Abstract
The formation of inclusion bodies (IBs) without enzyme activity in bacterial research is generally undesirable. Researchers have attempted to recovery the enzyme activities of IBs, which are commonly known as active IBs. Tyrosine phenol-lyase (TPL) is an important enzyme that can convert pyruvate and phenol into 3,4-dihydroxyphenyl-l-alanine (L-DOPA) and IBs of TPL can commonly occur. To induce the correct folding and recover the enzyme activity of the IBs, peptides, such as ELK16, DKL6, L6KD, ELP10, ELP20, L6K2, EAK16, 18A, and GFIL16, were fused to the carboxyl terminus of TPL. The results showed that aggregate particles of TPL-DKL6, TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 improved the enzyme activity by 40.9%, 50.7%, 48.9%, 86.6%, and 97.9%, respectively. The peptides TPL-DKL6, TPL-EAK16, TPL-18A, and TPL-GFIL16 displayed significantly improved thermostability compared with TPL. L-DOPA titer of TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16, with cells reaching 37.8 g/L, 53.8 g/L, 37.5 g/L, and 29.1 g/L, had an improvement of 111%, 201%, 109%, and 63%, respectively. A higher activity and L-DOPA titer of the TPL-EAK16 could be valuable for its industrial application to biosynthesize L-DOPA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Tyrosine phenol-lyase (TPL) (EC4.1.99.2), a tetrameric enzyme, can catalyze stereospecific isotope exchange of α-protons of various amino acids with pyridoxal-5′-phosphate (PLP) as the cofactor [20]. TPL has been mainly isolated and characterized from bacteria, such as Citrobacter freundii [38,46], Erwinia herbicola [58], and Fusobacterium nucleatum [62]. Monovalent cations, K+ or NH4+ are necessary for achieving high activity levels of TPL [33]. The biosynthesis based on the enzyme activity of TPL has attracted attention for its application in the production of enantiomerically pure α-deuterated (S)-amino acids, such as S-alkyl/aryl-cysteines [7] and l-dihydroxyphenylalanine (L-DOPA) [54]. L-DOPA, as a precursor of dopamine, has been regarded as the main medicine to treat Parkinson’s disease since the 1960s [42].TPL can catalyze the reversible hydrolytic cleavage of L-tyrosine to phenol and ammonium pyruvate [8], and due to the reversibility of this reaction, when the catechol is substituted for phenol, L-DOPA is synthesized. The presence of ammonium salt plays an important role in leading the direction of the reversible catalysis in the production of L-DOPA [57].
Metabolic engineering strategies for de novo synthesis of L-DOPA have mainly focused on directing major carbon fluxes toward desired L-tyrosine formation [50]. L-tyrosine, as a precursor for synthesis of L-DOPA, is usually difficult to be synthesized and accumulated by metabolic pathways [10]. There are inevitable issues for de novo synthesis of L-DOPA, such as low titer and browning caused by oxidation and remaining tyrosine; therefore, research is important to further improve the catalytic performance of TPL and the enzymatic synthesis of L-DOPA. In the last decade, production of pyruvic acid, which is the precursor for enzymatic synthesis of L-DOPA, has been significantly improved [64] [29,30,31]. This has significantly lowered the price of pyruvic acid and makes the enzyme synthesis of L-DOPA the most cost-effective method. Higher enzyme activity and operational stability are research hotspots in the development of the industrial application of TPL [39]. Thermostability and enzyme activity of the TPL from Symbiobacterium toebii were simultaneously improved by the application of random mutagenesis and the subsequent reassembly of the acquired mutations [43]. It was found that Ser51 of TPL participated in stabilizing the ammonium form of Lys257, by studying the kinetics for the complexes of the mutant form with competitive inhibitors [1]. It was also found that Phe-448 and Phe-449 in TPL contributed to β-elimination catalysis reactions by introducing ground state destabilization in the substrate [39].
Heterogeneous proteins expressed in recombinant bacteria aggregate as insoluble protein clusters were defined as inclusion bodies (IBs) [26]. These IBs are formed due to unfolded or highly misfolded polypeptides that fail to reach their normal conformation [48]. Refolding of proteins from IBs is affected by several factors, including solubilization of IBs by denaturants, removal of the denaturant, and induction of refolding by small molecule additives [13,26]. Some protein aggregates, known as active IBs, become functionally active proteins due to a correct fold with the assistance of self-assembly peptides (SAPs) [53,61] and there are also studies showing that these SAPs could enhance the thermostability of the target enzyme [22]. SAPs, as specific type of peptide, can spontaneously assemble into nanostructures. Moreover, SAPs displayed several advantages, such as small size, convenient operations, and cost-saving. In last several years, these SAPs have been applied to recover the biological activity of bacterial IBs. Active IBs induced by terminally attaching SAPs have also been successfully applied in protein purification with high yields and purity [9,49,60].
