Abstract
There is mounting evidence that reproductively isolated, but morphologically weakly differentiated species (so-called cryptic species) represent a substantial part of biological diversity, especially in bryophytes. We assessed the evolutionary history and ecological differentiation of a species pair, Dicranum brevifolium and D. septentrionale, which have overlapping ranges in the Holarctic. Despite their morphological similarity, we found similar genetic differentiation as between morphologically well-differentiated Dicranum species. Moreover, we detected gene tree discordance between plastid and nuclear markers, but neither of the two datasets resolved the two as sister species. The signal in trnL–trnF better reflects the morphological and ecological affinities and indicates a close relationship while ITS sequence data resolved the two taxa as phylogenetically distantly related. The discordance is probably unrelated to the ecological differentiation of D. septentrionale to colonise subneutral to alkaline substrates (vs. acidic in D. brevifolium), because this ability is rare in the genus and shared with D. acutifolium. This taxon is the closest relative of D. septentrionale according to the trnL–trnF data and does not share the discordance in ITS. We furthermore demonstrate that beside D. acutifolium, both D. septentrionale and D. brevifolium occur in the Alps but D. brevifolium is most likely rarer. Based on morphological analyses including factor analysis for mixed data of 45 traits we suggest treating the latter two as near-cryptic species and we recommend verifying morphological determinations molecularly.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The application of molecular methods has opened a completely new perception of biological diversity on Earth. In bryophytes, the second largest group of land plants after vascular plants, it has led to the discovery of substantial species diversity in many lineages that had not been recognised in earlier morphological concepts (e.g., Bakalin et al. 2020; Fernandez et al. 2006; Hedenäs and Eldenäs 2007; see Renner 2020 for a review). Such taxa, where molecular data indicate reproductive isolation without substantial morphological differentiation, are usually referred to as cryptic. Most commonly, cryptic taxa are phylogenetically closely related and often reflect differentiation along geographical or ecological gradients. The lack of morphological differentiation can then be explained by a recent origin form a common ancestor without substantial phenotypic change (Struck et al. 2018). By contrast, examples of more deeply divergent cryptic species are rare and can result from morphological stasis or convergence (Renner et al. 2013; Struck et al. 2018).
Here we assess an example of two deeply divergent species in the Dicranum acutifolium complex: Dicranum brevifolium (Lindb.) Lindb. and D. septentrionale Tubanova & Ignatova. The latter species was first reported through variation in sequences of the nuclear ITS region analysing accessions of D. brevifolium from Russia (Tubanova et al. 2010). Based on the molecular clustering of specimens, minor morphological differences were found between the two species; i.e., predominantly elongated distal and median lamina cells in D. septentrionale (vs. isodiametric in D. brevifolium) and usually longer basal cells. Subsequently, Lang et al. (2014) also considering specimens from Scandinavia, performed a cluster analysis of 34 morphological characters that failed to unambiguously discern the two species while the phylogenetic analysis of the concatenated dataset of the nuclear ITS region and five plastid loci indicated that the two taxa are not closely related (Lang et al. 2015). Dicranum septentrionale was resolved as falling outside of the clade containing the morphologically similar species D. brevifolium and D. acutifolium (Lindb. & Arnell) C.E.O.Jensen. These latter two species appeared to be genetically more closely related to species which differ strongly morphologically, e.g., D. angustum Lindb. and D. bonjeanii De Not.
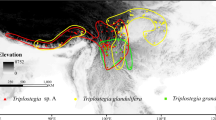
To date, no clear geographical pattern has been documented that could explain the genetic differentiation of D. brevifolium and D. septentrionale, both of which are known from across Eurasia (Lang et al. 2014; Tubanova et al. 2010; Fig. 1). Therefore, the genetic differentiation of the two species may coincide with ecological differences. Both are known to inhabit mountainous regions, and less commonly, lowland areas (Lang et al. 2014; Tubanova et al. 2010). However, they may occur on divergent substrates, but to date, this hypothesis has not been investigated. Different substrate affinities have often been observed between closely related plant species, including bryophytes (Kiebacher et al. 2022).
a Distribution of Dicranum brevifolium (yellow dots) and D. septentrionale (blue) in Eurasia according to Tubanova et al. (2010) and Lang et al. (2014) considering only molecularly analysed specimens. b Sampling of D. brevifolium s.l. (black dots) and D. acutifolium (asterisks) in the Alps. c Plant of Dicranum brevifolium s.l. in dry state, scale bare = 2 mm
In Central Europe, taxonomic assessment of the D. acutifolium complex is incomplete and ambiguous. The complex comprises the morphologically similar species D. acutifolium, D. bardunovii Tubanova & Ignatova., D. brevifolium and D. septentrionale (Lang et al. 2014). Dicranum brevifolium (sub D. muehlenbeckii subsp. brevifolium (Lindb.) J.J. Amann or D. muehlenbeckii var. brevifolium Lindb.) was mentioned from Switzerland by Limpricht (1886) and Amann et al. (1918) and by Breidler (1891) from Austria, but all authors claimed morphological differences to Nordic collections. Certainly, these records include D. acutifolium and D. septentrionale. Until recently, reliable records for D. acutifolium from the Alps were very sparse and its occurrence was not accepted in Grims (1999) for Austria and in Meinunger and Schröder (2007) for Germany. Only recently Hedenäs and Bisang (2004) or Schlüsslmayr (2019) seriously reported its presence in the Eastern Alps and so far, only one Central European sample of the complex from Switzerland (belonging to D. brevifolium) and one from Austria (belonging to D. septentrionale) have been molecularly examined (Lang et al. 2014).
