/10.1093/forestscience/37.5.1376
Sebbenn AM, Carvalho ACM, Freitas MLM et al (2011) Low levels of realized seed and pollen gene flow and strong spatial genetic structure in a small, isolated and fragmented population of the tropical tree Copaifera langsdorffii Desf. Heredity 106:134–145. https://doi.org/10.1038/hdy.2010.33
Smouse PE, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82:561–573. https://doi.org/10.1046/j.1365-2540.1999.00518.x
Stacy EA (2001) Cross-fertility in two tropical tree species: evidence of inbreeding depression within populations and genetic divergence among populations. Am J Bot 88:1041–1051. https://doi.org/10.2307/2657086
Streiff R, Ducousso A, Lexer C et al (1999) Pollen dispersal inferred from paternity analysis in a mixed oak stand of Quercus robur L. and Q. petraea. (Matt) Liebl Mol Ecol 8:831–841. https://doi.org/10.1046/j.1365-294X.1999.00637.x
Su H, Qu L-J, He K et al (2003) The Great Wall of China: a physical barrier to gene flow? Heredity 90:212–219. https://doi.org/10.1038/sj.hdy.6800237
Sullivan AR, Lind JF, McCleary TS et al (2013) Development and characterization of genomic and gene-based microsatellite markers in north American red oak species. Plant Mol Biol Rep 31:231–239. https://doi.org/10.1007/s11105-012-0495-6
Sullivan AR, Owusu SA, Weber JA, Hipp AL, Gailing O (2016) Hybridization and divergent selection in multispecies oak communities. Bot J Linn Soc 181:99–114. https://doi.org/10.1111/boj.12393
Torimaru T, Wang X-R, Fries A et al (2009) Evaluation of pollen contamination in an advanced scots pine seed orchard. Silvae Genet 58:262–269. https://doi.org/10.1515/sg-2009-0033
Uyenoyama MK (1986) Inbreeding and the cost of meiosis: the evolution of selfing in populations practicing biparental inbreeding. Evolution 40:388–404. https://doi.org/10.1111/j.1558-5646.1986.tb00479.x
Vakkari P, Rusanen M, Heikkinen J et al (2020) Patterns of genetic variation in leading-edge populations of Quercus robur: genetic patchiness due to family clusters. Tree Genet Genomes 16:73. https://doi.org/10.1007/s11295-020-01465-9
Valbuena-Carabaña M, González-Martínez SC, Sork VL et al (2005) Gene flow and hybridisation in a mixed oak forest (Quercus pyrenaica Willd. and Quercus petraea (Matts.) Liebl.) in central Spain. Heredity 95:457–465. https://doi.org/10.1038/sj.hdy.6800752
Vor T, Spellmann H, Bolte A, Ammer C (2015) Potentials and risks of introduced tree species. Göttinger Forstwissenschaften 7:296. Germany
Weising K, Gardner RC (1999) A set of conserved PCR primers for the analysis of simple sequence repeat polymorphisms in chloroplast genomes of dicotyledonous angiosperms. Genome 42:9–19. https://doi.org/10.1139/g98-104
Wilson MC, Chen X-Y, Corlett RT et al (2016) Habitat fragmentation and biodiversity conservation: key findings and future challenges. Landsc Ecol 31:219–227. https://doi.org/10.1007/s10980-015-0312-3
Young AG, Clarke GM (2000) Introduction: genetics, demography and the conservation of fragmented populations. In: Genetics, Demography and Viability of Fragmented Populations. Cambridge: Cambridge University Press; 2000 pp 1–6. https://doi.org/10.1017/CBO9780511623448
Young A, Boyle T, Brown T (1996) The population genetic consequences of habitat fragmentation for plants. Trends Ecol Evol 11:413–418. https://doi.org/10.1016/0169-5347(96)10045-8
Zeng X, Fischer GA (2020) Wind pollination over 70 years reduces the negative genetic effects of severe forest fragmentation in the tropical oak Quercus bambusifolia. Heredity 124:156–169. https://doi.org/10.1038/s41437-019-0258-3
Zhang R, Hipp AL, Gailing O (2015) Sharing of chloroplast haplotypes among red oak species suggests interspecific gene flow between neighboring populations. Botany 93:691–700. https://doi.org/10.1139/cjb-2014-0261
Acknowledgements
We thank Alexandra Dolynska for her assistance in the laboratory and Jeremias Götz and Gerold Dinkel for their technical support collecting and planting the seeds. Gudrun Diederich for her help and support in the maintenance of the plants.
Funding
The funding for this research was provided by the Georg-August University of Göttingen. John Carlson has been funded as a Mercator fellow by the DFG grant GA 714/7 − 1 (reference number 429696097).
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
T.DF.: Laboratory procedures, data acquisition, data analysis, writing - original draft. K.B.: Data evaluation, writing - review and editing. O.G.: Conceptualization, funding acquisition, writing - review and editing. J.C.: Conceptualization, review, and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no known conflict of interest to disclose.
Additional information
Communicated by Judy Simon.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dominguez-Flores, T., Budde, K.B., Carlson, J.E. et al. Distance-dependent mating but considerable pollen immigration in an isolated Quercus rubra planting in Germany. Eur J Forest Res (2024). https://doi.org/10.1007/s10342-024-01704-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10342-024-01704-x