Abstract
Climate change and environmental issues compel us to find alternatives to the production of molecules of interest from petrochemistry. This study aims at understanding the production of butyrate, hydrogen, and CO2 from the oxidation of lactate with acetate in Clostridium tyrobutyricum and thus proposes an alternative carbon source to glucose. This specie is known to produce more butyrate than the other butyrate-producing clostridia species due to a lack of solvent genesis phase. The recent discoveries on flavin-based electron bifurcation and confurcation mechanism as a mode of energy conservation led us to suggest a new metabolic scheme for the formation of butyrate from lactate-acetate co-metabolism. While searching for genes encoding for EtfAB complexes and neighboring genes in the genome of C. tyrobutyricum, we identified a cluster of genes involved in butyrate formation and another cluster involved in lactate oxidation homologous to Acetobacterium woodii. A phylogenetic approach encompassing other butyrate-producing and/or lactate-oxidizing species based on EtfAB complexes confirmed these results. A metabolic scheme on the production of butyrate, hydrogen, and CO2 from the lactate-acetate co-metabolism in C. tyrobutyricum was constructed and then confirmed with data of steady-state continuous culture. This in silico metabolic carbon flux analysis model showed the coherence of the scheme from the carbon recovery, the cofactor ratio, and the ATP yield. This study improves our understanding of the lactate oxidation metabolic pathways and the role of acetate and intracellular redox balance, and paves the way for the production of molecules of interest as butyrate and hydrogen with C. tyrobutyricum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Many saccharolytic species of the genus Clostridium are known to produce CO2, butyric acid, and H2 as the main metabolic end products of butyric acid fermentation during the acidogenesis phase (Lee et al. 2008). Over the past decades, butyric acid fermentation has attracted great research and development (R and D) interest as it offers an alternative for the production of (i) butyric acid, a platform molecule widely used in food, cosmetic, and oil industries and mainly produced through petrochemical means, and (ii) hydrogen, a potential environmentally friendly energy carrier for substitution of fossil fuels and mainly produced from water electrolysis (Zeng and Zhang 2010; Cai et al. 2010; Dwidar et al. 2012; Bao et al. 2020).
C. tyrobutyricum produces higher amounts of butyric acid and hydrogen as the main fermentation products from carbohydrates than other species like Clostridium acetobutylicum or Clostridium butyricum (Cummins and Johnson 1971; Jang et al. 2013, 2014). As a consequence, this bacteria has been extensively explored by metabolic engineering and in fermentation for its potential to produce butyric acid and hydrogen from sugars and lignocellulosic biomass hydrolysates containing glucose and xylose (Liu et al. 2006; Jo et al. 2010; Jiang et al. 2013; Du et al. 2015; Yu et al. 2015; Fu et al. 2017; Suo et al. 2018a, b, 2019; Li et al. 2020). Among species usable in an industry, C. tyrobutyricum seems to be the best candidate in particular due to its sole acidogenesis phase (Jaros et al. 2013). C. tyrobutyricum is also able to metabolize lactate to butyrate and hydrogen (Cheng et al. 2013).
Lactic acid production by fermentation has attracted a lot of interest as it offers an alternative to the environmental pollution caused by the petrochemical industry (Abdel-Rahman et al. 2013). Lactic acid has a wide range of applications such as in food, beverages, cosmetics, pharmaceuticals, and chemical industry, and hence its demand is continuously increasing (Jantasee et al. 2017). From an industrial perspective, the use of lactate could be an attractive alternative and cost-effective to the use of carbohydrate to produce a molecule of interest, especially since it has been shown that lactate co-metabolism with acetate could stimulate butyrate and bio-hydrogen productions (Matsumoto and Nishimura 2007). But the mechanism of lactate oxidation in C. tyrobutyricum has not yet been identified and the functioning of the metabolic pathway has not been clarified. Moreover, studies on lactate oxidation metabolism could bring a solution to the late blowing defect in the cheese industry caused by contamination by C. tyrobutyricum (Klijn et al. 1995; D’incecco et al. 2018).
The first suggestion of lactate oxidation mechanism was proposed in C. acetobutylicum. Lactate oxidation to pyruvate would be catalyzed with iNAD+ lactate dehydrogenase, but the genes involved have not yet been identified (Cheng et al. 2013). The discovery of flavin-based electron bifurcation (FBEB) complexes has improved our understanding of energy conservation mechanisms in strict anaerobic bacteria and archaea (Thauer et al. 2008; Wang et al. 2010; Buckel and Thauer 2018a). This conservation of energy results in a net exergonic reaction with minimal negative free energy change involving an electron transfer between exergonic and endergonic reactions. FBEBs complexes are divided into four distinct families, including that of the electron transfer flavoprotein (Etfs) complex. This complex consists of two subunits encoded by the etfA and etfb genes. It was first described in Clostridium kluyveri (Bcd/EtffAB) as its genes are clustered with a butyryl-CoA dehydrogenase gene (Li et al. 2008). This key reaction in the production of butyrate involves the exergonic reduction of crotonyl-CoA to butyryl-CoA with NADH and the endergonic reduction of ferredoxins with NADH, the equation (Eq. 1) of which is as follows:
Genes encoding BCD/EtfAB are clustered with HBD ones (3-hydroxybutyryl-CoA dehydrogenase) also found in other Clostridium genomes (Bennett and Rudolph 1995; Lee et al. 2016). It is common to find genes coding for the Etfs complex in the same genetic locus as a CoA dehydrogenase gene in bacterial genomes. This is the case for A. woodii with a lactate dehydrogenase (LDH) gene. This LDH/EtfAB complex explains the biochemical reaction of lactate oxidation (Weghoff et al. 2015). It involves the exergonic reaction of the ferredoxin oxidation reduced with NAD+ coupled with the endergonic reaction of lactate reduction to pyruvate with NAD+ giving the following equation (Eq. 2):
This FBEB complex leads to decrease in the ΔG°' of the oxidation of lactate from + 25 to + 6 kJ mol−1 making the reaction energetically more favorable. Genes encoding the LDH/EtfAB complex are grouped with those encoding a lactate permease, a lactate racemase, and a transcriptional regulator forming a lactate oxidation operon. Orthologous operons have been identified in other Firmicutes species, some of which belong to the Clostridium genus such as Clostridium botulinum and Clostridiuml jundhalii. From phylogenetic and structural analyses of different EtfA and EtfB subunits belonging to the domain of bacteria and archaea, five Etf groups (G1–G5) have been distinguished. The G2 group is mostly represented by the Firmicutes and divided into two sub-groups. The G2A subgroup includes the Etfs involved in butyrate synthesis, and the G2B subgroup comprises those in lactate oxidation (Garcia Costas et al. 2017).