By analysing multiple accessions of D. acutifolium and D. brevifolium s.l. (D. brevifolium + D. septentrionale) from different substrates in the Alps molecularly and morphologically and including available sequence data from previous work in the phylogenetic analyses we address the discordance between the phylogenetic and morphological signal in D. brevifolium and D. septentrionale.
Specifically, (i) we test the consistency of the phylogenetic signal in nuclear and plastid sequence data and (ii) if specimens from the Alps can be assigned to one of the two species based on morphological characteristics. Furthermore, (iii) we explore whether the two taxa differ regarding substrate specificity and (iv) we provide information about their occurrence in the Alps.
Material and methods
Taxon sampling and morphological analyses
To provide specimens < 20 years old (suitable for molecular examination) we sent requests to GJO, W and Z + ZT and contacted seven private collectors active in the Alps (see acknowledgements) who we expected to hold collections of the complex. In this way we succeeded in providing 15 specimens from the Austrian (4 specimens) and Swiss (11) Alps which according to an initial morphological examination were assigned to D. brevifolium s.l. Furthermore, we considered two specimens of D. acutifolium (Fig. 1). To examine the specimens of D. brevifolium s.l. morphologically we applied two approaches. (a) We asked an experienced bryologist to determine the specimens using the distinguishing characters presented by Tubanova et al. (2010) and Lang et al. (2014) prioritising the shape of the median and upper lamina cells and the length of the basal cells as pointed out by Tubanova et al. (2010). In case of overlapping characteristics or discordance between the characters, the determinator had the option to assign multiple taxon names, starting with the best fit taxon name. (b) We compiled a data set of 45 morphological traits comprising the characters presented by Tubanova et al. (2010) and Lang et al. (2014) and further characters commonly used for species distinction in the genus (see e.g., Hedenäs and Bisang 2004) and analysed it using multivariate statistics. (i) We measured length and width of 5 leaves of one well-grown shoot of each specimen and estimated distal dorsal costa and upper leaf margin ornamentation and the shape of the base of these leaves. (ii) We measured length and width of 10 basal cells of each of the 5 leaves and counted the number of pores per cells. And (iii), we visually estimated the portion of elongate, isodiametric, oblate and triangular lamina cells at ½–\({\raise0.5ex\hbox{$\scriptstyle 1$} \kern-0.1em/\kern-0.15em \lower0.25ex\hbox{$\scriptstyle 3$}}\) above the leaf base. We calculated minimum, maximum and average values, as well as leaf and basal cell length/width ratio per specimen to compile the spreadsheet of traits and analysed it using factor analysis for mixed data (FAMD; Pagès 2004) with the FAMD function in the R package FactoMineR (Le et al. 2008). The technique combines principal component and multiple correspondence analyses and allows to simultaneously analyse quantitative and qualitative variables. The spreadsheet including a more detailed description how the traits were estimated is available in Supplementary Information 1. We ran the analysis in R version 4.1.3 (R Core Team 2022) and summarized results graphically using the plotly package (Sievert 2020). Furthermore, we tested for species specific differences individually for each trait using the Wilcoxon-signed-rank test implemented in the ggpubr package (Kassambara 2023).
Marker selection
To identify plastid markers that are sufficiently variable to resolve the species of the D. acutifolium complex, we used the dataset of Lang et al. (2015) and generated phylogenetic trees independently for each of the five loci examined (rpoB, rps19–rpl2, psbA–trnH, trnL–trnF, trnT–rps4). For simplicity, we reduced the set of accessions to all representatives of D. brevifolium, D. acutifolium and D. septentrionale and at least two (if available) representatives of the other species and Holomitrium arboreum Mitten as an outgroup species. We aligned the sequences using the AlignSeqs function of the DECIPHER package (Wright 2015) with default settings in R version 4.1.3 (R Core Team 2022) and subsequent manual corrections. We scored indel data using the simple coding method (Simmons and Ochoterena 2000) and ran Bayesian Inference (BI) analyses using MrBayes v.3.2.7a (Ronquist et al. 2012). We specified a GTR + G + I model, a sample frequency of 100, stop rule set to yes with critical value for the topological convergence diagnostic set to 0.01 and default settings for all other parameters. Only trnL–trnF differentiated D. brevifolium, D. acutifolium and D. septentrionale from each other and from other species (Figs. S1−5 in Supplementary Information 2), thus we proceeded to sequence this locus and the nuclear ITS region. We a priori selected the latter because it had more parsimony informative sites than the five plastid markers combined (Lang et al. 2015).
DNA extraction and sequencing
The protocols to generate the sequence data followed previous work (Kiebacher et al. 2021, 2022). In brief, genomic DNA was extracted using the NaOH method (Werner et al. 2002), we targeted the nuclear ITS region including partial sequence of 18S rRNA gene, ITS1, 5.8S rRNA gene, ITS2 and partial sequence of 26S rRNA gene using the primers m-18-S (Spagnuolo et al. 1999) and m-25-R (Stech and Frahm 1999), and the plastid spacer trnL–trnF including partial sequences of the trnL- and trnF-genes using the primers TabC and TabF (Taberlet et al. 1991). Reagents and volumes for the PCR reaction were the same as is described in Kiebacher et al. (2021), except that we used a 2.5 µg/ml solution of bovine serum albumin instead of water. Cycling for ITS followed Kiebacher et al. (2021) and for the trnL–trnF locus it started with 3-min initialisation at 94 °C, followed by 40 cycles of 1 min at 94 °C, 1 min at 49 °C, and 1 min at 72 °C, and a final extension step of 5 min at 72 °C. Purification of the PCR product, sequencing and editing of raw sequences were again performed following Kiebacher et al. (2021). Amplification of both loci failed for three of the specimens of D. brevifolium s.l. which we excluded from the subsequent analyses. For two specimens of D. brevifolium s.l. and one of D. acutifolium, only ITS sequences could be generated.
