Abstract
Consensus PCR assays that can be used to sensitively detect several herpesvirus (HV) species across the different subfamilies were developed in this study. Primers containing degenerate bases were designed to amplify regions of the DNA polymerase (DPOL) gene of alpha- and gamma-HVs, and the glycoprotein B (gB) gene of beta-HVs in a singleplex, non-nested touchdown PCR format. The singleplex touchdown consensus PCR (STC-PCR) was used to amplify the DNA of eight human and 24 animal HVs. The assay was able to detect the lowest DNA dilution of 10−5 for alpha-HVs and 10−3 for beta- and gamma-HVs. In comparison, lowest detection limits of 10−5, 10−3, and 10−2 were obtained for alpha-, beta-, and gamma-HVs respectively when a nested PCR was used. The findings in this study suggest that the STC-PCR assays can be employed for the molecular surveys and clinical detection of novel and known HVs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Herpesviruses (HVs) are known to have a wide host range, infecting both vertebrate and invertebrate species [1, 2]. The virus is made up of a linear, monopartite, double-stranded DNA genome that encodes up to 300 genes and ranges from 124 to 241 kbp in length [3]. Herpesviruses are divided into three subfamilies, the Alpha-, Beta-, and Gamma-herpesvirinae on the basis of biological and molecular properties [2]. A common feature among all of the sub-groups of HVs is their ability to cause latent infection in infected hosts, which can be reactivated to cause serious illness in immunocompromised hosts [2]. Clinical diseases associated with active or recrudescent HV infections vary according to the hosts and the infecting viral species. For instance, the human HVs (HHV-1 to HHV-8) are members of Alpha-, Beta- and Gamma-herpesvirinae and have been associated with gingivostomatitis, herpetic keratitis, encephalitis, varicella, mononucleosis, lymphoproliferative malignancy, roseola and sarcoma [2]. The HVs of ruminants belong to the subfamilies Alpha- and Gamma-herpesvirinae, and infections are associated with rhinotracheitis (ovine HV1, caprine HV1), herpes mammalitis (bovine HV2), meningoencephalitis (bovine HV5), fatal systemic infection (caprine HV1), malignant catarrhal fever (ovine HV2, alcelaphine HV1, 2), ocular disease (cervine HV1) and fatal neurological disorder (bubaline HV1) [4]. The avian and reptilian HVs have so far only been assigned to the subfamily Alphaherpesvirinae causing clinical and economic important diseases such as Marek’s disease (gallid HV2) and infectious laryngotracheitis (gallid HV1) in poultry, duck plaque enteritis (anatid HV1) in waterfowl, Pacheco’s disease (psittacid HV1) in psittacines and fibropapillomatosis (chelonid HV5) in sea turtles [5,6,7,8,9,10]. Mixed infections of HV species can occur in susceptible hosts leading to a variety of clinical symptoms that may be difficult to diagnose or treat [11,12,13,14,15]. Therefore, there is a need for a sensitive assay that can reliably detect HV species of more than one subfamily in the same clinical samples.
Molecular surveys often employ consensus PCR assays to detect known and novel HVs [16,17,18,19]. In fact, several new HV species of mammals, reptiles and avians have been discovered using this approach [18, 20,21,22,23,24,25]. Despite these valuable outcomes, some of the existing consensus PCR assays have variable sensitivity to different HV subfamilies and require a nested PCR format, which can be costly and prone to contamination. Here, we have designed singleplex touchdown consensus PCRs (STC-PCRs) that amplify regions of the DNA polymerase (DPOL) gene of alpha- and gamma-HVs and glycoprotein B (gB) gene of beta-HVs. This non-nested PCR assay was successfully used to detect a wide range of HVs across a broad range of herpesviruses in two independent laboratories.
Materials and methods
Primer design
Degenerate consensus primers were designed for each subfamily based on the alignment of full and partial nucleotide sequences of HVs obtained from GenBank (Supplementary file 1, Table S1). The HV sequences were imported into Geneious 11.1.5 (https://www.geneious.com), and alignments were conducted with ClustalW 2.1 using the default parameters. Primers (Table 1) were manually generated from the conserved regions following visual inspections of the alignments.
