Abstract
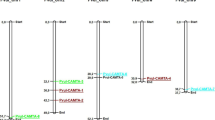
The precise regulation of gene expression is required for the determination of cell fate, differentiation, and developmental programs in eukaryotes. The Polycomb Group (PcG) genes are the key transcriptional regulators that constitute the repressive system, with two major protein complexes, Polycomb Repressive Complex 1 (PRC1) and Polycomb Repressive Complex 2 (PRC2). Previous studies have demonstrated the significance of these proteins in regulation of normal growth and development processes. However, the role of PcG in adaptation of crops to abiotic stress is still not well understood. The present study aimed to a comprehensive genome-wide identification of the PcG gene family in one of the economically important staple crops, Oryza sativa. Here, a total of 14 PcG genes have been identified, which were distributed over eight chromosomes. Protein structure analysis revealed that both the complexes have distinct domain and motifs that are conserved within the complexes. In silico promoter analysis showed that PcG gene promoters have abundance of abiotic stress-responsive elements. RNA-seq based expression analysis revealed that PcG genes are differentially expressed in different tissues and responded variably in different environmental stress. Validation of gene expression by qRT-PCR showed that most of the genes were upregulated at 1-h time point in shoot tissue and at 24-h time point in root tissue under the drought and salinity stress conditions. These findings provide important and extensive information on the PcG family of O. sativa, which will pave the path for understanding their role in stress signaling in plants.









Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Ach RA, Taranto P, Gruissem W (1997) A conserved family of WD-40 proteins binds to the retinoblastoma protein in both plants and animals. Plant Cell 9(9):1595–1606. https://doi.org/10.1105/tpc.9.9.1595
Agarwal P, Khurana P (2020) TaZnF, a C3HC4 type RING zinc finger protein from Triticum aestivum is involved in dehydration and salinity stress. J Plant Biochem Biotechnol 29(3):395–406. https://doi.org/10.1007/s13562-019-00546-8
Aggarwal S, Kumar A, Jain M, Sudan J, Singh K, Kumari S, Mustafiz A (2020) C-terminally encoded peptides (CEPs) are potential mediators of abiotic stress response in plants. Physiol Mol Biol Plants 26(10):2019–2033. https://doi.org/10.1007/s12298-020-00881-4
Aubert D, Chen L, Moon Y-H, Martin D, Castle LA, Yang C-H, Sung ZR (2001) EMF1, a novel protein involved in the control of shoot architecture and flowering in Arabidopsis. Plant Cell 13(8):1865–1876. https://doi.org/10.1105/TPC.010094
Baile F, Gómez-Zambrano Á, Calonje M (2022) Roles of Polycomb complexes in regulating gene expression and chromatin structure in plants. Plant Commun 3(1):100267. https://doi.org/10.1016/j.xplc.2021.100267
Bemer M, Grossniklaus U (2012) Dynamic regulation of Polycomb group activity during plant development. Curr Opin Plant Biol 15(5):523–529. https://doi.org/10.1016/j.pbi.2012.09.006
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B Methodol 57(1):289–300. https://doi.org/10.1111/j.2517-6161.1995.tb02031.x
Calonje M, Sanchez R, Chen L, Sung ZR (2008) EMBRYONIC FLOWER1 participates in polycomb group-mediated AG gene silencing in Arabidopsis. Plant Cell 20(2):277–291. https://doi.org/10.1105/tpc.106.049957