Molecular data set and analyses
For the phylogenetic analyses we considered the set of accessions used for marker selection and extended it with accessions retrieved from Genbank. Specifically, we included accessions of D. bardunovii, D. howellii Renauld & Cardot, D. ignatovii Tubanova & Fedosov, D. japonicum Mitten, D. laevidens R.S.Williams and D. muehlenbeckii Bruch & Schimp. which were identified as similar to ITS or trnL–trnF sequences of our accessions of D. brevifolium s.l. based on BLAST searches using the megablast algorithm (https://blast.ncbi.nlm.nih.gov/Blast.cgi) with default settings. From this raw set of accessions, we excluded sequences with low coverage (less than 70% of the target locus) which applied to one trnL–trnF (GenBank accession number DQ462591) and one ITS sequence of D. septentrionale (HQ830340). The final dataset comprised 31 species of Dicranum, each represented by at least two accessions, and the outgroup taxon Chorisodontium aciphyllum (Hook.f. & Wilson) Broth. Dicranum brevifolium s.l. was represented by the 14 accessions from the Alps (two of which were from previous studies) and five accessions of D. brevifolium and nine of D. septentrionale from other regions. We aligned trnL–trnF sequences using the same function and software as used for marker selection and ITS sequences using the online interface of MAFFT v7 (Katoh and Standley 2013) applying the E-INS-i strategy, default settings for all other parameters and subsequent manual corrections. We scored indels for both alignments using simple coding (Simmons and Ochoterena 2000) and performed BI and Maximum Likelihood (ML) analyses. Bayesian Inference was performed with the same software and settings as for marker selection and convergence of the runs was checked using Tracer v1.7.2 (Rambaut et al. 2018). For ML analyses we used RAxML v8.2.4 (Stamatakis 2014), also specified the GTR + G + I model and stopped bootstrap analysis automatically using the autoMRE command. Support for the nodes of the best scoring tree out of 50 independent ML runs was assessed using the thorough bootstrapping algorithm with the extended majority rule bootstopping criterion. We summarised the support of nodes from the two approaches using TreeGraph 2 (Stöver and Müller 2010) displaying the BI topology.
Substrate reaction
To classify the substrate of specimens of D. brevifolium s.l., including specimens for which sequence data were generated in previous studies, we defined two classes of substrate reaction, acidic (A) and subneutral to alkaline (N). To assign specimens to either of the two we used the bedrock type at the collection site and anticipated that substantial content of carbonate results in subneutral to alkaline conditions and that low or no content of carbonate results in acidic conditions. Bedrock type was derived from label information, geological maps, field observations of the collectors, and the evaluation of soil reaction affinities of associated species. For Switzerland, we used the geological map GeoCover of the Swiss Federal Office of Topography Swisstopo (available online; www.swisstopo.admin.ch) which combines information from different sources listed in Swisstopo (2021). For Austria, we used Rockenschaub and Nowotny (2009) and for Sweden we used the online bedrock map provided by SGU Geological Survey of Sweden (https://apps.sgu.se/kartvisare/kartvisare-berg-50-250-tusen.html). We succeeded in classifying 23 of the 29 specimens of D. brevifolium s.l. considered. For the remaining specimens, information on bedrock type was not available from labels and neither the locality nor the geological maps were accurate enough to reliably classify them. For other Dicranum species, we classified the substrate reaction requirements according to our expertise and indications in the literature (Chien et al. 1999; Dierssen 2001; Hill et al. 2007; Ireland 2007; Urmi 2010; van Zuijlen et al. 2023), which required an additional intermediate class for species that occur on moderately acidic to subneutral substrates.
Results
Molecular and morphological species assignment
The trnL–trnF alignment comprised 471 positions, of which 48 were variable and 47 were parsimony-informative. The alignment of the nrITS region comprised 883 positions, 122 were variable and 91 were parsimony-informative.
In agreement with the morphological diagnosis the phylogenetic analysis of ITS data resolved the two accessions of D. acutifolium among accessions of this species examined in earlier studies (Fig. 2). Of the 12 Alpine accessions of D. brevifolium s.l. for which we could generate sequences, four were resolved among accessions of D. brevifolium and eight among accessions of D. septentrionale (Table 1, Fig. 2). This molecular species assignment was consistent with the assignment in trnL–trnF except for one accession of D. acutifolium and two of D. brevifolium for which trnL–trnF sequences could not be generated (Table 1, Fig. 3). In the morphological determination, four accessions of D. brevifolium s.l. were assigned to D. brevifolium, four to D. septentrionale and for the four others, the morphological affinities were ambiguous (Table 1). The unambiguous determinations matched the molecular assignment in no more than two cases for D. septentrionale (Table 1). The four specimens with ambiguous morphological affinities were molecularly either resolved among accessions of D. brevifolium (2 specimens) or D. septentrionale (2 specimens).