DNA preparation and Singleplex Touchdown Consensus PCR
The human and animal HV DNAs tested in this study are shown in Table 2. Viral DNAs were extracted from infected tissues or culture supernatants using the DNeasy Blood and Tissue Kit (Qiagen) as recommended by the manufacturer. Additional DNA extracts were obtained from the Victorian Infectious Diseases Reference Laboratory (VIDRL).
Following assay optimisation (data not shown), the STC-PCR was used to amplify HV DNA in a 20-μL reaction. The reaction mix contained 2 μL of DNA template, 1 μM (beta-HV) or 2 μM (alpha-HV and gamma-HV) primers (Table 1), 200 μM of each deoxynucleotide triphosphate (dNTP), 1.5 mM MgCl2, 0.5 U of HotStarTaq polymerase and 1× PCR buffer (Qiagen). The assays were successfully evaluated with ready-to-use pre-mixes including the GoTaq Hot Start Green Master Mix (Promega) and the HotStarTaq Plus master mix (Qiagen) to ensure the assays could be used across a range of PCR chemistries (data not shown). PCR enhancers, including 5% dimethyl sulfoxide (DMSO) and tetramethylammonium chloride (15 mM; TMAC), were also added to the reaction mix. A Touchdown PCR protocol was carried out as outlined in Table 3. The PCR products were analysed on a 1.5% agarose gel made up of 1× TBE buffer and 1× GelRed nucleic acid stain (Biotium).
The specificity of the herpesvirus consensus assays was evaluated by testing a large number of alphaherpesviruses (n = 22), betaherpesviruses (n = 3) and gammaherpesviruses (n = 6). The assay performance was compared to another commonly used herpesvirus nested consensus PCR [19]. The STC-PCR relative sensitivity was tested by assaying a series of 10-fold dilutions of the DNA extracts of representative HVs from each subfamily, and comparing the limit of detection (LOD) with the VanDevanter assay [19]. The assay specificity was also checked by testing the consensus primer pair of one subfamily with the HV DNA templates of other subfamilies. To assess the reproducibility of the assay, herpesviruses were tested using the assays at two independent laboratories, with 22 viruses tested at James Cook University (Townsville, Queensland) and 15 viruses tested at the Australian Centre for Disease Preparedness (Geelong, Victoria).
Results
Overall, a total of 56 primers targeting the conserved regions of different HV genes were designed and tested with a wide range of HV DNAs. Of these, the three primer pairs reported in this study (Table 1; Supplementary file 1, Figure S1) were found to sensitively amplify the DNA sequences of 32 HV species (Table 2). In addition, appropriately sized (specific) single bands were seen (for most of the HVs tested) on agarose gel following electrophoresis (Fig. 1). The addition of 5% DMSO and 15 mM TMAC greatly improved the sensitivity, specificity, and reproducibility of the PCR reaction (Supplementary file 1, Figure S2).
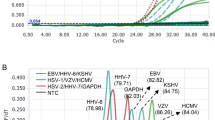
Electrophoresis of PCR products of HV DNAs obtained by STC-PCR in a 1.5% agarose gel. Lane 1 and 18 contain a 100 bp DNA marker; Lane 2 = Bovine alphaherpesvirus 1; lane 3 = Chelonid alphaherpesvirus 5, lane 4 = Macropodid alphaherpesvirus 1; lane 5 = Macropodid alphaherpesvirus 2; lane 6 = Human alphaherpesvirus 1; lane 7 = Human alphaherpesvirus 2; lane 8 = Human alphaherpesvirus 3; lane 9 = Equid alphaherpesvirus 4; lane 10 = Meleagrid alphaherpesvirus 1; lane 11 = Gallid alphaherpesvirus 2; lane 12 = Felid alphaherpesvirus 1; lane 13 = Human betaherpesvirus 5; lane 14 = Human betaherpesvirus 6; lane 15 = Human betaherpesvirus 7; lane 16 = Human gammaherpesvirus 4; lane 17 = Human gammaherpesvirus 8
The detection limits (relative) of the STC-PCR were comparable or lower when compared to the previously reported nested-PCR (Table 2), except for human betaherpesvirus 6, for which the nested PCR detected the viral DNA at one 10-fold dilution lower (Supplementary file 1, Figure S3). The STC-PCR assays produced much ‘cleaner’ DNA gels than the nested-PCR, which consistently produced many non-specific bands.