Chang Y-N, Zhu C, Jiang J, Zhang H, Zhu J-K, Duan C-G (2020) Epigenetic regulation in plant abiotic stress responses. J Integr Plant Biol 62(5):563–580. https://doi.org/10.1111/jipb.12901
Chanvivattana Y, Bishopp A, Schubert D, Stock C, Moon Y-H, Sung ZR, Goodrich J (2004) Interaction of Polycomb-group proteins controlling flowering in Arabidopsis. Development 131(21):5263–5276. https://doi.org/10.1242/dev.01400
Chen D, Huang Y, Ruan Y, Shen W-H (2016) The evolutionary landscape of PRC1 core components in green lineage. Planta 243(4):825–846. https://doi.org/10.1007/s00425-015-2451-9
Chen D, Molitor A, Liu C, Shen W-H (2010) The Arabidopsis PRC1-like ring-finger proteins are necessary for repression of embryonic traits during vegetative growth. Cell Res 20(12):1332–1344. https://doi.org/10.1038/cr.2010.151
Chen L-J, Diao Z-Y, Specht C, Sung ZR (2009) Molecular evolution of VEF-domain-containing PcG genes in plants. Mol Plant 2(4):738–754. https://doi.org/10.1093/mp/ssp032
Conrad LJ, Khanday I, Johnson C, Guiderdoni E, An G, Vijayraghavan U, Sundaresan V (2014) The polycomb group gene EMF2B is essential for maintenance of floral meristem determinacy in rice. Plant J 80(5):883–894. https://doi.org/10.1111/tpj.12688
Czermin B, Melfi R, McCabe D, Seitz V, Imhof A, Pirrotta V (2002) Drosophila enhancer of Zeste/ESC complexes have a histone H3 methyltransferase activity that marks chromosomal Polycomb sites. Cell 111(2):185–196
Derkacheva M, Hennig L (2014) Variations on a theme: Polycomb group proteins in plants. J Exp Bot 65(10):2769–2784. https://doi.org/10.1093/jxb/ert410
Derkacheva M, Steinbach Y, Wildhaber T, Mozgová I, Mahrez W, Nanni P, Bischof S, Gruissem W, Hennig L (2013) Arabidopsis MSI1 connects LHP1 to PRC2 complexes. EMBO J 32(14):2073–2085. https://doi.org/10.1038/emboj.2013.145
Francis NJ, Saurin AJ, Shao Z, Kingston RE (2001) Reconstitution of a functional core polycomb repressive complex. Mol Cell 8(3):545–556
Gendall AR, Levy YY, Wilson A, Dean C (2001) The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 107(4):525–535
Goodrich J, Puangsomlee P, Martin M, Long D, Meyerowitz EM, Coupland G (1997) A Polycomb-group gene regulates homeotic gene expression in Arabidopsis. Nature 386(6620):44–51. https://doi.org/10.1038/386044a0
Han G, Lu C, Guo J, Qiao Z, Sui N, Qiu N, Wang B (2020) C2H2 zinc finger proteins: master regulators of abiotic stress responses in plants. Front Plant Sci 11:115. https://doi.org/10.3389/fpls.2020.00115
Hennig L, Bouveret R, Gruissem W (2005) MSI1-like proteins: an escort service for chromatin assembly and remodeling complexes. Trends Cell Biol 15(6):295–302. https://doi.org/10.1016/j.tcb.2005.04.004
Huang Y, Jiang L, Liu B-Y, Tan C-F, Chen D-H, Shen W-H, Ruan Y (2019) Evolution and conservation of polycomb repressive complex 1 core components and putative associated factors in the green lineage. BMC Genomics 20:533. https://doi.org/10.1186/s12864-019-5905-9
Jung YJ, Lee IH, Nou IS, Lee KD, Rashotte AM, Kang KK (2013) BrRZFP1 a Brassica rapa C3HC4-type RING zinc finger protein involved in cold, salt and dehydration stress. Plant Biol (Stuttg) 15(2):274–283. https://doi.org/10.1111/j.1438-8677.2012.00631.x
Katz A, Oliva M, Mosquna A, Hakim O, Ohad N (2004) FIE and CURLY LEAF polycomb proteins interact in the regulation of homeobox gene expression during sporophyte development. Plant J 37(5):707–719