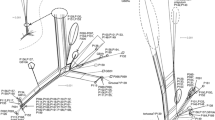
Bayesian inference from plastid trnL–trnF sequence data. Numbers above branches are Bayesian posterior probabilities ≥ 0.5, numbers below branches are bootstrap support values ≥ 50 of branches obtained from maximum likelihood analysis of the same dataset. New accessions are in bold, accessions retrieved from GenBank are followed by the accession number. For voucher information see Table 1. Colours indicate substrate reaction requirements of species and for Dicranum brevifolium and D. septentrionale the substrate reaction at the collection sites of each specimen: brown, acidic; ochre, moderately acidic to subneutral; green, subneutral to alkaline; blank fields, unknown
Bayesian inference from nuclear ITS sequence data. Numbers above branches are Bayesian posterior probabilities ≥ 0.5, numbers below branches are bootstrap support values ≥ 50 of branches obtained from maximum likelihood analysis of the same dataset. New accessions are in bold, accessions retrieved from GenBank are followed by the accession number. For voucher information see Table 1, for colour coding see Fig. 2
Gene tree topology
The analysis of trnL–trnF sequence data resolved all species of the D. acutifolium complex in a supported clade (BI posteriori probability 0.99/ML bootstrap support 67; Fig. 2). They shared two synapomorphic single nucleotide polymorphisms (SNP’s) with respect to all other species considered. Within the clade, D. brevifolium and D. bardunovii lack synapomorphic mutations while D. acutifolium + D. septentrionale share two synapomorphic indels and D. septentrionale differs from D. acutifolium in two SNP’s. The phylogenetic tree of the nuclear ITS region revealed a different topology, it resolved D. septentrionale as rather distantly related to the other species of the complex (Fig. 3). Accessions of D. septentrionale were grouped in a supported clade (1/93) of a basal polytomy and differed from D. acutifolium, D. brevifolium and D. bardunovii in five indels and five SNP’s. The latter species appeared more closely related to eight other, morphologically well-differentiated Dicranum-species (e.g., D. scoparium Hedw., D. leioneuron Kindb) and were placed with these species in a supported clade (1/73). Within this clade they were grouped with weak support (0.6/-) together with D. angustum.
FAMD and trait analyses
The dimensions one and two of the FAMD analysis explained 37.3% and 14.5% of the variance respectively, revealed high within species variance and resolved the two species in two weakly separated clusters (Fig. 4a, Fig. S6 in Supplementary Information 2; a 3D plot of the first three dimensions is available in Supplementary Information 3). The analyses of individual traits failed to detect differences between the two species in the portion of elongate and isodiametric cells in the upper part of the lamina and revealed differences in leaf length, basal cell length, basal cell length/with ratio and the number of pores in basal cells, being generally larger in D. septentrionale (Fig. 4b, 5, Fig. S7 in Supplementary Information 2).
Factor analysis for mixed data a of 45 morphological traits of Dicranum brevifolium (yellow dots) and D. septentrionale (blue dots) and b differences in mean percentage of elongate (ECmean) and isodiametric cells (ICmean) in the upper lamina, maximum basal cell length (BCLmax), mean basal cell length/width ratio (BCLWRmean), maximum leaf length (LLmax) and minimum number of pores per basal cell (BCPmin). For voucher information of specimens see Table 1. Boxplots for all 45 traits are available in Fig. S7 in Supplementary Information 2
Substrate reaction
All seven (out of 10) accessions of D. brevifolium for which substrate reaction could be classified were collected from acidic substrates, and all 16 (out of 19) accessions of D. septentrionale were collected from subneutral to alkaline substrates (Table 1).
Discussion
Evolutionary history
We found substantial discordance between plastid and nuclear sequence data regarding the phylogenetic placement of D. septentrionale. While the signal in trnL–trnF suggests the monophyly of the D. acutifolium complex, ITS sequences suggest that D. septentrionale is more deeply divergent, i.e., the most recent common ancestor of the species of the complex is shared with many other morphologically substantially different taxa. We hypothesize that the trnL–trnF tree better represents the evolutionary history of the species of the complex, because it coincides with their morphological similarity. Furthermore, it explains the ecological similarity of D. septentrionale and D. acutifolium to colonise subneutral to alkaline substrates by a shared evolutionary history instead of assuming an additional transition to this rare trait in the genus (Figs. 2, 3). The discordance in ITS may be due to well-known evolutionary processes, specifically gene flow and incomplete lineage sorting. To disentangle these two processes in our example would require an extended molecular dataset, but, the rare occurrence of sporophytes in the species of the D. acutifolium complex, as well as their longevity suggest that incomplete lineage sorting is a more likely explanation (Copetti et al. 2017; Meleshko et al. 2018). For instance, a recent genome-wide study of long-lived and rather rarely sexually reproducing peatmosses detected incomplete lineage sorting as the main source of discordance between nuclear and organellar genomes (Meleshko et al. 2021). On the other hand, gene flow seems to predominate in short-lived species that frequently produce sporophytes (Košnar et al. 2012; Linde et al. 2020; McDaniel et al. 2010; Nieto-Lugilde et al. 2018).
Within the D. acutifolium complex, the trnL–trnF data suggest a sister position of D. acutifolium and D. septentrionale and not of D. brevifolium and D. septentrionale, although the latter two are morphologically very similar whereas D. acutifolium can usually be easily distinguished from them. Consequently, the similarity of D. brevifolium and D. septentrionale would represent an example of either stasis or convergence (Struck et al. 2018). As outlined above, the sister position of D. septentrionale and D. acutifolium is supported by their substrate specificity. Both occur on subneutral to alkaline substrates whereas D. brevifolium and D. bardunovii occur on acidic substrates. Since the preference for acidic substrates is a conservative trait in the genus and transitions to alkaline substrates are rare (Figs. 2, 3), the ability of D. septentrionale and D. acutifolium to colonise alkaline substrates probably originated from the same evolutionary event and is most likely unrelated to the discordance in ITS sequence data of D. septentrionale, because D. acutifolium does not share the discordance.