The aim of this study was to investigate lactate-acetate co-metabolism in C. tyrobutyricum using the LDH/EtfAB complex. Because C. tyrobutyricum is phylogenetically close to C. ljungdhalli, a comparison with A. woodii and C. kluyveri could provide answers to the metabolic mechanisms of lactate oxidation involved. The investigation was made in three complementary steps: (i) in silico analysis of putative lactate oxidation clusters, (ii) phylogenetic analysis, and (iii) metabolic flux analysis (MFA) including intracellular redox balance and carbon recovery, with data gathered from a steady-state continuous culture of C. tyrobutyricum using lactate and acetate as carbon substrates.
Methods
Identification of a predicted EtfAB and neighboring genes involved in lactate oxidation from Clostridium tyrobutyricum CIRM BIA 2237
Protein sequence research studies from predicted Etf genes were performed on C. tyrobutyricum CIRM BIA 2237 (NCBI RefSeq: NZ_CP038158.1). The etfA (locus_tag: AWO_RS04415) and etfB (AWO_RS0441) genes of A. woodii DSM 1030 (NC_016894) (Weghoff et al. 2015) were used as queries. Searches, data analyses, and visualizations were performed with NCBI blast. The etf sequences from C. ljungdahlii DSM13528 were also used as queries.
Phylogenetic analysis of Etf complex proteins from the selected bacterial species
EtfA and EtfB protein sequences were searched in the genome sequence of the following species: A. woodii DSM1030 (RefSeq: NC_016894), C. kluyveri DSM555 (RefSeq: NC_009706.1), Clostridium acetobutylicum ATCC 824 (RefSeq: NC_003030.1), Clostridium butyricum KNU-L09 (RefSeq: NZ_CP013252.1), Clostridium beijerenckii DSM791 (RefSeq: NZ_CP073653.1), C.tyrobutyricum CIRM BIA 2237 (RefSeq: NZ_CP038158.1), and Eubacterium limosum (RefSeq: NZ_CP019962.1).
A custom EtfA and EtfB protein database of the above listed species was prepared using the NCBI database by searching all electron transfer flavoprotein A or B genes from the genome sequences.
The selected EtfA and EtfB protein sequences forming an Etf complex in each genome were compilated by addition of both sequences giving Etf complex protein sequences.
Etf complexes were subjected to phylogenetic analysis, and protein sets were aligned using ClustalW with default parameters (cost matrix: ID, gap open cost: 8, gap extend cost: 0.1). A phylogenetic tree was built using Tree Builder with the following parameters: genetic distance model: Jukes-Cantor (Jukes and Cantor 1969), tree build method: UPGMA (Michener and Sokal 1957), resampling method: bootstrap, number of replicates: 1000, support threshold: 30% by using the Geneious v2022.0.2 software.
Bacteria and medium
C. tyrobutyricum CIRM BIA 2237 was obtained from National Institute of Agronomic Research and Environment collection. The strain was stored at – 80 °C in cryotubes. For pre-culture, 1 colony of C. tyrobutyricum on RCM (reinforced clostridial medium; Biokar, France) Petri dishes was used to inoculate 100 mL of an RCM-modified (glucose was replaced by sodium lactate) medium at 37 °C until obtaining OD600nm: 1.4–1.8 measured by a spectrometer (PRIM; Secomam, France). A medium with 14 g.L−1 sodium lactate (Sigma-Aldrich, France), yeast extract 3 g.L−1 (VWR, France), and 10 g.L−1 meat extract (VWR, France), 3 g.L−1 sodium acetate (Rectapur, France), 5 g.L−1 NaCl (Chem-LAB, Belgium), 0.05 g.L−1 cysteine hydrochloride (Sigma-Aldrich, France) was used in the pre-culture and continuous culture studies. Media for the pre-culture were autoclaved at 120 °C for 15 min and filter-sterilized (0.2 µm) for continuous cultures (Millipore, Merck, France).
