Abstract
Japanese giant salamander (Andrias japonicus) is one of the largest amphibian species in the world and an iconic species in Japan. However, as its distribution has recently declined across the country, rapid and extensive monitoring of the distribution is urgently needed for its efficient conservation. Here, we used environmental DNA (eDNA) analysis to assess the Japanese giant salamander’s distribution in western Japan and, for that purpose, we collected 410 water samples from 12 rivers. We then developed a new eDNA assay for multi-copy nuclear DNA (nuDNA) of the giant salamander and compared the eDNA detectability of the nuDNA marker with that of a previous mitochondrial DNA (mtDNA) marker. Throughout the survey, we detected target eDNA from 162 water samples using either of the markers, which generally corresponded to the known natural distribution of the species. Additionally, the use of the nuDNA marker allowed for higher detection rate of target eDNA than the mtDNA marker. Moreover, the detection rate of target eDNA decreased substantially in water samples with higher conductivity and also partly in those with higher pH, suggesting their negative impacts on the salamander’s ecology. Our results demonstrated that eDNA analysis with multi-copy nuDNA marker is highly useful for efficient and sensitive surveillance of Japanese giant salamander’s distribution. Our study provided the methodology for efficiently monitoring the Japanese giant salamander’s distribution via eDNA analysis and facilitating conservation activities for them.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Japanese giant salamander (Andrias japonicus) is one of the largest amphibian species in the world, with a maximum body length of 1.5 m, and has been registered as a special Japanese natural treasure since 1952 (https://kunishitei.bunka.go.jp/heritage/detail/401/3129). This iconic salamander is distributed in streams with relatively moderate slopes in western Japan and reproduces from August to September (Okada et al. 2015). The giant salamander’s eggs hatch in winter and the larvae metamorphose after approximately 3 years. Sexual maturity is considered to take 8 years or more (Browne et al. 2014). Japanese giant salamander forages mainly on crustaceans and small fish, reaching at the top of the trophic chain in stream ecosystems in Japan (Browne et al. 2014; Matsui and Hayashi 1992; Matsui et al. 2008). However, recent climate and land use changes, as well as competition with the alien Chinese-originated Andrias spp. (e.g., Andrias davidianus and Andrias sligoi; Turvey et al. 2019), can result in habitat degradation and reduction of distribution for Japanese giant salamander (Browne et al. 2014; Fukumoto et al. 2015; Matsui et al. 2008). Moreover, introgression due to hybridization between the endemic and alien giant salamanders causes genetic contamination and can further decrease the population of the endemic giant salamander (Matsui et al. 2008; Yoshikawa et al. 2011).
Considering these crises, the Japanese giant salamander is now listed as vulnerable in both the International Union for Conservation of Nature and Natural Resources (IUCN) and the Japanese National Red List (Ministry of the Environment 2020). Japanese giant salamanders have no land phase, hide in riverbed gaps between rocks or in underwater halls in vegetated riverbanks during the day, and are active at night (Fukumoto et al. 2015). Owing to their aquatic and nocturnal features, conventional capture-based surveys are time-consuming, labor-intensive, and sometimes dangerous for researchers. In addition, capture-based surveys are potentially harmful to individuals and their habitats. Therefore, it is unlikely that these conventional methods can sufficiently assess the giant salamander’s current distribution and abundance, which limits the availability of information pertaining sites where conservation of the giant salamander should be prioritized.
Environmental DNA (eDNA; extra-organismal DNA fragments in aquatic and terrestrial environments) analysis has recently attracted considerable attention as a novel biomonitoring tool that can complement conventional methods (Clare et al. 2022; Deiner et al. 2017; Minamoto et al. 2012; Yao et al. 2022). In this analysis, target organisms’ presence and distribution can easily be estimated by detecting their eDNA fragments via PCR amplification. Thus, eDNA analysis allows for noninvasive, cost-effective, and sensitive biomonitoring (Czeglédi et al. 2021; Darling and Mahon 2011; Fediajevaite et al. 2021; Jo et al. 2020b) and several studies have demonstrated the usefulness of eDNA analysis for the surveillance of amphibian species (Bálint et al. 2018; Biggs et al. 2015; Ficetola et al. 2008; Iwai et al. 2019; Jo et al. 2020c; Li et al. 2021; Pilliod et al. 2013). Fukumoto et al. (2015) previously used eDNA analysis to survey the distribution of Japanese giant salamanders along the Katsura River basin, Japan, demonstrating the suitability of the analysis for rapid and extensive biomonitoring. Fukumoto et al. (2015) also stated that further application of eDNA analysis over a broader spatial scale would enhance its usefulness in the context of aquatic conservation.