Interestingly, the genetic distances between morphologically poorly differentiated species such as D. brevifolium and D. septentrionale are similar or even larger as between morphologically well-differentiated species (e.g., D. elongatum Schleich. ex Schwägr. and D. fragilifolium Lindb. in trnL–trnF; D. leioneuron and D. scoparium in ITS). Overall, most other Dicranum species considered are morphologically well-differentiated. All but four (D. bardunovii, D. dispersum, D. ignatovii, D. septentrionale) were described more than 100 years ago and accepted in most taxonomic concepts since their description (e.g., Hill et al. 2006; Hodgetts et al. 2020; Ireland 1971; Limpricht 1886; Mönkemeyer 1927; Nyholm 1954; Suzuki 2016), hence, the genetic distance does not necessarily reflect the morphological distance. Possibly, in the absence of morphological differentiation, a morphological optimum for a particular habitat has already been reached and is selectively maintained, whereas adaptive radiation is still possible at the physiological level. Moreover, it is likely that (near-) cryptic taxa can also be detected in other Dicranum species if a larger number of specimens are studied along their ecological gradients.
Substrate specificity and cryptic species
Dicranum septentrionale occurs on subneutral to alkaline substrates, whereas its near-cryptic counterpart D. brevifolium occurs on acidic substrates. The two species thus represent ecological vicariants. Cryptic or near-cryptic species associated with ecological vicariance have been observed in various groups of plants, animals and fungi (e.g., Boissin et al. 2011; Douglas et al. 2011; Douhan et al. 2008; Reis et al. 2020) but were rarely reported from bryophytes (Kiebacher et al. 2022). Only recently, Kiebacher et al. (2022) described a similar example in the epilithic moss Lewinskya killiasii (Müll. Hal.) Kiebacher, Köckinger & Jan Kučera (Orthotrichaceae). The two subspecies of this taxon occur on siliceous and carbonate rocks, respectively. Divergent substrate preference is probably a common phenomenon among (near-)cryptic bryophytes but this has not been exhaustively demonstrated, possibly because many examples are still unknown and hidden in species considered insensitive to substrate reaction. In contrast, cryptic speciation has often been shown to correlate with biogeographical patterns (e.g., Bakalin et al. 2020; Hedenäs 2017; Hutsemékers et al. 2012; Shaw 2001). Due to the long evolutionary history of many bryophyte species (Laenen et al. 2014), presumably both processes in concert often contributed to the cryptic diversity observed today (Hedenäs 2017; Hedenäs et al. 2020). In the case of D. brevifolium and D. septentrionale, however, a clear biogeographical pattern could not be observed. Both taxa occur throughout Russia (Tubanova et al. 2010), Scandinavia (Lang et al. 2014) and the Alps and the small overlap of the distribution areas observed in Russia (Tubanova et al. 2010) can now be explained by the different geology of these regions. The assessment of the ecological differentiation of D. brevifolium and D. septentrionale was largely limited to the Alps and Scandinavia (Table 1), because it was difficult to recover reliable substrate information of individual specimens from Russia. However, Tubanova et al. (2010) noted that in the Kola Peninsula D. brevifolium is much more common than D. septentrionale, whereas the latter is more common in the Kamchatka Peninsula, and this coincides with the geology of the two regions. The Kola Peninsula is dominated by acidic bedrock types (Mitrofanov et al. 1995) and the Kamchatka peninsula by alkaline basalts rich in magnesia (Portnyagin et al. 2015). Thus, this provides partial support for our observations in the Alps and Scandinavia.
Occurrence and morphological differentiation in the Alps
Our data confirm the presence of three species of the Dicranum acutifolium complex, D. acutifolium, D. brevifolium and D. septentrionale, in the Swiss and Austrian Alps. Dicranum septentrionale is probably the most frequent species of the complex in the region, whereas D. brevifolium hitherto is confirmed only from Central Switzerland. Our search for recent collections of D. brevifolium s.l. revealed only a few specimens from siliceous bedrock, suggesting that D. brevifolium is rare and that it should be prioritised in conservation. The same may apply to Scandinavia and Europe in general because D. brevifolium s.l. is normally given as basiphilous (Hedenäs and Bisang 2004; Nyholm 1987). Dicranum acutifolium seemingly occurs scattered in the limestone ranges of both Austria and Switzerland (Köckinger et al. 2021, Köckinger in litt). Additional research is required to examine the distribution of these species at a more detailed geographical resolution.
Dicranum acutifolium is mostly recognizable by its narrow and hardly-crisped leaves in dry condition and the elongated, variably shaped cells in the median and upper part of the lamina (Hedenäs and Bisang 2004; Tubanova et al. 2010). In contrast, D. brevifolium and D. septentrionale are characterised by strongly-crisped leaves in dry condition (Fig. 1c) and the median and upper cells rather uniformly shaped (oblate, quadrate or short rectangular). On the other hand, our data indicate that the morphological distinction of D. brevifolium and D. septentrionale in the Alps is not straightforward. This is especially true for the proportion of elongate and isodiametric upper lamina cells, which was proposed as a critical difference (Tubanova et al. 2010) based on specimens from Russia. This trait was similar in Alpine specimens of the two species (Figs. 4b, 5) and is possibly strongly influenced by environmental conditions. For example, the predominantly elongate upper lamina cells in the D. brevifolium collection from Avers (Lüth 8238; Table 1; CH3 in Fig. 5; Supplementary Information 1) could be due to the unusually humid microhabitat, partly shaded by vascular plants. In part, the analyses of individual traits also confirmed differences observed in specimens from Russia. Dicranum brevifolium, on average, has shorter basal cells, a smaller length/with ratio of basal cells and fewer pores per basal cell, and our data additionally suggest that its leaves are shorter (Tubanova et al. 2010; Fig. 4, Fig. S7 in Supplementary Information 2; Tubanova et al. 2010). However, due to overlaps in essentially all traits and the low number of specimens at hand for the analyses, these differences should be used with caution to distinguish the species and we recommend verifying morphological determinations molecularly, especially if the substrate is in conflict with the morphological diagnosis (e.g., D. brevifolium on calcareous bedrock). In such cases, it is more reliable to label specimens as D. brevifolium s.l.