Kim J-M, Sasaki T, Ueda M, Sako K, Seki M (2015) Chromatin changes in response to drought, salinity, heat, and cold stresses in plants. Front Plant Sci 6:114. https://doi.org/10.3389/fpls.2015.00114
Kiyosue T, Ohad N, Yadegari R, Hannon M, Dinneny J, Wells D, Katz A, Margossian L, Harada JJ, Goldberg RB, Fischer RL (1999) Control of fertilization-independent endosperm development by the MEDEA polycomb gene in Arabidopsis. Proc Natl Acad Sci U S A 96(7):4186–4191
Köhler C, Hennig L, Bouveret R, Gheyselinck J, Grossniklaus U, Gruissem W (2003) Arabidopsis MSI1 is a component of the MEA/FIE Polycomb group complex and required for seed development. EMBO J 22(18):4804–4814. https://doi.org/10.1093/emboj/cdg444
Kumar D, Das PK, Sarmah BK (2018) Reference gene validation for normalization of RT-qPCR assay associated with germination and survival of rice under hypoxic condition. J Appl Genet 59(4):419–430. https://doi.org/10.1007/s13353-018-0466-1
Lafos M, Kroll P, Hohenstatt ML, Thorpe FL, Clarenz O, Schubert D (2011) Dynamic regulation of H3K27 trimethylation during Arabidopsis differentiation. PLoS Genet 7(4):e1002040. https://doi.org/10.1371/journal.pgen.1002040
Lewis EB (1978) A gene complex controlling segmentation in Drosophila. Nature 276(5688):565–570
Li W, Wang Z, Li J, Yang H, Cui S, Wang X, Ma L (2011) Overexpression of AtBMI1C, a polycomb group protein gene, accelerates flowering in Arabidopsis. PLoS ONE 6(6):e21364. https://doi.org/10.1371/journal.pone.0021364
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25(4):402–408. https://doi.org/10.1006/meth.2001.1262
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Luo M, Bilodeau P, Dennis ES, Peacock WJ, Chaudhury A (2000) Expression and parent-of-origin effects for FIS2, MEA, and FIE in the endosperm and embryo of developing Arabidopsis seeds. Proc Natl Acad Sci 97(19):10637–10642. https://doi.org/10.1073/pnas.170292997
Luo M, Bilodeau P, Koltunow A, Dennis ES, Peacock WJ, Chaudhury AM (1999) Genes controlling fertilization-independent seed development in Arabidopsis thaliana. Proc Natl Acad Sci U S A 96(1):296–301
Luo M, Liu X, Singh P, Cui Y, Zimmerli L, Wu K (2012) Chromatin modifications and remodeling in plant abiotic stress responses. Biochim Biophys Acta 1819(2):129–136. https://doi.org/10.1016/j.bbagrm.2011.06.008
Luo M, Platten D, Chaudhury A, Peacock WJ, Dennis ES (2009) Expression, imprinting, and evolution of rice homologs of the polycomb group genes. Mol Plant 2(4):711–723. https://doi.org/10.1093/mp/ssp036
Mohd-Sarip A, Venturini F, Chalkley GE, Verrijzer CP (2002) Pleiohomeotic can link polycomb to DNA and mediate transcriptional repression. Mol Cell Biol 22(21):7473–7483. https://doi.org/10.1128/MCB.22.21.7473-7483.2002
Mozgova I, Hennig L (2015) The polycomb group protein regulatory network. Annu Rev Plant Biol 66:269–296. https://doi.org/10.1146/annurev-arplant-043014-115627
Müller J, Hart CM, Francis NJ, Vargas ML, Sengupta A, Wild B, Miller EL, O’Connor MB, Kingston RE, Simon JA (2002) Histone methyltransferase activity of a Drosophila Polycomb group repressor complex. Cell 111(2):197–208
Nagar P, Kumar A, Jain M, Kumari S, Mustafiz A (2020) Genome-wide analysis and transcript profiling of PSKR gene family members in Oryza sativa. PLoS ONE 15(7):e0236349. https://doi.org/10.1371/journal.pone.0236349