Expanding the eDNA application in Fukumoto et al. (2015), the present study conducted an eDNA-based surveillance of Japanese giant salamander’s distribution in western Japan. We collected 410 water samples from 12 rivers and analyzed the presence or absence of target eDNA in the water samples. Although most eDNA studies have targeted mitochondrial DNA (mtDNA) as a genetic marker (Biggs et al. 2015; Jerde et al. 2011; Sakata et al. 2021), recent studies suggested the use of nuclear DNA (nuDNA), particularly multiple copies of ribosomal RNA (rRNA) genes, as a more sensitive genetic marker for eDNA analyses than mtDNA (Dysthe et al. 2018; Jo et al. 2020a; Minamoto et al. 2017). Despite multiple mitochondrial genomes per cell (tens to thousands of mtDNA copies), the multi-copy nuDNA marker has exhibited similar or higher eDNA detection sensitivity likely due to the tandem-repeated sequences of rRNA genes in eukaryotic cells (Bylemans et al. 2017; Jo et al. 2020a; Minamoto et al. 2017). We thus designed a novel eDNA assay for the multi-copy nuDNA of the Japanese giant salamander and compared the detection rate of target eDNA between mtDNA and nuDNA markers. We further examined the relationship between target eDNA detection rate and abiotic parameters to discuss the effects of environmental conditions on Japanese giant salamander distribution and their eDNA detection.
Methods
Field survey and water sampling
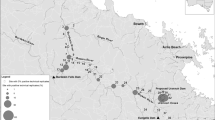
We collected water samples from 336 sites along 12 river systems in western Japan from June to December 2014 and June to November 2015 (Fig. 1). Water sampling was conducted three times in the Hatsuka River (Hyogo Pref.), twice in the Sayo River (Hyogo Pref.), and once in other rivers, resulting in a total of 410 water samples (Table S1). The rivers surveyed in our study were located in prefectures where the presence of the target species was confirmed by a previous national census and an individual study (Biodiversity Center of Japan 2001; Matsui et al. 2008). Considering the potential downstream transport of eDNA particles (Jo and Yamanaka 2022), most sampling sites were located 5–10 km apart along the mainstem, with some exceptions at the confluence of the tributaries. For each survey, we collected water samples using a 1-L plastic container from the river surface. The plastic containers were in advance carefully bleached with 0.1% sodium hypochlorite solution for at least 5 min (Jo et al. 2020b) and washed twice with the river water immediately before water sampling. Disposable gloves were put on during water sampling and were replaced at each sampling site. When collecting the water samples, environmental parameters (water temperature, pH, and electrical conductivity [EC]) were measured using Hanna Combo 2 (HI 98130; Hanna Instruments, Japan) at each site (Table S1). Water samples were transported to the laboratory (Kobe University, Hyogo, Japan) at low temperatures in a cooler box and filtered within a day following sampling (Jo et al. 2020c), except for water samples from Maruyama River, which were frozen and filtered a few days after sampling.
Map of all water sampling sites in this study with a photograph of Japanese giant salamander (at night survey in the Sayo River). Water samples were collected from 336 sites at 12 rivers in western Japan (shown as circles). Note that the detailed location with and without target eDNA detection is not disclosed because of the rarity and conservation importance of the target species
We filtered water samples using a 47-mm diameter glass microfiber filter GF/F (nominal pore size 0.7 μm; GE Healthcare Life Science, Little Chalfont, UK). Each 1 L of water sample was filtered through one or two GF/F filters, depending on filter clogging. In each river system, one or two 1-L units of distilled water were filtered as a filtration negative control to assess the potential for contamination of target eDNA during and after water filtration. Before and after every use, all filtering equipment (i.e., filter funnels [Magnetic Filter Funnel, 500 mL capacity; Pall Corporation, Westborough, MA, U.S.], 1-L beakers, and tweezers) were bleached with 0.1% sodium hypochlorite solution for at least 5 min. All filtered samples were stored at − 20 °C until DNA extraction.
DNA extraction and real-time PCR
Total eDNA on the filter was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Germany), following the method described by Minamoto et al. (2019). The presence of Japanese giant salamander eDNA in the water samples was evaluated using the StepOnePlus Real-Time PCR system (Applied Biosystems). We designed the primers and probe set for the nuclear DNA of the Japanese giant salamander, which amplifies 133 bp fragments of rRNA genes ranging from 18S to internal transcribed spacer-1 (ITS1) regions (Table 1; Table S2; Appendix S1). Each 20 μL TaqMan reaction contained 2 μL of template DNA, a final concentration of 900 nM of forward and reverse primers, and 125 nM of TaqMan probe in a 1 × TaqMan Gene Expression Master Mix (Thermo Fisher, U.S.). Thermal conditions of real-time PCR were as follows: 2 min at 50 °C, 10 min at 95 °C, and 55 cycles of 15 s at 95 °C and 60 s at 60 °C. Exuviae-derived DNA from target species (0.2 ng/template) and 2 μL of pure water were simultaneously analyzed as PCR-positive and -negative controls, respectively. All PCRs for the eDNA extracts, positive controls, and negative controls were performed in four replicates. Target eDNA was considered present in each water sample if any of the PCR replicates were clearly amplified. The primers and probe set developed in Fukumoto et al. (2015) was used as the assay for amplifying mtDNA of the target species (Table 1). The water samples collected in 2014 were analysed using both mtDNA and nuDNA markers, while those collected in 2015 were analyzed using only nuDNA markers because of the higher eDNA detection rate of the nuDNA marker (see below).