The occurrence of (near-)cryptic species that are ecologically and genetically differentiated indicates that morphological concepts should not uncritically be interpreted to reflect the species diversity of bryophytes. Because (near-)cryptic species can have similar genetic differentiation as morphologically well differentiated ones, and may occupy different niches, they should not be ignored in conservation perspectives. To uncover hidden diversity in bryophytes it seems worthwhile to focus on species with presumably low substrate specificity.
Data availability
Sequence data are available in GenBank (https://www.ncbi.nlm.nih.gov/genbank), the alignments are available at the Zenodo repository (https://zenodo.org; https://doi.org/10.5281/zenodo.7768294) and voucher specimens are deposited in Z + ZT.
References
Amann J, Meylan C, Culmann P (1918) Flore des mousses de la Suisse. Deuxième partie Bryogéographie de la Suisse. Herbier Boissier, Genève
Bakalin VA, Vilnet AA, Choi SS, Nguyen VS (2020) Blepharostoma trichophyllum s.l. (Marchantiophyta): the complex of sibling species and hybrids. Plants 9:1–28. https://doi.org/10.3390/plants9111423
Boissin E, Stöhr S, Chenuil A (2011) Did vicariance and adaptation drive cryptic speciation and evolution of brooding in Ophioderma longicauda (Echinodermata: Ophiuroidea), a common atlanto-mediterranean ophiuroid? Mol Ecol 20:4737–4755. https://doi.org/10.1111/j.1365-294X.2011.05309.x
Breidler J (1891) Die Laubmoose Steiermarks. Mitteilungen Des Naturwissenschaftlichen Vereins Für Steiermark 28:3–234
Chien G, Vitt DH, He S (1999) 17. Dicranum Hedw., Sp. Musc. Frond. 126. 1801. Moss flora of China (English version), vol 1. Missouri Botanical Garden Press, St. Louis, pp 163–192
Copetti D, Búrquez A, Bustamante E et al (2017) Extensive gene tree discordance and hemiplasy shaped the genomes of North American columnar cacti. Proc Natl Acad Sci USA 114:12003–12008. https://doi.org/10.1073/pnas.1706367114
Dierssen K (2001) Distribution, ecological amplitude and phytosociological-characterization of European bryophytes. Bryophyt Bibl 56:1–289
Douglas NA, Wall WA, Xiang QY et al (2011) Recent vicariance and the origin of the rare, edaphically specialized Sandhills lily, Lilium pyrophilum (Liliaceae): Evidence from phylogenetic and coalescent analyses. Mol Ecol 20:2901–2915. https://doi.org/10.1111/j.1365-294X.2011.05151.x
Douhan GW, Smith ME, Huyrn KL et al (2008) Multigene analysis suggests ecological speciation in the fungal pathogen Claviceps purpurea. Mol Ecol 17:2276–2286. https://doi.org/10.1111/j.1365-294X.2008.03753.x
Fernandez CC, Shevock JR, Glazer AN, Thompson JN (2006) Cryptic species within the cosmopolitan desiccation-tolerant moss Grimmia laevigata. Proc Natl Acad Sci USA 103:637–642. https://doi.org/10.1073/pnas.0510267103
Grims F (1999) Die Laubmoose Österreichs. Catalogus Florae Austriae II. Teil, Bryophyten (Moose), Heft 1, Musci (Laubmoose). Österreichische Akademie der Wissenschaften, Vienna
Hedenäs L (2017) Scandinavian Oncophorus (Bryopsida, Oncophoraceae): species, cryptic species, and intraspecific variation. Eur J Taxon 2017:1–34. https://doi.org/10.5852/ejt.2017.315
Hedenäs L, Bisang I (2004) Key to European Dicranum species. Herzogia 17:179–197
Hedenäs L, Eldenäs P (2007) Cryptic speciation, habitat differentiation, and geography in Hamatocaulis vernicosus (Calliergonaceae, Bryophyta). Plant Syst Evol 268:131–145. https://doi.org/10.1007/s00606-007-0529-y
Hedenäs L, Kuznetsova OI, Ignatov MS (2020) A revision of the genus Tomentypnum (Amblystegiaceae) in northern Eurasia. Bryologist 123:377–395. https://doi.org/10.1639/0007-2745-123.3.377
Hill MO, Bell N, Bruggeman-Nannenga M et al (2006) An annotated checklist of the mosses of Europe and Macaronesia. J Bryol 28:198–267. https://doi.org/10.1179/174328206X119998
Hill M, Preston C, Bosanquet S, Roy D (2007) Bryoatt-attributes of British and Irish mosses, liverworts and hornworts. Centre for Ecology and Hydrology, Cambridgeshire
Hodgetts NG, Söderström L, Blockeel TL et al (2020) An annotated checklist of bryophytes of Europe, Macaronesia and Cyprus. J Bryol 42:1–116. https://doi.org/10.1080/03736687.2019.1694329
Hutsemékers V, Vieira CC, Ros RM et al (2012) Morphology informed by phylogeny reveals unexpected patterns of species differentiation in the aquatic moss Rhynchostegium riparioides s.l. Mol Phylogenet Evol 62:748–755. https://doi.org/10.1016/j.ympev.2011.11.014
Ireland RR (1971) Dicranum. Moss Flora of the Pacific Northwest. Hattori Botanical Laboratory, Nichinan, pp 72–81
Ireland RR (2007) Dicranum Hedwig, sp. Musc. Frond. 126. 1801. Flora of North America, vol 27. Bryophytes: Mosses. Oxford University Press, New York, pp 397–419
Kassambara A (2023) ggpubr: 'ggplot2' Based Publication Ready Plots. R package version 0.6.0. https://rpkgs.datanovia.com/ggpubr/