Nallamilli BRR, Zhang J, Mujahid H, Malone BM, Bridges SM, Peng Z (2013) Polycomb group gene OsFIE2 regulates rice (Oryza sativa) seed development and grain filling via a mechanism distinct from Arabidopsis. PLoS Genet 9(3):e1003322. https://doi.org/10.1371/journal.pgen.1003322
Ojolo SP, Cao S, Priyadarshani SVGN, Li W, Yan M, Aslam M, Zhao H, Qin Y (2018) Regulation of plant growth and development: a review from a chromatin remodeling perspective. Front Plant Sci 9:1232. https://doi.org/10.3389/fpls.2018.01232
Qin F, Sakuma Y, Tran L-SP, Maruyama K, Kidokoro S, Fujita Y, Fujita M, Umezawa T, Sawano Y, Miyazono K-I, Tanokura M, Shinozaki K, Yamaguchi-Shinozaki K (2008) Arabidopsis DREB2A-interacting proteins function as RING E3 ligases and negatively regulate plant drought stress-responsive gene expression. Plant Cell 20(6):1693–1707. https://doi.org/10.1105/tpc.107.057380
Roy S (2016) Function of MYB domain transcription factors in abiotic stress and epigenetic control of stress response in plant genome. Plant Signal Behav 11(1):e1117723. https://doi.org/10.1080/15592324.2015.1117723
Saeed AI, Sharov V, White J, Li J, Liang W, Bhagabati N, Braisted J, Klapa M, Currier T, Thiagarajan M, Sturn A, Snuffin M, Rezantsev A, Popov D, Ryltsov A, Kostukovich E, Borisovsky I, Liu Z, Vinsavich A, … Quackenbush J (2003) TM4: a free, open-source system for microarray data management and analysis. BioTechniques 34(2):374–378. https://doi.org/10.2144/03342mt01
Sanchez-Pulido L, Devos D, Sung ZR, Calonje M (2008) RAWUL: a new ubiquitin-like domain in PRC1 ring finger proteins that unveils putative plant and worm PRC1 orthologs. BMC Genomics 9:308. https://doi.org/10.1186/1471-2164-9-308
Santos AP, Serra T, Figueiredo DD, Barros P, Lourenço T, Chander S, Oliveira MM, Saibo NJM (2011) Transcription regulation of abiotic stress responses in rice: a combined action of transcription factors and epigenetic mechanisms. OMICS 15(12):839–857. https://doi.org/10.1089/omi.2011.0095
Schatlowski N, Stahl Y, Hohenstatt ML, Goodrich J, Schubert D (2010) The CURLY LEAF interacting protein BLISTER controls expression of polycomb-group target genes and cellular differentiation of Arabidopsis thaliana. Plant Cell 22(7):2291–2305. https://doi.org/10.1105/tpc.109.073403
Shen Q, Lin Y, Li Y, Wang G (2021) Dynamics of H3K27me3 modification on plant adaptation to environmental cues. Plants 10(6):1165. https://doi.org/10.3390/plants10061165
Simon JA, Kingston RE (2013) Occupying chromatin: Polycomb mechanisms for getting to genomic targets, stopping transcriptional traffic, and staying put. Mol Cell 49(5):808–824. https://doi.org/10.1016/j.molcel.2013.02.013
Strejčková B, Čegan R, Pecinka A, Milec Z, Šafář J (2020) Identification of polycomb repressive complex 1 and 2 core components in hexaploid bread wheat. BMC Plant Biol 20(Suppl 1):175. https://doi.org/10.1186/s12870-020-02384-6
Tavares L, Dimitrova E, Oxley D, Webster J, Poot R, Demmers J, Bezstarosti K, Taylor S, Ura H, Koide H, Wutz A, Vidal M, Elderkin S, Brockdorff N (2012) RYBP-PRC1 complexes mediate H2A ubiquitylation at polycomb target sites independently of PRC2 and H3K27me3. Cell 148(4):664–678. https://doi.org/10.1016/j.cell.2011.12.029