We estimated the limit of detection (LOD; defined as the lowest quantity of target DNA that could be detected in one of the PCR replicates) of each genetic marker. The LOD is based on detection/non-detection criteria and represents the ability of an assay to detect the low concentrations of target DNA fragments (Klymus et al. 2020). A dilution series of the A. japonicus tissue DNA (10–3–102 pg/2 µL PCR template) was prepared for estimating the LODs. PCRs for the dilution series were performed in six replicates and other PCR conditions were similar as described above.
Statistical analyses
All analyses were performed using R version 4.2.1 (R Core Team 2022) and the significance level (α) was set at 0.05. We compared the detection rate of target eDNA between mtDNA and nuDNA markers using water samples collected in 2014 (n = 246). We used a generalized linear mixed model (GLMM) with a binomial distribution to assess the effects of genetic markers and environmental parameters (EC and pH) on target eDNA detection using the lmerTest package (Kuznetsova et al. 2017). In the GLMM, the presence/absence of target eDNA in each water sample (1/0) was included as a dependent variable, and the type of genetic marker (mtDNA or nuDNA), pH, and EC were included as fixed effects. Given that water sampling was replicated in some rivers, each river was included as a random effect, and site IDs were nested within the river groups. Additionally, using the water samples collected in 2015 (n = 164), we also performed a binomial GLMM to assess the effects of environmental parameters (pH and EC) on target eDNA detection using the same formula as above (except for using only nuDNA markers). For both GLMMs, we confirmed that multicollinearity among the variables was negligible (variance inflation factor, VIF = 1.008–1.050). Water temperature was not used for the analyses because of its temporal variability within a day (Table S1).
Results
All PCR-positive controls were successfully amplified, and no filtration and PCR-negative controls were amplified throughout the study (Table S1). The LODs were estimated at 0.01 pg (per reaction) for both the genetic markers (Table S3). However, when 0.1 and 0.01 pg of tissue DNA were used, four and one replicates were amplified for the mtDNA marker and six and four replicates were amplified for the nuDNA marker.
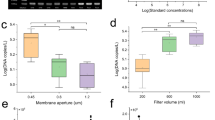
In 2014 surveys, we collected 246 water samples in total from seven rivers and detected Japanese giant salamander eDNA in 74 samples using nuDNA marker (74/246 = 30.1%), 48 samples using mtDNA markers (48/246 = 19.5%), and 85 samples using either of the markers (85/246 = 34.6%; Table 2). Of the 246 samples, 37 showed target eDNA detections with both markers, 37 showed target eDNA detections with only the nuDNA marker, and 11 showed target eDNA detections with only the mtDNA marker. In contrast, the target eDNA was not detected in 161 samples using either of the markers (Table S1; results of target eDNA detections from each river are summarized in Fig. S1). The number of samples with target eDNA detection was higher for nuDNA than for mtDNA markers in all surveyed rivers. A binomial GLMM based on the 2014 dataset showed a higher eDNA detection rate of the nuDNA marker (P < 0.001) and significant negative effects of pH (P < 0.001) and EC (P = 0.024) on eDNA detection (Table 3). For the samples with eDNA detections, the pH and EC values were 7.08 ± 0.57 and 0.056 ± 0.031, respectively (Fig. 2). For the samples without eDNA detections, by contrast, the pH and EC values were 7.54 ± 0.74 and 0.076 ± 0.044, respectively.
In 2015 surveys, we collected 164 water samples in total from six rivers and detected Japanese giant salamander eDNA in 76 samples using nuDNA marker (76/164 = 46.3%; Table 2). A binomial GLMM based on the 2015 dataset showed a significantly negative effect of EC on eDNA detection (P < 0.001), whereas the pH effect was not significant (P = 0.673; Table 3; Fig. 3). For the samples with eDNA detections, the pH and EC values were 7.26 ± 0.73 and 0.051 ± 0.023, respectively. For the samples without eDNA detections, the pH and EC values were 7.31 ± 0.60 and 0.087 ± 0.037, respectively. In summary, during the 2-year survey from 2014 to 2015, target eDNA was detected in 150 of the 410 water samples (36.6%) using nuDNA markers and from 162 of the 410 water samples (39.5%) using either of the genetic markers (Table S1).
Discussion
We succeeded in estimating the distributional information of Japanese giant salamanders in western Japan using eDNA analysis. The estimated distribution of the Japanese giant salamander generally corresponded to the known natural distribution of the species, as reported in previous studies (Biodiversity Center of Japan 2001; Matsui et al. 2008). Although Fukumoto et al. (2015) examined the distribution of Japanese giant salamander using eDNA analysis in a single river system over two days, our study was able to survey the distribution much more broadly over 19 days, further indicating the usefulness of eDNA analysis for rapid and extensive surveillance of aquatic rare species distribution. Moreover, we showed a higher sensitivity of the nuDNA marker for detecting giant salamander eDNA, which would allow for efficiently surveying the Japanese giant salamander’s entire distribution using eDNA analysis and facilitating their conservation.