In this study, TPL was overexpressed in Escherichia coli BL21(DE3) as a heterogeneous protein. Besides the soluble expression of TPL, insoluble TPL aggregates without enzyme activity also existed, which might be resulted from undesirable misfolded polypeptides. As the waste byproducts during protein expression, IBs are recognized as the major bottleneck in recombinant protein expression [19] and are discarded from further processing or are eventually used as a pure protein by in vitro refolding and recovery [37]. As whole-cell biocatalyst, L-DOPA titer could be seriously affected by TPL IBs without activity. Therefore, it is necessary to recover active proteins from the TPL IBs and improve the catalytic performance by fusing SAPs to the C-terminus of TPL. The short peptides ELK16, DKL6, L6KD, ELP10, ELP20, L6K2, EAK16, 18A, and GFIL16 were each attached to the C-terminus of TPL, respectively. Amino acid sequences of peptides were showed in Table S1. It was found that TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 showed obvious advantages, such as IBs with enzyme activity and improved thermostability and the L-DOPA titer of TPL-EAK16 increased to 53.8 g/L. As a result, fusion of peptides to the C-terminus of TPL has been identified as achieving improved enzyme performance. The enzyme activity of TPL IBs was first studied and activated using peptides, demonstrating the potential of peptides as novel IB-inducing fusion tags in vivo. In addition, peptides could also be successfully applied in the production and purification of proteins.
Materials and methods
Construction of the plasmids
Strains of E. coli BL21(DE3) and E. coli JM109, pET28a-TPL plasmid were preserved in the laboratory [14]. The nucleotide sequence of TPL gene has been recorded in National Center for Biotechnology Information (NCBI) and GenBank accession number was MN205565. Proteins expressed by plasmid pET28a-TPL in E. coli BL21 were designated as TPL. Plasmid pET28a-TPL was amplified using the polymerase chain reaction (PCR) with pET-TPL-F/R primers (Table S2). Genes of peptides were synthesized by GenScript (Nanjing, China). The synthesized genes were amplified by PCR with primers (Table S2) and assembled into pET28a-TPL with homologous recombination to yield vector pET28a-TPL-ELK16, pET28a-TPL-DKL6, pET28a-TPL-L6KD, pET28a-TPL-ELP10, pET28a-TPL-ELP20, pET28a-TPL-L6K2, pET28a-TPL-EAK16, pET28a-TPL-18A, and pET28a-TPL-GFIL16. The corresponding proteins expressed by these plasmids were designated as TPL-ELK16, TPL-DKL6, TPL-L6KD, TPL-ELP10, TPL-ELP20, TPL-L6K2, TPL-EAK16, TPL-18A, and TPL-GFIL16, respectively.
Expression of TPL
The growth of recombinant cells was carried out in terrific broth (TB) medium supplemented with 50 mg/L kanamycin with 400 rpm at 37 °C in 3 L bioreactor (T&J Bio-engineering Co., LTD, Shanghai, China). Isopropyl-β-d-thiogalactoside (IPTG) with a final concentration of 0.2 mM was added to the culture to induce the expression of proteins and the cells were cultured for a further 10 h at 20 °C.
Preparation and SDS-PAGE analysis of TPL
The collected cell pellets were washed three times and resuspended with buffer B1 [50 mM KH2PO4–K2HPO4, 1% (v/v) glycerol, (pH 8.5)]. Cell suspensions were lysed by ultrasonic disruption. The resulting cell lysates were centrifuged at 10,000 rpm for 10 min at 4 °C to separate the soluble fractions and insoluble fractions. The amounts of fusion proteins in both soluble and insoluble fractions were analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Protein concentration was normalized using an enhanced BCA protein assay kit (P0009; Beyotime Biotechnology, Jiangsu, China).
The resulting TPLs all containing a 6 × His fusion tag, were purified by an His-Trap FF 5-mL column with AKTA Pure system (GE Healthcare, Piscataway, NJ, USA). The purification operation was the same as a previous study [14].
Enzyme activity assay of TPL
The relative enzyme activity of supernatant and precipitates of TPLs was determined with the β-elimination reactions of L-tyrosine [2]. L-tyrosine was decomposed into phenol, pyruvate, and ammonia, and the resulting pyruvate concentration was measured by Agilent 1200–series high-performance liquid chromatography (HPLC) [28]. To determine the optimal pH value of TPLs at 20 °C, L-tyrosine was dissolved in 50 mM KH2PO4-K2HPO4 (pH 6 to 8) and 50 mM glycine–NaOH (pH 8–10) buffer, respectively. The enzyme activity of intracellular supernatants from TPL were defined as 100%. The enzyme activity of TPLs was determined by the same method at different temperatures, as one unit of TPL activity, defined as the amount of enzyme producing 1 µM pyruvate per min. The half-life of TPLs at 20 °C, 40 °C was determined as before [14]. A reference standard of 100% TPLs without maintenance at 20 °C, 40 °C, or 60 °C was established and measured at the corresponding reaction temperature.
The kinetic parameters of TPLs were determined at 20 °C, with 900 μL substrate and 100 μL enzyme liquid-containing 30 µM PLP. The substrate was L-DOPA with a gradient concentration from 0.1 to 6 mM. Lineweaver–Burk plots were generated for the analysis of Km, Vmax, and kcat/Km [3].