Turck F, Roudier F, Farrona S, Martin-Magniette M-L, Guillaume E, Buisine N, Gagnot S, Martienssen RA, Coupland G, Colot V (2007) Arabidopsis TFL2/LHP1 specifically associates with genes marked by trimethylation of histone H3 lysine 27. PLoS Genet 3(6):e86. https://doi.org/10.1371/journal.pgen.0030086
Tuteja N (2007) Abscisic acid and abiotic stress signaling. Plant Signal Behav 2(3):135–138. https://doi.org/10.4161/psb.2.3.4156
Wu JI, Lessard J, Crabtree GR (2009) Understanding the words of chromatin regulation. Cell 136(2):200–206. https://doi.org/10.1016/j.cell.2009.01.009
Xu L, Shen W-H (2008) Polycomb silencing of KNOX genes confines shoot stem cell niches in Arabidopsis. Curr Biol 18(24):1966–1971. https://doi.org/10.1016/j.cub.2008.11.019
Yang CH, Chen LJ, Sung ZR (1995) Genetic regulation of shoot development in Arabidopsis: role of the EMF genes. Dev Biol 169(2):421–435. https://doi.org/10.1006/dbio.1995.1158
Yoshida S, Forno DA, Cock JH (1971) Laboratory manual for physiological studies of rice. Laboratory Manual for Physiological Studies of Rice. https://www.cabdirect.org/cabdirect/abstract/19721703488. Accessed 25 Nov 2019
Yuan L, Liu X, Luo M, Yang S, Wu K (2013) Involvement of histone modifications in plant abiotic stress responses. J Integr Plant Biol 55(10):892–901. https://doi.org/10.1111/jipb.12060
Zhang X, Germann S, Blus BJ, Khorasanizadeh S, Gaudin V, Jacobsen SE (2007) The Arabidopsis LHP1 protein colocalizes with histone H3 Lys27 trimethylation. Nat Struct Mol Biol 14(9):869–871. https://doi.org/10.1038/nsmb1283
Zhao Y, Zhang J, Sun Z, Tang Y, Wu Y (2021) Genome-wide identification and analysis of the Polycomb group family in Medicago truncatula. Int J Mol Sci 22(14):7537. https://doi.org/10.3390/ijms22147537
Zhu Y, Hu X, Duan Y, Li S, Wang Y, Rehman AU, He J, Zhang J, Hua D, Yang L, Wang L, Chen Z, Li C, Wang B, Song C-P, Sun Q, Yang S, Gong Z (2020) The Arabidopsis Nodulin Homeobox Factor AtNDX interacts with AtRING1A/B and negatively regulates abscisic acid signaling[OPEN]. Plant Cell 32(3):703–721. https://doi.org/10.1105/tpc.19.00604
Funding
The study was supported by funding from South Asian University, New Delhi, to AM for consumables and the infrastructure. NY received financial assistance from the Council of Scientific and Industrial Research.
Author information
Authors and Affiliations
Contributions
The idea, concept, and design of experiments, analysis, interpretation of data, fund, and project management were done by AM. NY has done the bioinformatics analyses and wet laboratory experiments. NY drafted the original manuscript. NY, AM, PN, and RR critically analyzed the data and revised the manuscript. AK helped in retrieving the RNA-seq data. AR helped in designing and initiating few experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 37 kb)
ESM 2
(XLSX 2706 kb)

ESM 3
Fig. S1 Figure showing the tertiary structure of all the PcG gene family members in Oryza sativa. (PNG 2974 kb)

ESM 4
Fig. S2 Figure showing the distribution of conserved motif of a PRC1 members and b PRC2 members in Arabidopsis thaliana. (PNG 1137 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yadav, N., Nagar, P., Rakhi, R. et al. Transcript profiling of Polycomb gene family in Oryza sativa indicates their abiotic stress-specific response. Funct Integr Genomics 22, 1211–1227 (2022). https://doi.org/10.1007/s10142-022-00906-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10142-022-00906-z