Subfamily assay specificity tests showed that the primer pair of one subfamily did occasionally cross-amplify HV DNAs of the other subfamilies (Supplementary file 1, Figure S4). For instance, the alpha-HV primer pair (AlphaFWD1 and AlphaREV2) amplified the DNA of HHV-6 (faint band observed), a member of the subfamily Betaherpesvirinae, but did not amplify any gamma-HV DNA (Supplementary file 1, Figure S4). The beta-HV primer pair amplified the DNA of a gamma-HV, HHV-4 (faint band), but none of the alpha-HVs. The gamma-HV primers produced varying sized bands for some alpha-HVs including crocodyline HV1 (CrHV-1), meleagridid HV1 (MeHV-1), equine HV4 (EHV-4), bovine HV1 (BoHV-1), HHV-1 and HHV-2 (Supplementary file 1, Figure S4). None of the beta-HV DNA was amplified by the gamma-HV primers (Supplementary file 1, Figure S4).
Discussion
Despite the biological and evolutionary divergence of HVs across the three subfamilies, many evolutionarily conserved core genes still persist [26, 27]. These genes encode proteins that play essential roles in viral entry, nucleic acid synthesis and metabolism, capsid maturation, and virion egress [26]. The DPOL and gB genes are among the most highly conserved genes of HVs and have previously been used as biomarkers for the detection of HVs [18, 19]. In this present study, a singleplex PCR assay targeting conserved genes (DPOL or gB genes) at the subfamily level was developed and successfully used to amplify a broad spectrum of human and animal HV DNAs. Also, the assay produced bright single bands on an electrophoretic gel, which is essential for downstream amplicon sequencing and identification of novel and known HVs.
The addition of 5% DMSO and 15 mM TMAC enhanced the STC-PCR by increasing product yield and ensuring assay reproducibility. High GC content is a common feature of HV genomes [28], and this could pose a challenge during amplification. As previously observed [29, 30], DMSO assists in reducing complex secondary structures and high melting temperature (Tm) associated with GC-rich templates, which in turn reduces duplex stability and allows efficient PCR. TMAC is often recommended when using degenerate primers and helps prevent mispriming by improving stringency of the PCR [31, 32].
In a previous study by VanDevanter et al. [19], a nested PCR using degenerate primers was found to have LODs ranging from a single copy to 100 copies of HV Polymerase DNA per 100 ng of human DNA. Therefore, the sensitivity of the STC-PCR relative to the nested PCR was determined using 10-fold dilutions of representative HVs. The assays were comparable or more sensitive than the nested assay across almost all of the herpesviruses tested. With the improved sensitivity, coupled with cost and time savings, the STC-PCRs can be employed for the epidemiological and clinical detection of known and novel HVs. Some cross-amplification between herpesvirus subfamilies was observed with the STC-PCR due to the high conservation of the targeted DPOL and gB genes at the family level. We consider this cross-amplification a universal feature of the STC-PCR for HV detection; therefore, positive results (amplicons) should be sequenced for onward identification and classification of the detected HVs.