The results of a binomial GLMM showed that the nuDNA marker developed in this study yielded higher detection of Japanese giant salamander eDNA than the mtDNA marker developed by Fukumoto et al. (2015). The result was also supported by its higher detection sensitivity in the LOD test, where the target DNA detection rate with low quantity (0.1 and 0.01 pg) was higher for the nuDNA marker than the mtDNA marker. Given that the previous reports have shown higher fish eDNA detection sensitivity through multi-copy nuDNA markers (Dysthe et al. 2018; Minamoto et al. 2017), our results supported the higher eDNA detection sensitivity of multi-copy nuDNA markers and indicated their usefulness for efficient eDNA-based biomonitoring for amphibian species.
Alternatively, the result may simply be accounted for by the difference in PCR amplicon lengths between the markers (116 bp for nuDNA and 133 bp for mtDNA) because longer DNA fragments are expected to degrade faster and persist shorter in the water (Hänfling et al. 2016; Jo et al. 2017). However, it is unlikely that such small differences in PCR amplification lengths between the markers would significantly affect the yields of target eDNA (Bylemans et al. 2018; Jo et al. 2020a). Altogether, the multi-copy nuDNA marker would allow for reducing the risk of false-negative eDNA detections and sensitively estimating the salamander’s distribution.
The binomial GLMM results also showed a significant negative effect of EC on the detection of the Japanese giant salamander eDNA in both years. High EC is an index of water quality decline, relating to the total dissolved solids and ion concentration in the water and negatively impacting the occurrence, abundance, and reproductive success of aquatic species including some salamanders and hellbenders (Alavi and Cosson 2006; Bodinof Jachowski et al. 2016; Bowles et al. 2006; Keitzer et al. 2013). Some studies have previously documented negative relationships between EC and detection rate of fish and amphibian eDNA (Jo et al. 2020b, c; Pitt et al. 2017), which supports our findings. Alternatively, high EC values can also relate to PCR inhibition (Schrader et al. 2012) and prevent the eDNA detection (Harper et al. 2019; Wineland et al. 2019), although Pitt et al. (2017) confirmed no evidence of PCR inhibition in their river water samples with higher EC values (> 0.5 mS/cm) than ours (< 0.25 mS/cm). We thus conclude that high EC is likely affecting the giant salamander’s occurrence at some sites, rather than detections of the target eDNA when the species are present there. This problem can be addressed to some extent by using a site occupancy model, which separates the probability of eDNA occurrence (not species occurrence) from that of eDNA detection (Jo et al. 2020c; Wineland et al. 2019), though the site occupancy modeling could not be applied to this study due to a lack of sampling replicates per survey site.
By contrast, the effect of pH on the eDNA detection was inconsistent between the survey years. In the 2014 survey, Japanese giant salamander eDNA tended to be detected less frequently in samples with higher pH. A higher pH (> 8.5) is also considered to reflect water quality (Boczkaj and Fernandes 2017) and can be observed in rivers with high algal productivity and limestone rocks (Bhateria and Jain 2016). However, it is less likely that eDNA persistence varies within the range of pH measured in this study (Jo et al. 2022; Strickler et al. 2015). Thus, the result may also indicate that alkaline conditions affect the giant salamander’s occurrence rather than target eDNA detection. For example, eDNA detection rate was low on average in the Sayo River, which had the highest pH and EC among the surveyed rivers. In contrast, the effect of pH on target eDNA detection was not significant in the samples collected in 2015. It is not clear what caused this difference, but this implies that, although EC was a strong variable determining the giant salamander’s distribution, a higher pH may not always be observed at sites that the salamanders do not prefer to inhabit. Their distribution can also be determined by other environmental factors (e.g., annual mean temperature, precipitation, elevation, land use, and geological features; Houlahan et al. 2000; Okada et al. 2008; Willson and Dorcas 2003). Precipitation can be related to the flow condition of the river, likely affecting the habitat suitability of river-dwelling species, including the target species, and land use and elevation can be related to anthropogenic impacts on amphibian habitats (Johnson et al. 2011). Other environmental parameters, such as biochemical oxygen demand (BOD) and dissolved organic carbon (DOC), may also associate with the giant salamander’s survival and distribution in the studied rivers. Future studies should measure relevant environmental variables simultaneously with eDNA sampling (Jo and Yamanaka 2022).
Although the presence/absence of target eDNA in a water sample was the focus of this study, we may have been able to infer a more detailed ecology of the giant salamanders in the studied rivers by quantifying target eDNA concentrations. Environmental DNA concentrations can represent their relative abundance and activity in the environment (Iwai et al. 2019; Jo et al. 2020a; Pilliod et al. 2013; Spear et al. 2015). Continuous eDNA-based quantitative monitoring will inform time-series changes not only regarding the giant salamander’s distribution but also their relative abundance in rivers. Moreover, as external fertilization (Kawamichi and Ueda 1998) can increase the relative concentration of nuDNA to mtDNA in the water, the ratio of nuclear to mitochondrial eDNA concentrations could be used to estimate the timing and location of their spawning (Bylemans et al. 2017; Wu et al. 2022). These efforts will advance our understanding of the life history and reproductive ecology of giant salamanders, providing useful information for conservation of endangered aquatic species with external fertilization.