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kiebacher T, Köckinger H, Blom HH (2021) Schistidium foraminis-martini sp. nov. (Grimmiaceae), a high mountain calcicole from the European Alps molecularly related to S. agassizii. Bryophyt Divers Evol 44:1–11. https://doi.org/10.11646/bde.44.1.1
Kiebacher T, Köckinger H, Kučera J (2022) Adaptive divergence in the neglected alpine moss Lewinskya killiasii (Orthotrichaceae), an example of vicariance on different rock types in bryophytes. Bot J Linnean Soc 198:456–481. https://doi.org/10.1093/botlinnean/boab060
Köckinger H, Lüth M, Kiebacher T (2021) Dicranum acutifolium (Lindb. & Arnell) C.E.O.Jensen. In: Swissbryophytes Working Group (Hrsg.), Moosflora der Schweiz. https://doi.org/10.5167/uzh-204111.www.swissbryophytes.ch
Košnar J, Herbstová M, Kolář F et al (2012) A case study of intragenomic ITS variation in bryophytes: assessment of gene flow and role of polyploidy in the origin of European taxa of the Tortula muralis (Musci: Pottiaceae) complex. Taxon 61:709–720. https://doi.org/10.1002/tax.614001
Laenen B, Shaw B, Schneider H et al (2014) Extant diversity of bryophytes emerged from successive post-Mesozoic diversification bursts. Nat Commun 5:1–6. https://doi.org/10.1038/ncomms6134
Lang AS, Tubanova DY, Stech M (2014) Species delimitations in the Dicranum acutifolium complex (Dicranaceae, Bryophyta) using molecular markers. J Bryol 36:279–290. https://doi.org/10.1179/1743282014Y.0000000119
Lang AS, Bocksberger G, Stech M (2015) Phylogeny and species delimitations in European Dicranum (Dicranaceae, Bryophyta) inferred from nuclear and plastid DNA. Mol Phylogenet Evol 92:217–225. https://doi.org/10.1016/j.ympev.2015.06.019
Le S, Josse J, Husson F (2008) FactoMineR: an R package for multivariate analysis. J Stat Softw 25:1–18. https://doi.org/10.18637/jss.v025.i01
Limpricht KG (1886) Die Laubmoose Deutschlands, Oesterreichs und der Schweiz. I. Abteilung: Sphagnaceae, Andreaeaceae, Archidiaceae, Bryinae. Lieferung VI. Eduard Kummer, Leibzig
Linde AM, Sawangproh W, Cronberg N et al (2020) Evolutionary history of the Marchantia polymorpha complex. Front Plant Sci 11:1–16. https://doi.org/10.3389/fpls.2020.00829
Lüth M (2019) Mosses of Europe—a photograhic flora (3 Volumes). M. Lüth, Freiburg i. B.
McDaniel SF, von Stackelberg M, Richardt S et al (2010) The speciation history of the Physcomitrium - Physcomitrella species complex. Evolution (n Y) 64:217–231. https://doi.org/10.1111/j.1558-5646.2009.00797.x
Meinunger L, Schröder W (2007) Verbreitungsatlas der Moose Deutschlands. Regensburgische Botanische Gesellschaft, Regensburg
Meleshko O, Stenøien HK, Speed JDM et al (2018) Is interspecific gene flow and speciation in peatmosses (Sphagnum) constrained by phylogenetic relationship and life-history traits? Lindbergia 41:1–14. https://doi.org/10.25227/linbg.01107
Meleshko O, Martin MD, Korneliussen TS et al (2021) Extensive genome-wide phylogenetic discordance is due to incomplete lineage sorting and not ongoing introgression in a rapidly radiated bryophyte genus. Mol Biol Evol. https://doi.org/10.1093/molbev/msab063
Mitrofanov FP, Pozhilenko VI, Smolkin VF et al (1995) Geology of the Kola Peninsula (Baltic Shild). Russian Akademy of Sciences Kola Science Centre Geological Institute, Apatity
Mönkemeyer W (1927) Die Laubmoose Europas. Rabenhorst’s Kryptogamen-Flora. 4. Erg. Bd. Akademische Verlagsgesellschaft, Leibzig
Nieto-Lugilde M, Werner O, McDaniel SF et al (2018) Peripatric speciation associated with genome expansion and female-biased sex ratios in the moss genus Ceratodon. Am J Bot 105:1009–1020. https://doi.org/10.1002/ajb2.1107
Nyholm E (1954) Illustrated moss flora of Fennoscandia. II. Musci. CWK Gleerup, Lund
Nyholm E (1987) Illustrated flora of Nordic mosses. Fasc. 1. Fissidentaceae-Seligeriaceae. Nordic Bryological Society, Copenhagen and Lund
Pagès J (2004) Analyse factorielle de donnees mixtes. Revue Statistique Appliquee 4:93–111
Portnyagin M, Duggen S, Hauff F et al (2015) Geochemistry of the late Holocene rocks from the Tolbachik volcanic field, Kamchatka: quantitative modelling of subduction-related open magmatic systems. J Volcanol Geothermal Res 307:133–155. https://doi.org/10.1016/j.jvolgeores.2015.08.015
R Core Team (2022) R: a language and environment for statistical computing. https://www.r-project.org/
Rambaut A, Drummond AJ, Xie D et al (2018) Posterior summarization in Bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Reis CA, Dias C, Araripe J et al (2020) Multilocus data of a manakin species reveal cryptic diversification moulded by vicariance. Zool Scr 49:129–144. https://doi.org/10.1111/zsc.12395
Renner MAM (2020) Opportunities and challenges presented by cryptic bryophyte species. Telopea (syd) 23:41–60. https://doi.org/10.7751/TELOPEA14083