Herpesviruses have been shown to be important pathogens across a large range of vertebrate hosts [1]. Recent initiatives to investigate viral diversity in wildlife hosts have utilised universal PCR assays to discover novel viruses, some with potential clinical and zoonotic concerns [33, 34]. For instance, universal PCR was used to identify six novel herpesviruses in multi-infected samples of chimpanzees (Pan troglodytes verus) [35]. Similarly, novel herpesviruses associated with respiratory disease in birds and hepatitis and enteritis in monitor lizards have been detected using universal PCR approaches [36, 37]. Although universal PCR assays have been an invaluable tool for these viral discovery initiatives, many of these assays can be problematic due to poor sensitivity, low specificity and contamination issues (especially with nested assays). Here, we have designed and evaluated novel singleplex universal PCR assays that will be useful for detection of known and novel herpesviruses from human and animal clinical samples.
Data Availability
All data analysed during this study are included in this published article [Supplementary file 1].
Code Availability
Not applicable.
References
Kaján GL, Doszpoly A, Tarján ZL, Vidovszky MZ, Papp T (2020) Virus–host coevolution with a focus on animal and human DNA viruses. J Mol Evol 88(1):41–56. https://doi.org/10.1007/s00239-019-09913-4
Whitley R (1996) Herpesviruses. In: Baron S (ed) Medical Microbiology, 4th edn. University of Texas Medical Branch at Galveston, Galveston (TX), p 68
Chaitanya K (2019) Structure and organization of virus genomes. In: Genome and Genomics. Springer, Singapore, pp 1–30
Engels M, Ackermann M (1996) Pathogenesis of ruminant herpesvirus infections. Vet Microbiol 53(1):3–15. https://doi.org/10.1016/S0378-1135(96)01230-8
Okoh GR, Horwood PF, Whitmore D, Ariel E (2021) Herpesviruses in reptiles. Front Vet Sci 8. https://doi.org/10.3389/fvets.2021.642894
Dhama K, Kumar N, Saminathan M, Tiwari R, Karthik K, Kumar MA et al (2017) Duck virus enteritis (duck plague) - a comprehensive update. Vet Q 37(1):57–80. https://doi.org/10.1080/01652176.2017.1298885
Boodhoo N, Gurung A, Sharif S, Behboudi S (2016) Marek’s disease in chickens: a review with focus on immunology. Vet Res 47(1):119. https://doi.org/10.1186/s13567-016-0404-3
Gowthaman V, Kumar S, Koul M, Dave U, Murthy T, Munuswamy P et al (2020) Infectious laryngotracheitis: Etiology, epidemiology, pathobiology, and advances in diagnosis and control - a comprehensive review. Vet Q 40(1):140–161. https://doi.org/10.1080/01652176.2020.1759845
Katoh H, Ogawa H, Ohya K, Fukushi H (2010) A review of DNA viral infections in psittacine birds. J Vet Med Sci 72(9):1099–1106. https://doi.org/10.1292/jvms.10-0022
Jones K, Ariel E, Burgess G, Read M (2016) A review of fibropapillomatosis in green turtles (Chelonia mydas). Vet J 212:48–57. https://doi.org/10.1016/j.tvjl.2015.10.041
Kaneko H, Kawana T, Ishioka K, Ohno S, Aoki K, Suzutani T (2008) Evaluation of mixed infection cases with both herpes simplex virus types 1 and 2. J Med Virol 80(5):883–887. https://doi.org/10.1002/jmv.21154
Taj A, Jamil N (2018) Co-occurrence of Herpes simplex virus 1 and 2 in patients suspected with different neurological ailments in Karachi. Clin Microbiol Infect Dis 3(1):1–5. https://doi.org/10.15761/CMID.1000135
Razonable RR, Paya CV (2002) The impact of human herpesvirus-6 and -7 infection on the outcome of liver transplantation. Liver Transpl 8(8):651–658. https://doi.org/10.1053/jlts.2002.34966