Continuous culture setup
Continuous culture was carried out in a 2-L bioreactor with 750 mL working volume (New Brunswick, Scientific, Discovery 100, USA). The bioreactor was autoclaved with distilled water at 120 °C for 15 min. After cooling, water was replaced by the filter-sterilized medium with lactate and acetate. The culture was controlled at a temperature of 37 °C, and a pH of 5.8. The stirring rate was controlled at 150 rpm, which was high enough to keep a homogenous medium without leading to significant foaming. Oxidoreduction potential (Eh) was measured with redox probe (Pt4805-DPAS-SC K8S; Mettler Toledo, France). The initial batch culture was started with 700 mL of a medium inoculated with 50 mL pre-culture, and switched to the continuous culture at the end of the growth phase. During fermentation, nitrogen was sparged into the headspace of the bioreactor at a flow rate of 10 mL.min−1 to maintain the anaerobic state of the bioreactor. The feed medium flow rate was controlled by a peristaltic pump (MINIPULS 3; Gilson, France). A total volume of 750 mL was maintained using a balance (Lynx; Mettler Toledo, France) coupled with a monitoring reactor unit (MRU) (New Brunswick, USA).
Analytical methods
Short-chain fatty acids were analyzed by HPLC with ion exclusion (HPLC system; Diomex (Thermo Fisher), Photodiode Array Detector Ultimate 3000 DAD, and a 300 × 7.8 mm Aminex HPX-87H column with a guard column). HPLC conditions for determination of organic acids were: column temperature of 30 °C; flow rate, 0.6 mL.min−1; injected volume, 20 µL; mobile phase, 9 mM sulfuric acid in water.
In silico model construction and metabolic flux
The anaerobic lactate oxidation metabolic network for the C. tyrobutyricum CIRM BIA 2237 strain was developed for MFA from the identification of EtfAB complex in C. tyrobutyricum CIRM BIA 2237 and phylogenetic analysis steps and completed with pathway information collected from other studies reported in the literature and the KEGG database (Lee et al. 2016; Kanehisa et al. 2017). The metabolic network of C. tyrobutyricum CIRM BIA 2237 was mathematically constructed with four key reactions (Table 1), gathering 11 reactions and seven intracellular metabolites from C. tyrobutyricum metabolism (Fig. 1). The lactate and acetate uptakes and the production of butyrate, which were measured during the steady-state phase of the continuous culture, were used as constraints in MFA. The steady-state phase was identified from the stability of the cell density determined by the OD600nm method with stable lactate and acetate consumption values and redox potential value (Eh). The model construction and the in silico analysis were carried out from the reactions in Table 1 using a previous method (Tsai and Lee 1988; Orth et al. 2010) and applied on https://smart-biotech.online/research/mfa/clostridium.tyrobutyricum.lactate.
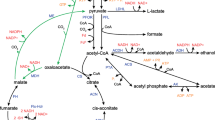
Proposed scheme for the metabolic pathways involving lactate and acetate conversion in C. tyrobutyricum. LctP, lactate permease; LDH, lactate dehydrogenase; PFOR, pyruvate ferredoxine:oxidoreductase; PTA, phosphotransacetylase; ACK, acetate kinase; THL, thiolase; HBD, 3-hydroxybutyryl-CoA dehydrogenase; CRT, crotonase; BCD, butyryl-CoA dehydrogenase; CAT, CoA-transferase; HYD, hydrogenase
From the metabolic steady state for intracellular metabolites, the metabolic fluxes (\(v\)) are constrained by the stoichiometry matrix (S) giving the following equation:
Metabolic carbon fluxes were calculated with measured rates (r) of lactate and acetate consumption, butyrate production, estimating CO2 production rate from lactate-acetate metabolic scheme (Fig. 1), and lactate consumption rate, giving the matrix system in Eq. (4), where xi are intracellular flow rates.
Considering the steady-state continuous culture, the intracellular flow rates of acetyl-CoA and butyryl-CoA are assumed as equal to 0.
An important application of MFA includes determining ATP yields and parameters of redox homeostasis (cofactor redox balance and metabolites redox balance, both being involved in cellular redox balance).
ATP yield (YATP, g.mol−1) is the ratio of biomass (g DW.L−1) produced from ATP (mol.L−1). For cofactor recovery, NADH ratio (Eq. 5) and Fd ratio (Eq. 6) were calculated separately and compiled together, giving Eq. 7. All these ratios were calculated on the basis of intracellular flux:
Redox balance was calculated from the oxidation state of substrates and products (lactate, 0; acetate, 0; carbon dioxide, + 2; hydrogen, − 1; and butyrate, − 2) (Riondet et al. 2000). The sum of the oxidation state of products should be equal to the sum of the oxidation state of substrates. Considering that the sum of the oxidation state of substrates equals 0, the following equation was used for redox balance (Eq. 8):
Dividing by rsubstrates allows to compare conditions with different substrates flow rates.
Results and discussion
Identification of genes involved in lactate oxidation
The discovery of complex FBEBs has led to a better understanding of the metabolic function of strict anaerobes and archaea (Buckel and Thauer 2018a), and to understand some metabolic activities like the ability to metabolize lactate with acetate or butyrate synthesis from some Clostridium species (Li et al. 2008; Detman et al. 2019).
We studied the genome of C. tyrobutyricum CIRM BIA 2237 to show the presence of EtfAB complexes. C. tyrobutyricum CIRM BIA 2237 has been isolated from silage and has already been used as a lactate oxidizer in different works (Roux and Bergere 1977). Its complete genome has been fully sequenced and annotated (Munier et al. 2019). Analysis of the chromosome (NZ_CP038158.1) of C. tyrobutyricum CIRMBIA 2237 identified three clusters of genes coding for EtfAB complexes (Fig. 2).