Whole-cell biosynthesis
The initial reaction mixture was composed of 18 g/L sodium pyruvate, 10 g/L catechol, 30 g/L ammonium salt, 4 g/L sodium sulfite, 2 g/L EDTA, and 30 µM PLP. The pH value of reaction mixture was adjusted to 8.5 with ammonium hydroxide. Sodium pyruvate was added at 12 g/L/h from 0 to 5 h. Catechol was added at 10 g/L/h during the first 2 h, at 8 g/L/h from 2 to 4 h, and at 4 g/L/h from 4 to 5 h. The reaction was at 15 °C. Ammonium salt can trigger the reaction to the direction of synthesizing L-DOPA due to the reversibility of reaction [34]. It was necessary to avoid light during the reaction to protect the production L-DOPA from being decomposed. Sample was extracted every 1 h and treated with the 50% (v/v) 2 M HCL.
HPLC analysis
L-DOPA was analyzed by an Agilent 1200-series HPLC system equipped with a reverse-phase Gemini NX-C18 column (4.6 × 250 mm) according to a previous protocol [15].
Transmission electron microscopic analysis
Morphometric analysis of the TPLs was performed using transmission electron microscopy (TEM). Solutions of 2.5% glutaraldehyde and 2% osmium tetroxide were used successively to fix the cells. After a graded ethanol serial dehydration step, the cells were embedded in epoxy resins. The fixed cells were sectioned, stained by uranyl acetate solution and lead citrate, and were observed using a Hitachi H-7650B (Hitachi, Japan) TEM at an accelerating voltage of 80 kV.
Results
Formation of active TPL IBs in E. coli
The TPL without a short peptide displayed two expression styles as supernatant and aggregate (52 kDa) (Fig. 1). SDS-PAGE results showed that TPL-ELK16, TPL-DKL6, TPL-L6K2, TPL-18A, and TPL-GFIL16 were mainly expressed in the insoluble fractions. The growth of E. coli BL21 containing fusion proteins was slow compared to the E. coli BL21 containing TPL (Fig. 2). The final OD600 of E. coli BL21 expressing TPL-ELK16, TPL-L6K2, TPL-18A, and TPL-GFIL16 was 18.7, 19.6, 18.4, and 19.2, which was 4.8, 3.9, 5.1, and 4.4 less, respectively, for the E. coli BL21 expressing TPL. To a certain extent, TPLs that were fused with peptide and expressed in the form of aggregate particles had a slight inhibitory effect on the E. coli BL21 cell growth.
SDS-PAGE analysis of fusion protein expression. Intracellular supernatant of TPL (lane 1); aggregate of TPL (lane 2), TPL-ELK16 (lane 4), TPL-DKL6 (lane 6), TPL-L6KD (lane 8), TPL-ELP10 (lane 10), TPL-ELP20 (lane 12), TPL-L6K2 (lane 14), TPL-EAK16 (lane 16) and TPL-18A (lane 18), TPL-GFIL16 (lane 20); supernatant of TPL-ELK16 (lane 3), TPL-DKL6 (lane 5), TPL-L6KD (lane 7), TPL-ELP10 (lane 9), TPL-ELP20 (lane 11), TPL-L6K2 (lane 13), TPL-EAK16 (lane 15), TPL-18A (lane 17), and TPL-GFIL16 (lane 19). Levels were normalized according to a 4-mg standard. Targets are denoted with a black arrow. M, standards (kDa)
The final OD600 of E. coli BL21 expressing TPLs. E. coli BL21(DE3) cells expressing TPL, TPL-ELK16, TPL-DKL6, TPL-L6KD, TPL-ELP10, TPL-ELP20, TPL-L6K2, TPL-EAK16, TPL-18A, and TPL-GFIL16 were cultivated at 37 °C for 2 h. The temperature was adjusted to 20 °C for 10 h. Experiments were performed in triplicate, and error bars represent the standard deviation
Catalytic properties of TPLs
The optimal pH value for the catalysis of TPL-ELK16, TPL-DKL6, TPL-L6KD, TPL-ELP10, TPL-ELP20, TPL-EAK16, and TPL-18A was 8.5. For the catalysis of TPL-L6K2 and TPL-GFIL16, the optimal pH value was 9.0 (Fig. 3). Anyway, alkaline buffer is a good fit for TPLs. Aggregate particles of TPL displayed only 7.3% enzyme activity in comparison with TPL supernatant (Fig. 4a); compared with aggregate particles of TPL-DKL6, TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16, this displayed 40.9%, 50.7%, 48.9%, 86.6%, and 97.9% enzyme activity, respectively. TPL displayed maximal enzyme activity of 5.1 IU/mL at 10 °C, (Fig. 4b), but this decreased by 55.8% at 20 °C. The obvious decrease of enzyme activity has been verified in other study [14], which could be contributed to the lower thermostability. TPL does not belong to enzyme with good thermostability. Therefore, reaction catalyzed by TPL was generally performed at 15 °C [62]. The maximal enzyme activity of TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 was 5.0 IU/mL, 5.4 IU/mL, 4.7 IU/mL, and 3.9 IU/mL, respectively, exhibiting improvement by 117.4%, 133.0%, 104.3%, and 69.5% relative to TPL at 20 °C. Using L-DOPA as the substrate, kinetic parameters showed that the Vmax and Km values of TPL were 0.04 mM/min/mg and 2.19 mM (Table S3). The Vmax of TPL-DKL6, TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 was higher by 24.4%, 24.4%, 12.2%, 9.8%, and 17%, respectively, relative to the TPL. The Km value of TPL-DKL6, TPL-ELP10, TPL-EAK16, and TPL-GFIL16 increased by 5.5%, 22.3%, 7.8%, and 1.8%, respectively, relative to the TPL. The kcat/Km value of TPL-DKL6, TPL-ELP10, TPL-EAK16, and TPL-GFIL16 increased by 25.4%, 24.0%, 11.1%, and 17.7%, respectively, relative to the TPL.
