Abstract
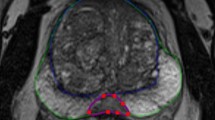
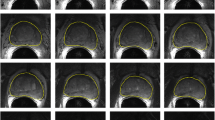
Three dimensional (3D) manual segmentation of the prostate on magnetic resonance imaging (MRI) is a laborious and time-consuming task that is subject to inter-observer variability. In this study, we developed a fully automatic segmentation algorithm for T2-weighted endorectal prostate MRI and evaluated its accuracy within different regions of interest using a set of complementary error metrics. Our dataset contained 42 T2-weighted endorectal MRI from prostate cancer patients. The prostate was manually segmented by one observer on all of the images and by two other observers on a subset of 10 images. The algorithm first coarsely localizes the prostate in the image using a template matching technique. Then, it defines the prostate surface using learned shape and appearance information from a set of training images. To evaluate the algorithm, we assessed the error metric values in the context of measured inter-observer variability and compared performance to that of our previously published semi-automatic approach. The automatic algorithm needed an average execution time of ∼60 s to segment the prostate in 3D. When compared to a single-observer reference standard, the automatic algorithm has an average mean absolute distance of 2.8 mm, Dice similarity coefficient of 82%, recall of 82%, precision of 84%, and volume difference of 0.5 cm3 in the mid-gland. Concordant with other studies, accuracy was highest in the mid-gland and lower in the apex and base. Loss of accuracy with respect to the semi-automatic algorithm was less than the measured inter-observer variability in manual segmentation for the same task.





Similar content being viewed by others
References
Siegel, R.L., K.D. Miller, and A. Jemal, Cancer statistics, 2015. CA Cancer J Clin, 2015. 65(1): p. 5–29.
Canadian Cancer Society’s Advisory Committee on Cancer Statistics, Canadian Cancer Statistics 2015, in Canadian Cancer Society. 2015
Kurhanewicz, J., D. Vigneron, P. Carroll, and F. Coakley, Multiparametric magnetic resonance imaging in prostate cancer: present and future. Curr Opin Urol, 2008. 18(1): p. 71–7.
Shukla-Dave, A. and H. Hricak, Role of MRI in prostate cancer detection. NMR Biomed, 2014. 27(1): p. 16–24.
Akin, O., E. Sala, C.S. Moskowitz, K. Kuroiwa, N.M. Ishill, D. Pucar, P.T. Scardino, and H. Hricak, Transition zone prostate cancers: features, detection, localization, and staging at endorectal MR imaging. Radiology, 2006. 239(3): p. 784–92.
Gilderdale, D.J., N.M. de Souza, G.A. Coutts, M.K. Chui, D.J. Larkman, A.D. Williams, and I.R. Young, Design and use of internal receiver coils for magnetic resonance imaging. Br J Radiol, 1999. 72(864): p. 1141–51.
Anwar, M., A.C. Westphalen, A.J. Jung, S.M. Noworolski, J.P. Simko, J. Kurhanewicz, M. Roach, 3rd, P.R. Carroll, and F.V. Coakley, Role of endorectal MR imaging and MR spectroscopic imaging in defining treatable intraprostatic tumor foci in prostate cancer: quantitative analysis of imaging contour compared to whole-mount histopathology. Radiother Oncol, 2014. 110(2): p. 303–8.
Gibson, E., G.S. Bauman, C. Romagnoli, D.W. Cool, M. Bastian-Jordan, Z. Kassam, M. Gaed, M. Moussa, J.A. Gomez, S.E. Pautler, J.L. Chin, C. Crukley, M.A. Haider, A. Fenster, and A.D. Ward, Toward prostate cancer contouring guidelines on magnetic resonance imaging: dominant lesion gross and clinical target volume coverage via accurate histology fusion. Int J Radiat Oncol Biol Phys, 2016. 96(1): p. 188–96.
Kim, Y., I.C. Hsu, J. Pouliot, S.M. Noworolski, D.B. Vigneron, and J. Kurhanewicz, Expandable and rigid endorectal coils for prostate MRI: impact on prostate distortion and rigid image registration. Med Phys, 2005. 32(12): p. 3569–78.
Husband, J.E., A.R. Padhani, A.D. MacVicar, and P. Revell, Magnetic resonance imaging of prostate cancer: comparison of image quality using endorectal and pelvic phased array coils. Clin Radiol, 1998. 53(9): p. 673–81.
Smith, W.L., C. Lewis, G. Bauman, G. Rodrigues, D. D'Souza, R. Ash, D. Ho, V. Venkatesan, D. Downey, and A. Fenster, Prostate volume contouring: a 3D analysis of segmentation using 3DTRUS, CT, and MR. Int J Radiat Oncol Biol Phys, 2007. 67(4): p. 1238–47.
Martin, S., V. Daanen, and J. Troccaz, Atlas-based prostate segmentation using an hybrid registration. Int J CARS, 2008. 3(6): p. 8.
Vikal, S., S. Haker, C. Tempany, and G. Fichtinger, Prostate contouring in MRI guided biopsy. Proc SPIE, 2009. 7259: p. 72594A.