Chromosome fragments (NZ_CP038158.1) of C. tyrobutyricum CIRM BIA 2237 containing EtfAB and FAD-binding oxidoreductase genes. EtfB (1: EZN00_RS10860, 2: EZN00_RS10795, 3: EZN00_RS08615), EtfA (1: EZN00_RS10865, 2: EZN00_RS10800, 3: EZN00_RS08620), FAD-binding oxidoreductase (1:: EZN00_RS10870; 3: EZN00_RS08625)
In fragment 1, genes encoding the EtfAB complex are next to a FAD-binding oxidoreductase. The EtfAB complex involved in fragment 2 is in a gene cluster involving an enoyl-CoA hydratase (also identified as crotonase), an acyl-CoA dehydrogenase, a 3-hydroxybutyryl-CoA dehydrogenase, and a FAD/NAD-binding protein. The predicted acyl-CoA dehydrogenase protein presents 100% identity with the butyryl-CoA dehydrogenase of C. tyrobutyricum KCTC 5387 and can thus be identified as a butyryl-CoA dehydrogenase. Therefore, the cluster of genes identified in fragment 2 is most likely involved in the production of the butyryl-CoA characteristic of butyrate-producing Clostridium.
In fragment 3, the EtfAB complex is included between the coding genes for a transcriptional regulator, a L-lactate permease, a lactate racemase, and a FAD-binding oxidoreductase. This gene cluster was also found in C. ljungdahlii DSM 13528, a species phylogenetically close to C. tyrobutyricum (Collins et al. 1994; Weghoff et al. 2015).
The protein sequences found in the lactate oxidation cluster present at least 38% of identity and 95% of query cover between C. tyrobutyricum CIRM BIA 2237, C. ljungdahlii DSM 13528, and A. woodii DSM 1030 (Fig. 3).
Putative lactate oxidation cluster of C. tyrobutyricum CIRM BIA 2237. Its identity percentage is compared with those from C. ljungdahlii DSM 13,528 et A. woodii DSM 1030. TR, transcriptional regulator (EZN00_RS08600, CLJU_RS10615, AWO_RS0440); L-lactate permease (EZN00_RS08605, CLJU_RS10610, and AWO_RS04425); lactate racemase (EZN00_RS08610, CLJU_RS10605, and AWO_RS04430); EtfB (EZN00_RS08615, CLJU_RS10600, and AWO_RS04410); EtfA (EZN00_RS08620, CLJU_RS10595, and AWO_RS04415); LDH, lactate dehydrogenase (EZN00_RS08625, CLJU_RS10590, and AWO_RS04420)
According to these results, C.tyrobutyricum would have the same mechanism of lactate oxidation as A. woodii, i.e., involving an EtfAB complex.
In order to support our proposal, a phylogenetic comparison was made based on EtfAB protein sequences of different species that can oxidize lactate and produce butyrate (E. limosum ATCC 8486, C. butyricum KNU-L09, C. beijerenckii DSM 791, and C. acetobutylicum ATCC 824), a reference species for the oxidation of lactate (A. woodii), and a reference species for the production of butyrate (C. kluyveri). This phylogenetic analysis was restricted to bacteria able to uptake lactate in the presence of acetate to produce butyrate. A search for EtfAB complexes was previously carried out by BLASTp in these different selected species. In the genomes of the selected species, at least two EtfAB complexes have been identified.
As shown in Fig. 4, the EtfAB complexes encoded by the etfA and etfB genes associated with the hbd form a distinct group with 100% bootstrap support.
Phylogenetic analysis of Etf complex protein sequences between different species. Orange, EtfAB coupled with acyl-CoA DH; red, EtfAB coupled with Hbd; green, EtfAB coupled FAD-binding oxidoreductase/LDH and lactate permease; blue, EtfAB-coupled FAD-binding oxidoreductase; black, out of any phylogenetic group
As already observed in other studies, the latter is related to EtfAB complexes associated with an acyl-CoA dehydrogenase gene (Detman et al. 2019). Concerning EtfAB complexes associated with a gene coding for GlcD or GlcD/LctP, two distinct additional groups are observable. The first forms a cluster involving the EtfAB complex associated with GclD/LctP in A. woodii and C. tyrobutyricum with 100% bootstrap support, and the second groups together with the EtfABGlcD_LctP complexes of C. butyricum KNU-L09 and C. beijerenckii DSM 791 as well as that associated with GlcD in C. tyrobutyricum CIRM BIA 2237.
Our results are in agreement with those of Garcia Costas et al. (2017) and Detman et al. (2019). The two G2 subgroups of EtfAB are found in this phylogenetics analysis. EtfAB protein sequences encoded by etfA and etfB associated with hbd or acyl-CoA DH belonging to subgroup G2A are involved in butyrate production. Those with the gene encoding for GlcD and/or LldP_GlcD including A. woodii formed subgroup G2B involved in lactate oxidation. The results of this phylogenetic analysis correspond to the ones obtained by comparisons of protein sequences with A. woodii and C. ljungdahlii.
Metabolic scheme with enzyme machinery of lactate and acetate transformation to butyrate
An update of the metabolic scheme involving the oxidation of lactate in C. tyrobutyricum could be proposed (Fig. 1). This scheme was built from previous studies on butyric acid fermentation from C. tyrobutyricum (Lee et al. 2016; Kanehisa et al. 2017), the database KEGG, and knowledge of flavin-based electron confurcation and regulation of energy conversion mechanisms (Buckel and Thauer 2018a).