Enzyme activity of TPL and TPL fused with peptides. Relative activity of supernatants and precipitates from TPL and TPL fused with peptides at 20 °C (a). Enzyme activity of TPL and TPL fused with peptides at 10 °C, 20 °C, 30 °C, 40 °C, and 50 °C (b). Experiments were performed in triplicate, and error bars represent the standard deviation
Thermostability of TPLs at different temperatures
The tests showed that TPL-DKL6, TPL-EAK16, TPL-18A, and TPL-GFIL16 displayed thermostability with a half-life of 14.8 min, 14.2 min, 14.1 min, and 15.8 min at 20 °C and 12.0 min, 13.9 min, 11.6 min, and 13.9 min at 40 °C, respectively. The residual activity for TPL remained at only 29.6% following incubation at 40 °C for 10 min (Fig. 5). The half-life of TPL was 13.8 min at 20 °C and 6.3 min at 40 °C. All the results demonstrated that the fusion of functional short peptides increased the thermostability of TPL significantly, especially TPL-18A and TPL-GFIL16.
L-DOPA titer by whole-cell biosynthesis
Whole-cell biosynthesis was carried out for 6 h at 20 °C and the L-DOPA titer was tested every hour (Fig. 6). As the reaction progressed, crystals of L-DOPA were observed. When the reaction time reached 5 h at 20 °C, L-DOPA biosynthesized by TPL cells was 17.9 g/L. In contrast, L-DOPA biosynthesized by TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 cells reached maximal values of 37.8 g/L, 53.8 g/L, 37.5 g/L, and 29.1 g/L, respectively, showing improvement by 111%, 201%, 109%, 63%, respectively, relative to TPL. The improved L-DOPA titer was correlated with improved enzyme activity. Moreover, the expression of soluble TPL-ELP10 and TPL-EAK16 has been enhanced, which could be observed on the SDS-PAGE (Fig. 1). IBs of TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 also displayed catalytic activity (Fig. 4a). Both the increased expression of soluble TPL and active inclusion bodies probably resulted in the increased L-DOPA titer.
Effect of peptides on L-DOPA production by whole-cell biosynthesis. Whole-cell L-DOPA production of TPL, TPL-ELK16, TPL-DKL6, TPL-L6KD, TPL-ELP10, TPL-ELP20, TPL-L6K2, TPL-EAK16, TPL-18A, and TPL-GFIL16 at 20 °C. Experiments were performed in duplicate, and error bars represent the standard deviation
Morphology analysis of TPLs
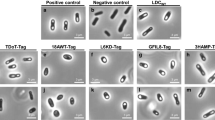
Negative stained TEM images showed homo-oligomeric structures for the freshly prepared TPL, TPL-ELK16, TPL-DKL6, TPL-L6KD, TPL-ELP10, TPL-ELP20, TPL-L6K2, TPL-EAK16, TPL-18A, and TPL-GFIL16 from E. coli BL21 (Fig. 7). The surfactant-like peptides DKL6, L6KD, and L6K2 can act as a pull-down handler for converting soluble proteins into active aggregates and ELP covalently attaches with the C-terminus residue of the xylanase via a random coil [[22]]. The self-assembly of EAK16-family peptides can induce fibrillary or globular assemblies [9]. ELK16, DKL6, L6KD, with ELP10 induce the formation of spherical TPL aggregates and L6K2, EAK16 induce irregular small TPL aggregates. It has been shown that 18A can self-assemble by coiled formation in the aqueous solution, and form discoidal particles, or helix fibrils [21]. Accordingly, elliptic aggregates of TPL-18A were formed. GFIL16 is comprised of hydrophobic residues and has strong hydrophobicity [49]. Therefore, results showed that GFIL16 induced the formation of large and amorphous TPL aggregates that were different from the others.