For the practical use of nuDNA in eDNA-based biomonitoring, the limitation of nuclear gene sequences available in databases (e.g., GenBank) is a primary drawback, which is especially the case for non-model organisms (Jo et al. 2022; Minamoto et al. 2017; Sigsgaard et al. 2020). Phylogenetics studies targeting bacteria and fungi have mainly used nuclear ribosomal DNA (rDNA) sequences (Handelsman 2004; Toju et al. 2012) and accordingly a significant amount of them has been accumulated. In contrast, studies targeting macro-organism’s (e)DNA including fish and amphibians have mainly used mtDNA fragments and research interest in nuDNA is still limited. Some recent studies have challenged eDNA-based genetic diversity monitoring targeting aquatic vertebrates, but most of them were also based on mtDNA (Sigsgaard et al. 2016; Wakimura et al. 2023; but see Andres et al. 2021). While recent substantial advance of next-generation sequencing (NGS) technology has drastically reduced sequencing costs (Goodwin et al. 2016), achieving database enrichment and developing a framework for sharing data among the science communities will require further collaborative efforts of multiple researchers and laboratories in the future.
Including our study, eDNA analysis has enabled the collection of broad-scale species distribution data on a considerably shorter timescale compared to conventional methods (Biggs et al. 2015; Deiner et al. 2017; Yao et al. 2022). Such an advantage of eDNA analysis can be quite useful not only for a rapid understanding of the species’ distribution and abundance, but also for revealing the relationship between their suitable habitats and environmental conditions. A few studies recently applied eDNA analysis to species distribution models (SDMs; a.k.a. ecological niche models) to link species occurrence records with environmental conditions and then estimated habitat suitability, defined as the occurrence probability of a species at a site with a given environmental condition (Hashemzadeh Segherloo et al. 2022; Riaz et al. 2020; Wilcox et al. 2018). Predicting species distribution has become increasingly important for its conservation owing to the impacts of recent climate change, including global warming caused by anthropogenic CO2 emissions. These effects are considered to cause habitat shifts and disturbances for various animals and plants, which may further accelerate in the future (Butchart et al. 2010; Cardinale et al. 2012; Ceballos et al. 2015). Combined with statistical approaches such as SDMs, eDNA analysis can contribute to biodiversity conservation and ecosystem management. Such studies would help to preserve the habitats of various rare species, including the giant salamander, and save them from extinction.
Data accessibility
The raw data from the real-time PCR experiments are included in the Supplemental Information.
References
Alavi SMH, Cosson J (2006) Sperm motility in fishes. (II) Effects of ions and osmolality: a review. Cell Biol Int 30:1–14
Andres KJ, Sethi SA, Lodge DM, Andrés J (2021) Nuclear eDNA estimates population allele frequencies and abundance in experimental mesocosms and field samples. Mol Ecol 30:685–697
Bálint M, Nowak C, Márton O, Pauls SU, Wittwer C, Aramayo JL, Schulze A, Chambert T, Cocchiararo B, Jansen M (2018) Accuracy, limitations and cost efficiency of eDNA-based community survey in tropical frogs. Mol Ecol Resour 18:1415–1426
Bhateria R, Jain D (2016) Water quality assessment of lake water: a review. Sustain Water Resour Manage 2:161–173
Biggs J, Ewald N, Valentini A, Gaboriaud C, Dejean T, Griffiths RA, Foster J, Wilkinson JW, Arnell A, Brotherton P, Williams P, Dunn F (2015) Using eDNA to develop a national citizen science-based monitoring programme for the great crested newt (Triturus cristatus). Biol Conserv 183:19–28
Biodiversity Center of Japan, Ministry of the Environment (2001) 5th National basic survey on natural environment. https://www.biodic.go.jp/reports2/5th/ap_rep/5_ap_rep.pdf. Accessed 6 December 2022
Boczkaj G, Fernandes A (2017) Wastewater treatment by means of advanced oxidation processes at basic pH conditions: a review. J Chem Eng 320:608–633
Bodinof Jachowski CM, Millspaugh JJ, Hopkins WA (2016) Current land use is a poor predictor of hellbender occurrence: why assumptions matter when predicting distributions of data-deficient species. Divers Distrib 22:865–880
Bowles BD, Sanders MS, Hansen RS (2006) Ecology of the Jollyville Plateau salamander (Eurycea tonkawae: Plethodontidae) with an assessment of the potential effects of urbanization. Hydrobiologia 553:111–120
Browne RK, Li H, Wang Z, Okada S, Hime P, McMillan A, Wu M, Diaz R, McGinnity D, Briggler JT (2014) The giant salamanders (Cryptobranchidae): part B. Biogeography, ecology and reproduction. Amphib Reptile Conserv 5:30–50