We constructed a model of central carbon metabolism in C. tyrobutyricum involving the conversion of lactate to pyruvate via a stable cytoplasmic complex formed of LDH with electron-transferring flavoprotein (EtfAB) using flavin-based electron confurcation. Unlike in glucose where two ATPs are produced during glycolysis, this scheme with the LDH/EtfAB complex proposes the acetate branch as the only metabolic pathway capable of producing ATP with the F0F1 ATPase and covering the cell’s energy needs. Moreover, the reaction of the electron confurcation involving oxidation of reduced ferredoxins will change the intracellular redox balance.
Electron confurcation is the reverse reaction of bifurcation coupling an exergonic reduction of NAD+ with reduced ferredoxin with an endergonic reduction of NAD+ and lactate in our case. As explained in Buckel and Thauer (2018b), two different electron donors, one from reduced ferredoxin (low potential donor) and one from lactate (high potential donor), act together to reduce the bifurcating NAD+.
In silico model construction and verification
To support the results obtained from the bioinformatics analyses, a MFA model was built. The metabolic flux network of C. tyrobutyricum CIRM BIA 2237 was constructed from the scheme in Fig. 1 without including Rnf complex and NfnAB because a previous work has shown that the genes coding for these complexes are poorly expressed during butyric fermentation (Lee et al. 2016). Therefore, a first MFA with only Etfs complexes and hydrogenases was carried out. Lactate and acetate utilization and production of butyrate, three of the main substrates and products of fermentation, were followed during the continuous culture (Table 2). These continuous data were used as constraints in the stochiometric matrix (Eq. 4). Batch data from the literature were also used as constraints in the MFA model as second verification (Bryant and Burkey 1956). Carbon recovery and butyrate/acetate flux ratio were used as criteria to evaluate the agreement between stochiometric matrix and experiments.
As can be seen in Table 2, data from fermentations with two different strains and culture processes lead to similar acetate/lactate, butyrate/lactate, and CO2/lactate ratios and consequently a butyrate/acetate ratio around 2, and a carbon recovery around 100%. Consequently, the following general equation of lactate-acetate co-metabolism in C. tyrobutyricum could be proposed:
The MFA model allows to quantify intracellular metabolic fluxes (all expressed in % mM–C) (Fig. 5); starting from 83 mM–C of lactate and 17 mM–C of acetate external for a total of 100 mM–C, carbon flux distribution from lactate is majorly flowed until acetyl-CoA (55 mM–C) node with a production of CO2 (28 mM–C). Flux from acetyl-CoA node was bifurcated into acetic acid formation (19 mM–C) and butyric acid synthesis (72 mM–C) via butyryl-CoA node. The assimilation of 17 mM–C of external acetate with the 19 mM–C of acetate formed justifies the 72 mM–C of the butyrate branch.
Scheme of the metabolic flux distribution involving lactate and acetate conversion by C. tyrobutyricum. Using an MFA model with 100 mM–C of substrates (83 mM–C from lactate and 17 mM–C from acetate). All values are expressed in % of total carbon. Hatched arrows represent multiple metabolic reactions
Thus, butyrate production is possible by addition of external acetate carbons and re-assimilation of acetate carbons formed in metabolic pathways. Moreover, we could observe that the consumed external acetate is essential in intra-cellular homeostasis. Indeed, by removing it from MFA, we found that the NADH ratio had fallen from 98 to 87%. This result was also observed in Staphylococcus aureus metabolism (Marshall et al. 2016).
Figure 5 is in agreement with Eq. 9; the sum of carbon fluxes giving acetyl-CoA at the origin of butyrate (72 mM–C) production respected a ratio of 3 coming from lactate (55 mM–C) and 1 from acetate (17 mM–C). Two acetyl-CoA are required for the production of one butyryl-CoA. To confirm the consistency of these observations with a necessary recovery of both CoA–SH and NADH, a statement from this stoichiometric balance was performed from intracellular metabolic rates calculated by MFA (Table 3), and the following routes were considered according to Fig. 1:
-
For NAD+/NADH ratio (Eqs. 5 and 7): LDH/EtfAB, HbBD, and BCD/EtfAB routes
-
For Fdox/Fdred ratio (Eqs. 6 and 7): LDH/EtfAB, PFOR, BCD/EtfAB, and HYD routes
A NADH ratio of 98% is obtained. However, without the HYD route, a 61% recovery could be calculated for ferredoxin ratio and causing an excess of 2.34 mM.h−1 of reduced ferredoxin. This result can be explained due to the absence of the hydrogen-produced value from the hydrogenase during fermentation (Fig. 1).
By respecting the reaction stoichiometry involving hydrogenase, 1.14 mM.h−1 of hydrogen would be generated considering the results obtained with the calculation of intracellular fluxes (Table 3). In order to confirm this result, a second approach was carried out using oxidation state balance (Eq. 8). Considering intracellular fluxes, 0.03 of oxidation state balance was obtained instead of 0.53 without hydrogen value. The first value is closer to neutrality, meaning that hydrogen production should be necessary to maintain redox homeostasis. Moreover, the HYD route improved the ferredoxin ratio to 100% and cofactor recovery to 99% (Table 3). These results are close to full balance; it suggests a low expression of genes coding for the Rnf and NfnAB complexes in consumption of lactate and acetate. Furthermore, with the agreement of these MFA results with consistency between the flow distribution and intracellular redox ratio close to 100% of recovery, the involvement of LDH/EtfAB in the lactate oxidation mechanism from C. tyrobutyricum would be metabolically possible.