Discussion
There are a few routine methods reported to improve the catalytic efficiency of TPL-mediated biosynthesis. It is common to genetically engineer the existing enzymes by rational protein design or random/saturated mutagenesis [41], which are labor-intensive and time-consuming. To improve thermostability and reusability, immobilization technology is a useful tool [35]. Conventional immobilization usually leads to the conformational changes or to prevention of the active pocket from being bound with substrates, and this could result in a decrease in enzyme activity [24]. TPL immobilized with wet nanoporous silica gels resulted in modified steady-state distribution of catalytic intermediates, but specific activities decreased [44]. However, SAPs were covalently attached with the C-terminus residue of the TPL via an irregular coil, overcoming the conflict between stability and enzyme activity. For example, 18A fused nitrilase aggregates (Nit-SEA) displayed enzyme activity and increased thermostability [55], which was the prospective results in this study. SAPs were able to serve as “pull-down” fusion tags to effectively induce the formation of cytoplasmic active IBs in E. coli, such as 18A, ELK16, DKL6, L6KD, and L6K2 [11]. EAK16 and GFIL16, which were composed of hydrophobic amino acids, played functional roles in the formation of active IBs due to strong hydrophobicity [1]. Accordingly, aggregate particles of TPL-DKL6, TPL-ELP10, TPL-EAK16, TPL-18A, and TPL-GFIL16 displayed 40.9%, 50.7%, 48.9%, 86.6%, and 97.9% enzyme activity relative to TPL supernatant, respectively. Moreover, TPL-DKL6, TPL-EAK16, TPL-18A, and TPL-GFIL16 displayed thermostability with prolonged half-life.
Previously investigated SAPs include ionic beta-strand peptide ELK16 [51], elastin-like polypeptide (ELP) [47], surfactant-like peptide (DKL6, L6KD, and L6K2) [63], charged and hydrophobic peptide EAK16 [9], modified apolipoprotein A-I mimetic amphipathic peptide 18A [25], and hydrophobic self-assembling peptide GFIL16 [49]. The SAPs that induced the formation of active IBs were regarded as the carrier for immobilization. IBs with high specific enzyme activity have been verified as potentially useful biocatalysts [36]. Target proteins were successfully released from active IBs upon cleavage of the intein between the peptide tag and the target protein [27]. It has become a promising method to develop a quick technique to enable protein expression and purification in bacteria. Pure enzyme and immobilized enzymes have been mainly used as biocatalyst in current industrial production. Compared to the traditional purification and immobilization, this process could achieve both time-saving and cost-saving in industrial application [52]. It has also been proven that one-step purification is feasible by inserting an appropriate self-cleavable site between the target protein and the peptide [17].
E. coli has served as a cell factory for recombinant protein expression for a long time and the cells expressing an enzyme can be used as biocatalysts [4,5], but overexpression of heterogeneous proteins can lead to the formation and accumulation of inactive IBs due to misfolded polypeptides [12,59]. Currently effective strategies have been developed to recover enzyme activity from aggregate particles [23,56]. Active IBs have shown unique advantages, such as easy separation, greater stability, and reusability, and offer robustness in applications [32]. Fusion of the N-terminal peptide GFIL8 to the Ulp1 results in increased production of active IBs with higher resistance to limited proteolysis and less leakage at different storage temperatures [18]. In this study, peptide-induced aggregates did not interfere with the correct folding of the target proteins, and active TPL IBs also contributed to the improved enzyme activity and thermostability.
In summary, more attention should be paid to recovering the enzyme activity of IBs with the assistance of SAPs [6,61]. To improve the biosynthesis of L-DOPA, TPL-EAK16 with obvious advantages was achieved with the help of EAK16 SAPs. The L-DOPA titer and productivity of TPL-EAK16 were 53.8 g/L and 10.76 g/L/h, respectively. Compared with the previous reported aggregation prone fusion partners [40,45], the peptides used in this study were much shorter in size and more easily modulated due to their simpler secondary structure [16]. However, it is difficult to control the amount and size of aggregate particles in the host cell. Spherical TPL aggregates were induced by ELK16, DKL6, L6KD, and ELP10. Aggregates of TPL-L6K2, TPL-EAK16 displayed small irregular shape. GFIL16 with strong hydrophobicity induced the formation of large and amorphous TPL aggregates. All the results demonstrated that TPL aggregate particles with enzyme activity induced by the SAPs had significance for application to other enzymes. Moreover, IBs, as biocatalysts, were easily separated and purified [11]. This method is, therefore, much easier, more economic, and time-efficient to scale up from an economic point of view.