Butchart SH, Walpole M, Collen B, Van Strien A, Scharlemann JP, Almond RE, Baillie JE, Bomhard B, Brown C, Bruno J, Carpenter KE, Carr GM, Chanson J, Chenery AM, Csirke J, Davidson NC, Dentener F, Foster M, Galli A, Galloway JN, Genovesi P, Gregory RD, Hockings M, Kapos V, Lamarque J, Leverington F, Loh J, McGeoch MA, McRae L, Minasyan A, Morcillo MH, Oldfield TE, Pauly D, Quader S, Revenga C, Sauer JR, Skolnik B, Spear D, Stanwell-Smith D, Stuart SN, Symes A, Tierney M, Tyrrell TD, Vié J, Watson R (2010) Global biodiversity: indicators of recent declines. Science 328:1164–1168
Bylemans J, Furlan EM, Hardy CM, McGuffie P, Lintermans M, Gleeson DM (2017) An environmental DNA-based method for monitoring spawning activity: a case study, using the endangered Macquarie perch (Macquaria australasica). Methods Ecol Evol 8:646–655
Bylemans J, Furlan EM, Gleeson DM, Hardy CM, Duncan RP (2018) Does size matter? An experimental evaluation of the relative abundance and decay rates of aquatic environmental DNA. Environ Sci Technol 52:6408–6416
Cardinale BJ, Duffy JE, Gonzalez A, Hooper DU, Perrings C, Venail P, Narwani A, Mace GM, Tilman D, Wardle DA, Kinzig AP, Daily GC, Loreau M, Grace JB, Larigauderie A, Srivastava DS, Naeem S (2012) Biodiversity loss and its impact on humanity. Nature 486:59–67
Ceballos G, Ehrlich PR, Barnosky AD, García A, Pringle RM, Palmer TM (2015) Accelerated modern human–induced species losses: entering the sixth mass extinction. Sci Adv 1:e1400253
Clare EL, Economou CK, Bennett FJ, Dyer CE, Adams K, McRobie B, Drinkwater R, Littlefair JE (2022) Measuring biodiversity from DNA in the air. Curr Biol 32:693–700
Czeglédi I, Sály P, Specziár A, Preiszner B, Szalóky Z, Maroda Á, Pont D, Meulenbroek P, Valentini A, Erős T (2021) Congruency between two traditional and eDNA-based sampling methods in characterising taxonomic and trait-based structure of fish communities and community–environment relationships in lentic environment. Ecol Indic 129:107952
Darling JA, Mahon AR (2011) From molecules to management: adopting DNA-based methods for monitoring biological invasions in aquatic environments. Environ Res 111:978–988
Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière-Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, de Vere N, Pfrender ME, Bernatchez L (2017) Environmental DNA metabarcoding: transforming how we survey animal and plant communities. Mol Ecol 26:5872–5895
Dysthe JC, Franklin TW, McKelvey KS, Young MK, Schwartz MK (2018) An improved environmental DNA assay for bull trout (Salvelinus confluentus) based on the ribosomal internal transcribed spacer I. PLoS ONE 13:e0206851
Fediajevaite J, Priestley V, Arnold R, Savolainen V (2021) Meta-analysis shows that environmental DNA outperforms traditional surveys, but warrants better reporting standards. Ecol Evol 11:4803–4815
Ficetola GF, Miaud C, Pompanon F, Taberlet P (2008) Species detection using environmental DNA from water samples. Biol Lett 4:423–425
Fukumoto S, Ushimaru A, Minamoto T (2015) A basin-scale application of environmental DNA assessment for rare endemic species and closely related exotic species in rivers: a case study of giant salamanders in Japan. J Appl Ecol 52:358–365
Goodwin S, McPherson JD, McCombie WR (2016) Coming of age: ten years of next-generation sequencing technologies. Nat Rev Genet 17:333–351
Handelsman J (2004) Metagenomics: application of genomics to uncultured microorganisms. Microbiol Mol Biol Rev 68:669–685
Hänfling B, Lawson Handley L, Read DS, Hahn C, Li J, Nichols P, Blackman RC, Oliver A, Winfield IJ (2016) Environmental DNA metabarcoding of lake fish communities reflects long-term data from established survey methods. Mol Ecol 25:3101–3119
Harper LR, Griffiths NP, Lawson Handley L, Sayer CD, Read DS, Harper KJ, Blackman RC, Li J, Hänfling B (2019) Development and application of environmental DNA surveillance for the threatened crucian carp (Carassius carassius). Freshw Biol 64:93–107
Hashemzadeh Segherloo I, Tabatabaei SN, Abdolahi-Mousavi E, Hernandez C, Normandeau E, Laporte M, Boyle B, Amiri M, GhaedRahmati N, Hallerman E, Bernatchez L (2022) eDNA metabarcoding as a means to assess distribution of subterranean fish communities: Iranian blind cave fishes as a case study. Environ DNA 4:402–416
Houlahan JE, Findlay CS, Schmidt BR, Meyer AH, Kuzmin SL (2000) Quantitative evidence for global amphibian population declines. Nature 404:752–755
Iwai N, Yasumiba K, Takahara T (2019) Efficacy of environmental DNA to detect and quantify stream tadpoles of Odorrana splendida. Royal Soc Open Sci 6:181798