Concerning ATP synthesis, an ATP yield at 0.35 mol.mol−1 of substrates was calculated. This result is similar to those obtained with propionate fermentation from Clostridium homopropionicum and Veillonella parvula from lactate uptake (Seeliger et al. 2002). Besides, an 8.8 g DW. mol−1 ATP yield is in agreement with the 9 g DW. mol−1 ATP estimated from C. kluyveri with ethanol and acetate as substrates and similar to or higher than C. butyricum (10.1 gDW. mol−1 ATP) and E. coli (7.9 g DW. mol−1 ATP) with glucose as substrate, further confirming our suggestion (Thauer et al. 1968; Crabbendam et al. 1985; Riondet et al. 2000). Thus, despite the constraints of continuous data and stoichiometry of metabolic pathways, we obtained a consistency between flow distribution, an intracellular redox recovery close to 100%, and a positive ATP generation value close to lactate oxidation in other bacteria. These results support the consistency of this new scheme proposed with LDH/EtfAB.
Conclusion
Bioinformatics allowed us to identify a gene cluster coding for the LDH/EtfAB complex that could be involved in lactate oxidation metabolism from C. tyrobutyricum. The metabolic flux analysis involving the LDH/EtfAB complex has confirmed our assumptions about the lactate oxidation mechanism in C. tyrobutyricum. In addition, the lactate/acetate ratio of 3:1 provides an explanation of how acetate is used as a co-substrate for the production of two butyrate molecules. The understanding of this metabolism could allow for the development of new strategies against this bacterium known to be involved in late blowing defect of cheese, or even exploit lactate and acetate as new resources to produce hydrogen or butyrate as molecules of interest.
Data availability
All the data generated or analyzed during this study are included in this published article. The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request and on the following website: https://smart-biotech.online/research/mfa/clostridium.tyrobutyricum.lactate
Abbreviations
- Etf :
-
Electron transfer flavoprotein
- FAD :
-
Flavin adenine dinucleotide
- GlcD :
-
Domain of FAD-dependent lactate dehydrogenase
- FAD/FMN :
-
containing dehydrogenase
- HPLC :
-
High-performance liquid chromatography
- LDH :
-
Lactate dehydrogenase
- LldP :
-
Lactate permease
- OD :
-
optical density
- R&D :
-
Research and development
- iNAD + :
-
Lactate dehydrogenase: independent NAD+ lactate dehydrogenase
- FBEB :
-
Flavin-based electron bifurcation
- Etf :
-
Electron transfer flavoprotein
- Bcd :
-
Butyryl-CoA dehydrogenase
- HBD :
-
3-Hydroxybutyryl-CoA dehydrogenase
- MFA :
-
Metabolic flux analysis
- RCM :
-
Reinforced clostridial medium
- KEGG :
-
Kyoto Encyclopedia of Genes and Genomes
- DH :
-
Dehydrogenase
- THL :
-
Thiolase
- PTA :
-
Phophotransacetylase
- CRT :
-
Crotonase
- ACK :
-
Acetate kinase
- PFOR :
-
Pyruvate ferredoxin:oxidoreductase
- Rnf :
-
Ferredoxin:NAD reductase
- NfnAB :
-
NADH-dependent ferredoxin:NADP reductase
- Hyd :
-
Hydrogenase
- DW :
-
Dry weight
References
Abdel-Rahman MA, Tashiro Y, Sonomoto K (2013) Recent advances in lactic acid production by microbial fermentation processes. Biotechnol Adv 31:877–902. https://doi.org/10.1016/j.biotechadv.2013.04.002
Bao T, Feng J, Jiang W et al (2020) Recent advances in n-butanol and butyrate production using engineered Clostridium tyrobutyricum. World J Microbiol Biotechnol 36:138. https://doi.org/10.1007/s11274-020-02914-2
Bennett GN, Rudolph FB (1995) The central metabolic pathway from acetyl-CoA to butyryl-CoA in Clostridium acetobutylicum. FEMS Microbiol Rev 17:241–249. https://doi.org/10.1111/j.1574-6976.1995.tb00208.x
Bryant MP, Burkey LA (1956) The characteristics of lactate-fermenting spore forming anaerobes from silage. J Bacteriol 71:43
Buckel W, Thauer RK (2018a) Flavin-based electron bifurcation, ferredoxin, flavodoxin, and anaerobic respiration with protons (Ech) or NAD+ (Rnf) as electron acceptors: a historical review. Front Microbiol 9:401. https://doi.org/10.3389/fmicb.2018.00401
Buckel W, Thauer RK (2018b) Flavin-based electron bifurcation, a new mechanism of biological energy coupling. Chem Rev 118:3862–3886. https://doi.org/10.1021/acs.chemrev.7b00707
Cai G, Jin B, Saint C, Monis P (2010) Metabolic flux analysis of hydrogen production network by Clostridium butyricum W5: effect of pH and glucose concentrations. Int J Hydrog Energy 35:6681–6690. https://doi.org/10.1016/j.ijhydene.2010.04.097
Cheng H-H, Whang L-M, Lin C-A et al (2013) Metabolic flux network analysis of fermentative hydrogen production: using Clostridium tyrobutyricum as an example. Bioresour Technol 141:233–239. https://doi.org/10.1016/j.biortech.2013.03.141