References
Barbolina MV, Kulikova VV, Tsvetikova MA, Anufrieva NV, Revtovich SV, Phillips RS, Gollnick PD, Demidkina TV, Faleev NG (2018) Serine 51 residue of Citrobacter freundii tyrosine phenol-lyase assists in C-α-proton abstraction and transfer in the reaction with substrate. Biochimie 147:63–69
Chandel M, Azmi W (2013) Purification and characterization of tyrosine phenol lyase from Citrobacter freundii. Appl Biochem Biotech 171:2040–2052
Crawford JJ, Hollett JW, Craig DB (2016) Determination of the inhibitor dissociation constant of an individual unmodified enzyme molecule in free solution. Electrophoresis 37:2217–2225
Cui Y, Meng Y, Zhang J, Cheng B, Yin H, Gao C, Xu P, Yang C (2017) Efficient secretory expression of recombinant proteins in Escherichia coli with a novel actinomycete signal peptide. Protein Expr Purif 129:69–74
Dai L, Tai C, Shen Y, Guo Y, Tao F (2019) Biosynthesis of 1,4-butanediol from erythritol using whole-cell catalysis. Biocatal Biotransform 37:92–96
Dasgupta A, Das D (2019) Designer peptide amphiphiles: self-assembly to applications. Langmuir 35:10704–10724
Demidkina TV, Barbolina MV, Faleev NG, Sundararaju B, Gollnick PD, Phillips RS (2002) Threonine-124 and phenylalanine-448 in Citrobacter freundii tyrosine phenol-lyase are necessary for activity with L-tyrosine. Biochem J 363:745–752
Do Q, Nguyen GT, Phillips RS (2016) Inhibition of tyrosine phenol-lyase by tyrosine homologues. Amino Acids 48:2243–2251
Emamyari S, Kargar F, Sheikh-hasani V, Emadi S, Fazli H (2015) Mechanisms of the self-assembly of EAK16-family peptides into fibrillar and globular structures: molecular dynamics simulations from nano- to micro-seconds. Eur Biophys J Biophy 44:263–276
Fordjour E, Adipah FK, Zhou S, Du G, Zhou J (2019) Metabolic engineering of Escherichia coli BL21 (DE3) for de novo production of L-DOPA from D-glucose. Microb Cell Fact 18:74
Garcia-Fruitos E, Vazquez E, Diez-Gil C, Luis Corchero J, Seras-Franzoso J, Ratera I, Veciana J, Villaverde A (2012) Bacterial inclusion bodies: making gold from waste. Trends Biotechnol 30:65–70
Gardner QA, Hassan N, Hafeez S, Arif M, Akhtar M (2019) Exploring the nature of inclusion bodies by MALDI mass spectrometry using recombinant proinsulin as a model protein. Int J Biol Macromol 139:647–653
Hamidi SR, Safdari Y, Arabi MS (2019) Test bacterial inclusion body for activity prior to start denaturing and refolding processes to obtain active eukaryotic proteins. Protein Expr Purif 154:147–151
Han H, Zeng W, Du G, Chen J, Zhou J (2020) Site-directed mutagenesis to improve the thermostability of tyrosine phenol-lyase. J Biotechnol 310:6–12
Ho PY, Chiou MS, Chao AC (2003) Production of L-DOPA by tyrosinase immobilized on modified polystyrene. Appl Biochem Biotechnol 111:139–152
Hong H, Fan H, Chalamaiah M, Wu J (2019) Preparation of low-molecular-weight, collagen hydrolysates (peptides): Current progress, challenges, and future perspectives. Food Chem 301:125222
Hwang PM, Pan JS, Sykes BD (2014) Targeted expression, purification, and cleavage of fusion proteins from inclusion bodies in Escherichia coli. FEBS Lett 588:247–252
Jiang L, Xiao W, Zhou X, Wang W, Fan J (2019) Comparative study of the insoluble and soluble Ulp1 protease constructs as carrier free and dependent protein immobilizates. J Biosci Bioeng 127:23–29
Koszagova R, Krajcovic T, Palencarova-Talafova K, Patoprsty V, Vikartovska A, Pospiskova K, Safarik I, Nahalka J (2018) Magnetization of active inclusion bodies: comparison with centrifugation in repetitive biotransformations. Microb Cell Fact 17:1–8
Koulikova VV, Zakomirdina LN, Gogoleva OI, Tsvetikova MA, Morozova EA, Komissarov VV, Tkachev YV, Timofeev VP, Demidkina TV, Faleev NG (2011) Stereospecificity of isotopic exchange of C-alpha-protons of glycine catalyzed by three PLP-dependent lyases: the unusual case of tyrosine phenol-lyase. Amino Acids 41:1247–1256
Lazar KL, Miller-Auer H, Getz GS, Orgel J, Meredith SC (2005) Helix-turn-helix peptides that form alpha-helical fibrils: turn sequences drive fibril structure. Biochemistry 44:12681–12689
Li C, Zhang G (2014) The fusions of elastin-like polypeptides and xylanase self-assembled into insoluble active xylanase particles. J Biotechnol 177:60–66
Li G-Y, Xiao Z-Z, Lu H-P, Li Y-Y, Zhou X-H, Tan X, Zhang X-Y, Xia X-L, Sun H-C (2016) A simple method for recombinant protein purification using self-assembling peptide-tagged tobacco etch virus protease. Protein Expr Purif 128:86–92. https://doi.org/10.1016/j.pep.2016.08.013