Renner MAM, Brown EA, Wardle GM (2013) Averaging v. outlier removal. Decrypting variance among cryptic Lejeunea species (Lejeuneaceae: Jungermanniopsida) using geometric morphometrics. Aust Syst Bot 26:13–30. https://doi.org/10.1071/SB12016
Rockenschaub M, Nowotny A (2009) Geologische Karte der Republik Österreich 1:50.000; Nr. 148, Brenner. Geologische Bundesanstalt, Wien
Ronquist F, Teslenko M, Van Der Mark P et al (2012) MrBayes 3.2: Efficient bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61: 39–542. https://doi.org/10.1093/sysbio/sys029
Schlüsslmayr G (2019) Die Moose des Dachsteingebirges. Stapfia 108:1–737
Shaw JA (2001) Biogeographic patterns and cryptic speciation in bryophytes. J Biogeogr 28:253–261. https://doi.org/10.1046/j.1365-2699.2001.00530.x
Sievert C (2020) Interactive web-based data visualization with R, plotly, and shiny. Chapman and Hall/CRC, Boca Raton
Simmons MP, Ochoterena H (2000) Gaps as characters in sequence-based phylogenetic analyses. Syst Biol 49:369–381. https://doi.org/10.1093/sysbio/49.2.369
Spagnuolo V, Caputo P, Cozzolino S et al (1999) Patterns of relationships in Trichostomoideae (Pottiaceae, Musci). Plant Syst Evol 216:69–79. https://doi.org/10.1007/BF00985101
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Stech M, Frahm J-P (1999) The status of Platyhypnidium mutatum Ochyra & Vanderpoorten and the systematic value of the Donrichardsiaceae based on molecular data. J Bryol 21:191–195. https://doi.org/10.1179/jbr.1999.21.3.191
Stöver BC, Müller KF (2010) TreeGraph 2: combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform 11:1–9. https://doi.org/10.1186/1471-2105-11-7
Struck TH, Feder JL, Bendiksby M et al (2018) Finding evolutionary processes hidden in cryptic species. Trends Ecol Evol 33:153–163. https://doi.org/10.1016/j.tree.2017.11.007
Suzuki T (2016) A revised new catalog of the mosses of Japan. Hattoria 7:9–223
Swisstopo, (2021) Landesgeologie - Service géologique national. Liste der erhältlichen Publikationen - Liste des publications disponibles, Bundesamt für Landestopografie Swisstopo, Wabern
Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17:1105–1109. https://doi.org/10.1007/BF00037152
Tubanova DY, Goryunov DV, Ignatova EA, Ignatov MS (2010) On the taxonomy of Dicranum acutifolium and D. fuscescens complexes (Dicranaceae, Bryophyta) in Russia. Arctoa 19:151–164. https://doi.org/10.15298/arctoa.19.13
Urmi E (2010) Teil II. Bryophyta (Moose). In: Landolt E (ed) Flora Indicativa. Ökologische Zeigerwerte und biologische Kennzeichen zur Flora der Schweiz und der Alpen. Haupt, Bern, pp 283–310
van Zuijlen K, Nobis MP, Hedenäs L et al (2023) Bryophytes of Europe Traits (BET) dataset: a fundamental tool for ecological studies. J Veg Sci 34:1–7. https://doi.org/10.1111/jvs.13179
Werner O, Ros RM, Guerra J (2002) Direct amplification and NaOH extraction: two rapid and simple methods for preparing bryophyte DNA for polymerase chain reaction (PCR). J Bryol 24:127–131. https://doi.org/10.1179/037366802125000980
Wright ES (2015) DECIPHER: harnessing local sequence context to improve protein multiple sequence alignment. BMC Bioinform 16:1–14. https://doi.org/10.1186/s12859-015-0749-z
Acknowledgements
We warmly thank H. Köckinger for the morphological examination of specimens and valuable comments on the manuscript and we acknowledge H. Hofmann, M. Lüth, H. Köckinger, M. Meier, N. Schnyder, E. Urmi, A. Vanderpoorten and the curators of GJO, L, W and Z + ZT for screening their collections and providing specimens, L. Hedenäs for giving substrate information of the Scandinavian specimens, E. Ignatova for providing a sequence of Dicranum acutifolium, S. Royek for her help with the morphological measurements and A. Hodgson for his linguistic editing. PS is grateful for funding within the framework of MAdLand (http://madland.science; project-number 440370236; PSLJ1111/1), priority programme 2237 of the German Research Foundation (DFG). Additional funding was received by PS from the Swiss National Science Foundation (grant nos. 160004, 184826, and 212509), the University Research Priority Program “Evolution in Action” of the University of Zurich and the Georges and Antoine Claraz Foundation.
Funding
Open access funding provided by University of Zurich.
Author information
Authors and Affiliations
Contributions
TK designed the study, generated and analysed the data and wrote the first manuscript. PS contributed to the conception and writing of the MS.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kiebacher, T., Szövényi, P. Morphological, genetic and ecological divergence in near-cryptic bryophyte species widespread in the Holarctic: the Dicranum acutifolium complex (Dicranales) revisited in the Alps. J Plant Res 137, 561–574 (2024). https://doi.org/10.1007/s10265-024-01534-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-024-01534-3