Jerde CL, Mahon AR, Chadderton WL, Lodge DM (2011) “Sight-unseen” detection of rare aquatic species using environmental DNA. Conserv Lett 4:150–157
Jo T, Yamanaka H (2022) Meta-analyses of environmental DNA downstream transport and deposition in relation to hydrogeography in riverine environments. Freshw Biol 67:1333–1343
Jo T, Murakami H, Masuda R, Sakata MK, Yamamoto S, Minamoto T (2017) Rapid degradation of longer DNA fragments enables the improved estimation of distribution and biomass using environmental DNA. Mol Ecol Resour 17:e25–e33
Jo T, Arimoto M, Murakami H, Masuda R, Minamoto T (2020a) Estimating shedding and decay rates of environmental nuclear DNA with relation to water temperature and biomass. Environ DNA 2:140–151
Jo T, Fukuoka A, Uchida K, Ushimaru A, Minamoto T (2020b) Multiplex real-time PCR enables the simultaneous detection of environmental DNA from freshwater fishes: a case study of three exotic and three threatened native fishes in Japan. Biol Invasions 22:455–471
Jo T, Tomita S, Kohmatsu Y, Osathanunkul M, Ushimaru A, Minamoto T (2020c) Seasonal monitoring of Hida salamander Hynobius kimurae using environmental DNA with a genus-specific primer set. Endanger Species Res 43:341–352
Jo TS, Tsuri K, Yamanaka H (2022) Can nuclear aquatic environmental DNA be a genetic marker for the accurate estimation of species abundance? Sci Nat 109:38
Johnson PT, McKenzie VJ, Peterson AC, Kerby JL, Brown J, Blaustein AR, Jackson T (2011) Regional decline of an iconic amphibian associated with elevation, land-use change, and invasive species. Conserv Biol 25:556–566
Kawamichi T, Ueda H (1998) Spawning at nests of extra-large males in the giant salamander Andrias japonicus. J Herpetol 32:133–136
Keitzer SC, Pauley TK, Burcher CL (2013) Stream characteristics associated with site occupancy by the eastern hellbender, Cryptobranchus alleganiensis alleganiensis, in southern West Virginia. Northeast Nat 20:666–677
Klymus KE, Merkes CM, Allison MJ, Goldberg CS, Helbing CC, Hunter ME, Jackson CA, Lance RF, Mangan AM, Monroe EM, Piaggio AJ, Stokdyk JP, Wilson CC, Richter CA (2020) Reporting the limits of detection and quantification for environmental DNA assays. Environ DNA 2:271–282
Kuznetsova A, Brockhoff PB, Christensen RH (2017) lmerTest package: tests in linear mixed effects models. J Stat Softw 82:1–26
Li W, Hou X, Xu C, Qin M, Wang S, Wei L, Wang Y, Liu X, Li Y (2021) Validating eDNA measurements of the richness and abundance of anurans at a large scale. J Animal Ecol 90:1466–1479
Matsui M, Hayashi T (1992) Genetic uniformity in the Japanese giant salamander, Andrias japonicus. Copeia 1992:232–235
Matsui M, Tominaga A, Liu WZ, Tanaka-Ueno T (2008) Reduced genetic variation in the Japanese giant salamander, Andrias japonicus (Amphibia: Caudata). Mol Phylogenet Evol 49:318–326
Minamoto T, Yamanaka H, Takahara T, Honjo MN, Kawabata Z (2012) Surveillance of fish species composition using environmental DNA. Limnology 13:193–197
Minamoto T, Uchii K, Takahara T, Kitayoshi T, Tsuji S, Yamanaka H, Doi H (2017) Nuclear internal transcribed spacer-1 as a sensitive genetic marker for environmental DNA studies in common carp Cyprinus carpio. Mol Ecol Resour 17:324–333
Minamoto T, Hayami K, Sakata MK, Imamura A (2019) Real-time polymerase chain reaction assays for environmental DNA detection of three salmonid fish in Hokkaido, Japan: application to winter surveys. Ecol Res 34:237–242
Ministry of the Environment, Japan (2020) Ministry of the Environment, Japan, Red list, 4th edn. http://www.env.go.jp/press/files/jp/114457.pdf. Accessed on 6 December 2020 (in Japanese)
Okada S, Utsunomiya T, Okada T, Felix ZI, Ito F (2008) Characteristics of Japanese giant salamander (Andrias japonicus) populations in two small tributary streams in Hiroshima Prefecture, western Honshu, Japan. Herpetol Conserv Biol 3:192–202
Okada S, Fukuda Y, Takahashi MK (2015) Paternal care behaviors of Japanese giant salamander Andrias japonicus in natural populations. J Ethol 33:1–7
Pilliod DS, Goldberg CS, Arkle RS, Waits LP (2013) Estimating occupancy and abundance of stream amphibians using environmental DNA from filtered water samples. Can J Fish Aquat Sci 70:1123–1130
Pitt AL, Shinskie JL, Tavano JJ, Hartzell SM, Delahunty T, Spear SF (2017) Decline of a giant salamander assessed with historical records, environmental DNA and multi-scale habitat data. Freshw Biol 62:967–976
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. Accessed 13 September 2023
Riaz M, Kuemmerlen M, Wittwer C, Cocchiararo B, Khaliq I, Pfenninger M, Nowak C (2020) Combining environmental DNA and species distribution modeling to evaluate reintroduction success of a freshwater fish. Ecol Appl 30:e02034