Dice, L.R., Measures of the amount of ecologic association between species. Ecology, 1945. 26(3): p. 297–302.
Toth, R. and A. Madabhushi, Multifeature landmark-free active appearance models: application to prostate MRI segmentation. IEEE Trans Med Imaging, 2012. 31(8): p. 1638–50.
Liao, S., Y. Gao, Y. Shi, A. Yousuf, I. Karademir, A. Oto, and D. Shen, Automatic prostate MR image segmentation with sparse label propagation and domain-specific manifold regularization. 2013, Springer. p. 511–523.
Cheng, R., B. Turkbey, W. Gandler, H.K. Agarwal, V.P. Shah, A. Bokinsky, E. McCreedy, S. Wang, S. Sankineni, M. Bernardo, T. Pohida, P. Choyke, and M.J. McAuliffe, Atlas based AAM and SVM model for fully automatic MRI prostate segmentation. Conf Proc IEEE Eng Med Biol Soc, 2014. 2014: p. 2881–5.
Cheng, R., H.R. Roth, L. Lu, S. Wang, B. Turkbey, W. Gandler, E.S. McCreedy, H.K. Agarwal, P. Choyke, and R.M. Summers, Active appearance model and deep learning for more accurate prostate segmentation on MRI, in SPIE Medical Imaging. 2016, International Society for Optics and Photonics. p. 97842I-97842I-9.
Guo, Y., Y. Gao, and D. Shen, Deformable MR prostate segmentation via deep feature learning and sparse patch matching. IEEE Trans Med Imaging, 2016. 35(4): p. 1077–89.
Qiu, W., J. Yuan, E. Ukwatta, Y. Sun, M. Rajchl, and A. Fenster, Prostate segmentation: an efficient convex optimization approach with axial symmetry using 3-D TRUS and MR images. IEEE Trans Med Imaging, 2014. 33(4): p. 947–60.
Mahapatra, D. and J.M. Buhmann, Prostate MRI segmentation using learned semantic knowledge and graph cuts. IEEE Trans Biomed Eng, 2014. 61(3): p. 756–64.
Makni, N., P. Puech, R. Lopes, A.S. Dewalle, O. Colot, and N. Betrouni, Combining a deformable model and a probabilistic framework for an automatic 3D segmentation of prostate on MRI. Int J Comput Assist Radiol Surg, 2009. 4(2): p. 181–8.
Litjens, G., R. Toth, W. van de Ven, C. Hoeks, S. Kerkstra, B. van Ginneken, G. Vincent, G. Guillard, N. Birbeck, J. Zhang, R. Strand, F. Malmberg, Y. Ou, C. Davatzikos, M. Kirschner, F. Jung, J. Yuan, W. Qiu, Q. Gao, P.E. Edwards, B. Maan, F. van der Heijden, S. Ghose, J. Mitra, J. Dowling, D. Barratt, H. Huisman, and A. Madabhushi, Evaluation of prostate segmentation algorithms for MRI: the PROMISE12 challenge. Med Image Anal, 2014. 18(2): p. 359–73.
Shahedi, M., D.W. Cool, C. Romagnoli, G.S. Bauman, M. Bastian-Jordan, E. Gibson, G. Rodrigues, B. Ahmad, M. Lock, A. Fenster, and A.D. Ward, Spatially varying accuracy and reproducibility of prostate segmentation in magnetic resonance images using manual and semiautomated methods. Med Phys, 2014. 41(11): p. 113503.
Chen, X. and U. Bagci, 3D automatic anatomy segmentation based on iterative graph-cut-ASM. Med Phys, 2011. 38(8): p. 4610–22.
Atkinson, A.C. and T.-C. Cheng, Computing least trimmed squares regression with the forward search. Statistics and Computing, 1999. 9(4): p. 251–263.
Woolson, R.F. and W.R. Clarke, Statistical methods for the analysis of biomedical data. Vol. 371. 2011 Wiley
Warfield, S.K., K.H. Zou, and W.M. Wells, Simultaneous truth and performance level estimation (STAPLE): an algorithm for the validation of image segmentation. IEEE Trans Med Imaging, 2004. 23(7): p. 903–21.
Acknowledgements
The authors gratefully acknowledge the late Dr. Cesare Romagnoli for his support and scientific contribution to this work.
This work was supported by the Ontario Institute for Cancer Research and the Ontario Research Fund. This work was also supported by Prostate Cancer Canada and is proudly funded by the Movember Foundation—Grant # RS2015-04. A. Fenster holds a Canada Research Chair in Biomedical Engineering and acknowledges the support of the Canada Research Chair Program. A. D. Ward holds a Cancer Care Ontario Research Chair in Cancer Imaging.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The study was approved by the research ethics board of our institution, and written informed consent was obtained from all patients prior to enrolment.
Rights and permissions
About this article
Cite this article
Shahedi, M., Cool, D.W., Bauman, G.S. et al. Accuracy Validation of an Automated Method for Prostate Segmentation in Magnetic Resonance Imaging. J Digit Imaging 30, 782–795 (2017). https://doi.org/10.1007/s10278-017-9964-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10278-017-9964-7