Lin P-C, Weinrich D, Waldmann H (2010) Protein biochips: oriented surface immobilization of proteins. Macromol Chem Phys 211:136–144
Lin Z, Zhou B, Wu W, Xing L, Zhao Q (2013) Self-assembling amphipathic alpha-helical peptides induce the formation of active protein aggregates in vivo. Faraday Discuss 166:243–256
Linova MY, Risor MW, Jorgensen SE, Mansour Z, Kaya J, Sigurdarson JJ, Enghild JJ, Karring H (2020) A novel approach for production of an active N-terminally truncated Ulp1 (SUMO protease 1) catalytic domain from Escherichia coli inclusion bodies. Protein Expr Purif 166:105507
Luan C, Xie YG, Pu YT, Zhang HW, Han FF, Feng J, Wang YZ (2014) Recombinant expression of antimicrobial peptides using a novel self-cleaving aggregation tag in Escherichia coli. Can J Microbiol 60:113–120
Luo Z, Liu S, Du G, Xu S, Zhou J, Chen J (2018) Enhanced pyruvate production in Candida glabrata by carrier engineering. Biotechnol Bioeng 115:473–482
Luo Z, Liu S, Du G, Zhou J, Chen J (2017) Identification of a polysaccharide produced by the pyruvate overproducer Candida glabrata CCTCC M202019. Appl Microbiol Biot 101:4447–4458. https://doi.org/10.1007/s00253-017-8245-1
Luo Z, Zeng W, Du G, Chen J, Zhou J (2019) Enhanced pyruvate production in Candida glabrata by engineering ATP futile cycle system. Acs Synth Biol 8:787–795. https://doi.org/10.1021/acssynbio.8b00479
Luo Z, Zeng W, Du G, Chen J, Zhou J (2020) Enhancement of pyruvic acid production in Candida glabrata by engineering hypoxia-inducible factor 1. Bioresour Technol. https://doi.org/10.1016/j.biortech.2019.122248
Mee C, Banki MR, Wood DW (2008) Towards the elimination of chromatography in protein purification: expressing proteins engineered to purify themselves. Chem Eng J 135:56–62
Milic D, Demidkina TV, Faleev NG, Matkovic-Calogovic D, Antson AA (2008) Insights into the catalytic mechanism of tyrosine phenol-lyase from X-ray structures of quinonoid intermediates. J Biol Chem 283:29206–29214
Min K, Park K, Park DH, Yoo YJ (2015) Overview on the biotechnological production of L-DOPA. Appl Microbiol Biotechnol 99:575–584
Mohamad NR, Marzuki NHC, Buang NA, Huyop F, Wahab RA (2015) An overview of technologies for immobilization of enzymes and surface analysis techniques for immobilized enzymes. Biotechnol Biotechnol Equip 29:205–220
Nahalka J, Nidetzky B (2007) Fusion to a pull-down domain: a novel approach of producing Trigonopsis variabilis D-amino acid oxidase as insoluble enzyme aggregates. Biotechnol Bioeng 97:454–461
Pekarsky A, Konopek V, Spadiut O (2019) The impact of technical failures during cultivation of an inclusion body process. Bioprocess Biosyst Eng 42:1611–1624
Phillips RS, Demidkina TV, Faleev NG (2003) Structure and mechanism of tryptophan indole-lyase and tyrosine phenol-lyase. BBA Proteins Proteom 1647:167–172
Phillips RS, Vita A, Spivey JB, Rudloff AP, Driscoll MD, Hay S (2016) Ground-state destabilization by Phe-448 and Phe-449 contributes to tyrosine phenol-lyase catalysis. ACS Catal 6:6770–6779
Qi M, Li L, Lu Y, Chen H, Zhang M, Wang M, Ge L, Yang J, Shi N, Chen T, Tang X (2019) Proteome profiling to identify peroxiredoxin 1 interacting protein partners in nicotine-associated oral leukoplakia. Arch Oral Biol 108:104537–104537
Qin N, Shen Y, Yang X, Su L, Tang R, Li W, Wang M (2017) Site-directed mutagenesis under the direction of in silico protein docking modeling reveals the active site residues of 3-ketosteroid-Δ1-dehydrogenase from Mycobacterium neoaurum. World J Microbiol Biotechnol 33:146
Rekdal VM, Bess EN, Bisanz JE, Turnbaugh PJ, Balskus EP (2019) Discovery and inhibition of an interspecies gut bacterial pathway for Levodopa metabolism. Science 364:1055. https://doi.org/10.1126/science.aau6323
Rha E, Kim S, Choi S-L, Hong S-P, Sung M-H, Song JJ, Lee S-G (2009) Simultaneous improvement of catalytic activity and thermal stability of tyrosine phenol-lyase by directed evolution. FEBS J 276:6187–6194
Roessl U, Nahalka J, Nidetzky B (2010) Carrier-free immobilized enzymes for biocatalysis. Biotechnol Lett 32:341–350
Sandoval JE, Huang Y-H, Muise A, Goodell MA, Reich NO (2019) Mutations in the DNMT3A DNA methyltransferase in acute myeloid leukemia patients cause both loss and gain of function and differential regulation by protein partners. J Biol Chem 294:4898–4910