Sakata MK, Watanabe T, Maki N, Ikeda K, Kosuge T, Okada H, Yamanaka H, Sado T, Miya M, Minamoto T (2021) Determining an effective sampling method for eDNA metabarcoding: a case study for fish biodiversity monitoring in a small, natural river. Limnology 22:221–235
Schrader C, Schielke A, Ellerbroek L, Johne R (2012) PCR inhibitors—occurrence, properties and removal. J Appl Microbiol 113:1014–1026
Sigsgaard EE, Nielsen IB, Bach SS, Lorenzen ED, Robinson DP, Knudsen SW, Pedersen MW, Jaidah MA, Orlando L, Willerslev E, Møller PR, Thomsen PF (2016) Population characteristics of a large whale shark aggregation inferred from seawater environmental DNA. Nat Ecol Evol 1:0004
Spear SF, Groves JD, Williams LA, Waits LP (2015) Using environmental DNA methods to improve detectability in a hellbender (Cryptobranchus alleganiensis) monitoring program. Biol Conserv 183:38–45
Strickler KM, Fremier AK, Goldberg CS (2015) Quantifying effects of UV-B, temperature, and pH on eDNA degradation in aquatic microcosms. Biol Conserv 183:85–92
Toju H, Tanabe AS, Yamamoto S, Sato H (2012) High-coverage ITS primers for the DNA-based identification of ascomycetes and basidiomycetes in environmental samples. PLoS ONE 7:e40863
Turvey ST, Marr MM, Barnes I, Brace S, Tapley B, Murphy RW, Zhao E, Cunningham AA (2019) Historical museum collections clarify the evolutionary history of cryptic species radiation in the world’s largest amphibians. Ecol Evol 9:10070–10084
Wakimura K, Uchii K, Kikko T (2023) Evaluation of genetic diversity in an endangered fish Gnathopogon caerulescens using environmental DNA and its potential use in fish conservation. Environ DNA 5:973–986. https://doi.org/10.1002/edn3.408
Wilcox TM, Young MK, McKelvey KS, Isaak DJ, Horan DL, Schwartz MK (2018) Fine-scale environmental DNA sampling reveals climate-mediated interactions between native and invasive trout species. Ecosphere 9:e02500
Willson JD, Dorcas ME (2003) Effects of habitat disturbance on stream salamanders: implications for buffer zones and watershed management. Conserv Biol 17:763–771
Wineland SM, Welch SM, Pauley TK, Apodaca JJ, Olszack M, Mosher JJ, Holmes JN, Waldron JL (2019) Using environmental DNA and occupancy modelling to identify drivers of eastern hellbender (Cryptobranchus alleganiensis alleganiensis) extirpation. Freshw Biol 64:208–221
Wu L, Yamamoto Y, Yamaguchi S, Minamoto T (2022) Spatiotemporal changes in environmental DNA concentrations caused by fish spawning activity. Ecol Indic 142:109213
Yao M, Zhang S, Lu Q, Chen X, Zhang SY, Kong Y, Zhao J (2022) Fishing for fish environmental DNA: ecological applications, methodological considerations, surveying designs, and ways forward. Mol Ecol 31:5132–5164
Yoshikawa N, Kaneko S, Kuwabara K, Okumura N, Matsui M, Isagi Y (2011) Development of microsatellite markers for the two giant salamander species (Andrias japonicus and A. davidianus). Curr Herpetol 30:177–180
Acknowledgements
We thank Mr. Hiromune Mitsuhashi (Museum of Nature and Human Activities, Hyogo Pref.) for helping select the sampling sites in the Hatsuka River, the members of the Association to Protect the Giant Salamander of the Sayo River and Ms. Mayu Fukamachi (Ajimu High School, Oita Pref.) for helping select the sampling sites and water sampling in the Sayo and Yakkan Rivers, respectively, Dr. Yasuoki Takami (Kobe University) for providing the comments and suggestions regarding the experimental design and interpretation of the results, and Dr. Teruhiko Takahara (Shimane University) for providing A. japonicus tissue DNA sample. We also thank the two anonymous reviewers and an editor who provided helpful advice to improve the manuscript. This study was supported by the Grant-in-Aid for JSPS Research Fellows (Grant Numbers JP22J00439 and JP22KJ3043), the JSPS KAKENHI (Grant number: JP26440238), and 2014 Academic Research Grant from Hyogo Science and Technology Association.
Funding
Open Access funding provided by Kobe University.
Author information
Authors and Affiliations
Contributions
SH and TM designed the study. SH, SY, KRK, ST, AU, and TM conducted field sampling. SH, TSJ, and MM performed molecular experiments and analyzed the data. SH, TSJ, MM, MI, and TM discussed the results. SH, TSJ, and TM wrote the first draft of the manuscript. All authors edited and provided feedback on the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Handling Editor: Hideyuki Doi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hidaka, S., Jo, T.S., Yamamoto, S. et al. Sensitive and efficient surveillance of Japanese giant salamander (Andrias japonicus) distribution in western Japan using multi-copy nuclear DNA marker. Limnology 25, 189–198 (2024). https://doi.org/10.1007/s10201-023-00740-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10201-023-00740-7