Collins MD, Lawson PA, Willems A et al (1994) The phylogeny of the genus Clostridium: proposal of five new genera and eleven new species combinations. Int J Syst Bacteriol 44:812–826. https://doi.org/10.1099/00207713-44-4-812
Crabbendam PM, Neijssel OM, Tempest DW (1985) Metabolic and energetic aspects of the growth of Clostridium butyricum on glucose in chemostat culture. Arch Microbiol 142:375–382. https://doi.org/10.1007/BF00491907
Cummins CS, Johnson JL (1971) Taxonomy of the Clostridia: wall composition and DNA homologies in Clostridium butyricum and other butyric acid-producing Clostridia. J Gen Microbiol 67:33–46. https://doi.org/10.1099/00221287-67-1-33
D’Incecco P, Pellegrino L, Hogenboom JA et al (2018) The late blowing defect of hard cheeses: behaviour of cells and spores of Clostridium tyrobutyricum throughout the cheese manufacturing and ripening. LWT - Food Sci Technol 87:134–141. https://doi.org/10.1016/j.lwt.2017.08.083
Detman A, Mielecki D, Chojnacka A et al (2019) Cell factories converting lactate and acetate to butyrate: Clostridium butyricum and microbial communities from dark fermentation bioreactors. Microb Cell Factories 18:36. https://doi.org/10.1186/s12934-019-1085-1
Du Y, Jiang W, Yu M et al (2015) Metabolic process engineering of Clostridium tyrobutyricum Δack-adhE2 for enhanced n-butanol production from glucose: effects of methyl viologen on NADH availability, flux distribution, and fermentation kinetics: metabolic process engineering of Clostridium tyrobutyricum. Biotechnol Bioeng 112:705–715. https://doi.org/10.1002/bit.25489
Dwidar M, Park J-Y, Mitchell RJ, Sang B-I (2012) The future of butyric acid in industry. Sci World J 2012:1–10. https://doi.org/10.1100/2012/471417
Fu H, Yu L, Lin M et al (2017) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production from glucose and xylose. Metab Eng 40:50–58. https://doi.org/10.1016/j.ymben.2016.12.014
Garcia Costas AM, Poudel S, Miller A-F et al (2017) Defining electron bifurcation in the electron-transferring flavoprotein family. J Bacteriol 199:e00440-e517. https://doi.org/10.1128/JB.00440-17
Jang Y-S, Woo HM, Im JA et al (2013) Metabolic engineering of Clostridium acetobutylicum for enhanced production of butyric acid. Appl Microbiol Biotechnol 97:9355–9363. https://doi.org/10.1007/s00253-013-5161-x
Jang Y-S, Im JA, Choi SY et al (2014) Metabolic engineering of Clostridium acetobutylicum for butyric acid production with high butyric acid selectivity. Metab Eng 23:165–174. https://doi.org/10.1016/j.ymben.2014.03.004
Jantasee S, Kienberger M, Mungma N, Siebenhofer M (2017) Potential and assessment of lactic acid production and isolation-a review: potential and assessment of lactic acid production. J Chem Technol Biotechnol 92:2885–2893. https://doi.org/10.1002/jctb.5237
Jaros AM, Rova U, Berglund KA (2013) Acetate adaptation of clostridia tyrobutyricum for improved fermentation production of butyrate. Springerplus 2:47. https://doi.org/10.1186/2193-1801-2-47
Jiang L, Song P, Zhu L et al (2013) Comparison of metabolic pathway for hydrogen production in wild-type and mutant Clostridium tyrobutyricum strain based on metabolic flux analysis. Int J Hydrog Energy 38:2176–2184. https://doi.org/10.1016/j.ijhydene.2012.11.050
Jo JH, Jeon CO, Lee SY et al (2010) Molecular characterization and homologous overexpression of [FeFe]-hydrogenase in Clostridium tyrobutyricum JM1. Int J Hydrog Energy 35:1065–1073. https://doi.org/10.1016/j.ijhydene.2009.11.102
Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Mammalian protein metabolism. Elsevier, pp 21–132
Kanehisa M, Furumichi M, Tanabe M et al (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res 45:D353–D361. https://doi.org/10.1093/nar/gkw1092
Klijn N, Nieuwenhof FF, Hoolwerf JD et al (1995) Identification of Clostridium tyrobutyricum as the causative agent of late blowing in cheese by species-specific PCR amplification. Appl Env Microbiol 61:2919–2924
Lee SY, Park JH, Jang SH et al (2008) Fermentative butanol production by Clostridia. Biotechnol Bioeng 101:209–228. https://doi.org/10.1002/bit.22003
Lee J, Jang Y-S, Han M-J et al (2016) Deciphering Clostridium tyrobutyricum metabolism based on the whole-genome sequence and proteome analyses. Mbio 7:e00743-e816. https://doi.org/10.1128/mBio.00743-16
Li F, Hinderberger J, Seedorf H et al (2008) Coupled ferredoxin and crotonyl coenzyme A (CoA) reduction with NADH catalyzed by the butyryl-CoA dehydrogenase/Etf complex from Clostridium kluyveri. J Bacteriol 190:843–850. https://doi.org/10.1128/JB.01417-07
Li W, Cheng C, Cao G et al (2020) Comparative transcriptome analysis of Clostridium tyrobutyricum expressing a heterologous uptake hydrogenase. Sci Total Environ 749:142022. https://doi.org/10.1016/j.scitotenv.2020.142022