Sundararaju B, Antson AA, Phillips RS, Demidkina TV, Barbolina MV, Gollnick P, Dodson GG, Wilson KS (1997) The crystal structure of Citrobacter freundii tyrosine phenol-lyase complexed with 3-(4'-hydroxyphenyl)propionic acid, together with site-directed mutagenesis and kinetic analysis, demonstrates that arginine 381 is required for substrate specificity. Biochemistry 36:6502–6510
Trabbic-Carlson K, Liu L, Kim B, Chilkoti A (2004) Expression and purification of recombinant proteins from Escherichia coli: Comparison of an elastin-like polypeptide fusion with an oligohistidine fusion. Protein Sci 13:3274–3284
Vallejo LF, Rinas U (2004) Strategies for the recovery of active proteins through refolding of bacterial inclusion body proteins. Microb Cell Fact 3:11
Wang X, Zhou B, Hu W, Zhao Q, Lin Z (2015) Formation of active inclusion bodies induced by hydrophobic self-assembling peptide GFIL8. Microb Cell Fact 14:88
Wei T, Cheng B-Y, Liu J-Z (2016) Genome engineering Escherichia coli for L-DOPA overproduction from glucose. Sci Rep 6:1–9
Wu W, Xing L, Zhou B, Lin Z (2011) Active protein aggregates induced by terminally attached self-assembling peptide ELK16 in Escherichia coli. Microb Cell Fact 10:9–16
Wingfield PT, Palmer I, Liang S-M (2014) Folding and purification of insoluble (inclusion body) proteins from Escherichia coli. Curr Protoc Protein Sci 78:6.5.1–6.5.27
Xi G, Esfandiary R, Sacramento CB, Jouihan H, Sharma A, Roth R, Linke T (2019) Refolding and purification of cGMP-grade recombinant human neurturin from Escherichia coli inclusion bodies. Protein Expr Purif 168:105552–105552
Xu B, Lei Q, Zeng W, Wei Y, Huang K, Zhou J (2019) High-density fermentation for preparing tyrosine phenol lyase and application in L-DOPA synthesis (高密度发酵产酪氨酸酚裂解酶及催化合成l-dopa). Food Ferment Ind 45:7–14
Yang X, Huang A, Peng J, Wang J, Wang X, Lin Z, Li S (2014) Self-assembly amphipathic peptides induce active enzyme aggregation that dramatically increases the operational stability of nitrilase. Rsc Adv 4:60675–60684
Ye X, Yu D, Wu Y, Han J, Li S, Wu Q, Li D, Qi J (2019) An efficient large-scale refolding technique for recovering biologically active recombinant human FGF-21 from inclusion bodies. Int J Biol Macromol 135:362–372
Yuan W, Zhong S, Xiao Y, Wang Z, Sun J (2019) Efficient biocatalyst of L-DOPA with Escherichia coli expressing a tyrosine phenol-lyase mutant from Kluyvera intermedia. Appl Biochem Biotech. https://doi.org/10.1007/s12010-019-03164-1
Zeng W, Xu B, Du G, Chen J, Zhou J (2019) Integrating enzyme evolution and high-throughput screening for efficient biosynthesis of l-DOPA. J Ind Microbiol Biot 46:1631–1641
Zhao F, Song Q, Wang B, Han Y, Zhou Z (2019) Purification and immobilization of alpha-amylase in one step by gram-positive enhancer matrix (GEM) particles from the soluble protein and the inclusion body. Appl Microbiol Biotechnol 104:643–652
Zhao Q, Zhou B, Gao X, Xing L, Wang X, Lin Z (2017) A cleavable self-assembling tag strategy for preparing proteins and peptides with an authentic N-terminus. Biotech J 12:1600656
Zhao Y, Yang W, Chen C, Wang J, Zhang L, Xu H (2018) Rational design and self-assembly of short amphiphilic peptides and applications. Curr Opin Colloid Interface Sci 35:112–123
Zheng R-C, Tang X-L, Suo H, Feng L-L, Liu X, Yang J, Zheng Y-G (2018) Biochemical characterization of a novel tyrosine phenol-lyase from Fusobacterium nucleatum for highly efficient biosynthesis of L-DOPA. Enzyme Microb Tech 112:88–93
Zhou B, Xing L, Wu W, Zhang X-E, Lin Z (2012) Small surfactant-like peptides can drive soluble proteins into active aggregates. Microb Cell Fact 11:10
Zhou J, Liu L, Chen J (2011) Improved ATP supply enhances acid tolerance of Candida glabrata during pyruvic acid production. J Appl Microbiol 110:44–53. https://doi.org/10.1111/j.1365-2672.2010.04865.x
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2019YFA09004800), the National Natural Science Foundation of China (31670095, 31770097), the National Science Fund for Excellent Young Scholars (21822806), the Fundamental Research Funds for the Central Universities (JUSRP51701A), and the National First-class Discipline Program of Light Industry Technology and Engineering (LITE2018-08).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Han, H., Zeng, W., Zhang, G. et al. Active tyrosine phenol-lyase aggregates induced by terminally attached functional peptides in Escherichia coli. J Ind Microbiol Biotechnol 47, 563–571 (2020). https://doi.org/10.1007/s10295-020-02294-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10295-020-02294-4