Liu X, Zhu Y, Yang S-T (2006) Butyric acid and hydrogen production by Clostridium tyrobutyricum ATCC 25755 and mutants. Enzyme Microb Technol 38:521–528. https://doi.org/10.1016/j.enzmictec.2005.07.008
Marshall DD, Sadykov MR, Thomas VC et al (2016) Redox imbalance underlies the fitness defect associated with inactivation of the Pta-AckA pathway in Staphylococcus aureus. J Proteome Res 15:1205–1212. https://doi.org/10.1021/acs.jproteome.5b01089
Matsumoto M, Nishimura Y (2007) Hydrogen production by fermentation using acetic acid and lactic acid. J Biosci Bioeng 103:236–241. https://doi.org/10.1263/jbb.103.236
Michener CD, Sokal RR (1957) A quantitative approach to a problem in classification. Evolution 11:130–162. https://doi.org/10.1111/j.1558-5646.1957.tb02884.x
Munier E, Licandro-Séraut H, Achilleos C, et al (2019) Whole-genome sequencing and annotation of Clostridium tyrobutyricum strain CIRMBIA 2237, isolated from silage. Microbiol Resour Announc 8:e00492–19, /mra/8/30/MRA.00492–19.atom. https://doi.org/10.1128/MRA.00492-19
Orth JD, Thiele I, Palsson BØ (2010) What is flux balance analysis? Nat Biotechnol 28:245–248. https://doi.org/10.1038/nbt.1614
Riondet C, Cachon R, Wache Y et al (2000) Extracellular oxidoreduction potential modifies carbon and electron flow in Escherichia coli. J Bacteriol 182:620–626. https://doi.org/10.1128/JB.182.3.620-626.2000
Roux C, Bergere JL (1977) Taxonomic characters of Clostridium tyrobutyricum. Ann Microbiol Paris 128A:267–276
Seeliger S, Janssen PH, Schink B (2002) Energetics and kinetics of lactate fermentation to acetate and propionate via methylmalonyl-CoA or acrylyl-CoA. FEMS Microbiol Lett 211:65–70. https://doi.org/10.1111/j.1574-6968.2002.tb11204.x
Suo Y, Fu H, Ren M et al (2018a) Enhanced butyric acid production in Clostridium tyrobutyricum by overexpression of rate-limiting enzymes in the Embden-Meyerhof-Parnas pathway. J Biotechnol 272–273:14–21. https://doi.org/10.1016/j.jbiotec.2018.02.012
Suo Y, Ren M, Yang X et al (2018b) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production with high butyrate/acetate ratio. Appl Microbiol Biotechnol. https://doi.org/10.1007/s00253-018-8954-0
Suo Y, Liao Z, Qu C et al (2019) Metabolic engineering of Clostridium tyrobutyricum for enhanced butyric acid production from undetoxified corncob acid hydrolysate. Bioresour Technol 271:266–273. https://doi.org/10.1016/j.biortech.2018.09.095
Thauer RK, Jungermann K, Henninger H et al (1968) The energy metabolism of Clostridium kluyveri. Eur J Biochem 4:173–180. https://doi.org/10.1111/j.1432-1033.1968.tb00189.x
Thauer RK, Kaster A-K, Seedorf H et al (2008) Methanogenic archaea: ecologically relevant differences in energy conservation. Nat Rev Microbiol 6:579–591. https://doi.org/10.1038/nrmicro1931
Tsai SP, Lee YH (1988) Application of metabolic pathway stoichiometry to statistical analysis of bioreactor measurement data. Biotechnol Bioeng 32:713–715. https://doi.org/10.1002/bit.260320517
Wang S, Huang H, Moll J, Thauer RK (2010) NADP+ reduction with reduced ferredoxin and NADP+ reduction with NADH are coupled via an electron-bifurcating enzyme complex in Clostridium kluyveri. J Bacteriol 192:5115–5123. https://doi.org/10.1128/JB.00612-10
Weghoff MC, Bertsch J, Müller V (2015) A novel mode of lactate metabolism in strictly anaerobic bacteria: a novel mode of lactate metabolism in anaerobes. Environ Microbiol 17:670–677. https://doi.org/10.1111/1462-2920.12493
Yu L, Zhao J, Xu M et al (2015) Metabolic engineering of Clostridium tyrobutyricum for n-butanol production: effects of CoA transferase. Appl Microbiol Biotechnol 99:4917–4930. https://doi.org/10.1007/s00253-015-6566-5
Zeng K, Zhang D (2010) Recent progress in alkaline water electrolysis for hydrogen production and applications. Prog Energy Combust Sci 36:307–326. https://doi.org/10.1016/j.pecs.2009.11.002
Acknowledgements
We would like to thank Floriane Revardeau from INRAE–URTAL, Poligny, France for her help in the short chain fatty acid analysis.
Funding
This work was supported by the Regional Council of Bourgogne–Franche Comté, the “Fonds Européen de Développement Régional (FEDER).”
Author information
Authors and Affiliations
Contributions
EM, RC, HL, and EB planned the work and conceived and designed the experiments. EM performed the experiments and the genome searching and phylogenetic analyses. EB performed the analyses of short chain fatty acids. EM, RC, EB, and HL analyzed the results. EM wrote the paper, and EB, RC, and HL revised it. All the authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Munier, E., Licandro, H., Beuvier, E. et al. Bioinformatics and metabolic flux analysis highlight a new mechanism involved in lactate oxidation in Clostridium tyrobutyricum. Int Microbiol 26, 501–511 (2023). https://doi.org/10.1007/s10123-022-00316-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00316-y