Gruffat H, Manet E (2018) Co-infection EBV/KSHV - Une alliance efficace [EBV/KSHV co-infection: an effective partnership]. Med Sci 34(1):79–82. https://doi.org/10.1051/medsci/20183401017
Olson D, Huntington MK (2009) Co-infection with cytomegalovirus and Epstein-Barr virus in mononucleosis: case report and review of literature. S D Med 62((9)):349 51-53
Li H, Keller J, Knowles DP, Crawford TB (2001) Recognition of another member of the malignant catarrhal fever virus group: an endemic gammaherpesvirus in domestic goats. J Gen Virol 82(1):227–232. https://doi.org/10.1099/0022-1317-82-1-227
Porto GS, Leme RA, Dall Agnol AM, Souza T, Alfieri AA, Alfieri AF (2021) Porcine lymphotropic herpesvirus (Gammaherpesvirinae) DNA in free-living wild boars (Sus scrofa Linnaeus, 1758) in Brazil. J Vet Sci 22(6):e81. https://doi.org/10.4142/jvs.2021.22.e81
Ehlers B, Borchers K, Grund C, Frölich K, Ludwig H, Buhk HJ (1999) Detection of new DNA polymerase genes of known and potentially novel herpesviruses by PCR with degenerate and deoxyinosine-substituted primers. Virus Genes 18(3):211–220. https://doi.org/10.1023/a:1008064118057
VanDevanter DR, Warrener P, Bennett L, Schultz ER, Coulter S, Garber RL et al (1996) Detection and analysis of diverse herpesviral species by consensus primer PCR. J Clin Microbiol 34(7):1666–1671. https://doi.org/10.1128/JCM.34.7.1666-1671.1996
Andersson KE, Adamovicz L, Mumm LE, Winter JM, Glowacki G, Teixeira-Neto R et al (2021) Detection of a novel herpesvirus associated with squamous cell carcinoma in a free-ranging Blanding's turtle. J Vet Diagn Invest 33(2):348–351. https://doi.org/10.1177/1040638721989302
Coverdill CC, Barnes JA, Garner MM, Hinton KL, Childress AL, Wellehan JF Jr (2016) Phylogenetic characterization of a novel herpesvirus found in the liver and lungs of a Chilean flamingo (Phoenicopterus chilensis). J Vet Diagn Invest 28(3):219–224. https://doi.org/10.1177/1040638716641157
Licheri M, Origgi FC (2020) Consensus PCR protocols for the detection of amphibian herpesviruses (Batrachovirus). J Vet Diagn Invest 32(6):864–872. https://doi.org/10.1177/1040638720951134
Maboni G, Kelly EJ, Clancy CS, De Luca E, Baldwin TJ, Van Wettere AJ et al (2022) Detection of asinine gammaherpesviruses in association with pulmonary fibrosis in free-ranging donkeys. J Vet Diagn Invest 34(1):167–171. https://doi.org/10.1177/10406387211052998
Sano K, Okazaki S, Taniguchi S, Masangkay JS, Puentespina R Jr, Eres E et al (2015) Detection of a novel herpesvirus from bats in the Philippines. Virus Genes 51(1):136–139. https://doi.org/10.1007/s11262-015-1197-6
Smith JA, Wellehan JF Jr, Pogranichniy RM, Childress AL, Landolfi JA, Terio KA (2008) Identification and isolation of a novel herpesvirus in a captive mob of eastern grey kangaroos (Macropus giganteus). Vet Microbiol 129(3-4):236–245. https://doi.org/10.1016/j.vetmic.2007.11.019
Mocarski ES Jr (2007) Comparative analysis of herpesvirus-common proteins. In: Arvin A, Campadelli-Fiume G, Mocarski E, Moore PS, Roizman B, Whitley R, Yamanishi K (eds) Human Herpesviruses, Biology, Therapy, and Immunoprophylaxis, Cambridge, p 4
Nicholas J (2000) Evolutionary aspects of oncogenic herpesviruses. Mol Pathol 53(5):222–237. https://doi.org/10.1136/mp.53.5.222
Brown JC (2007) High G+C content of herpes simplex virus DNA: proposed role in protection against retrotransposon insertion. Open Biochem J 1:33–42. https://doi.org/10.2174/1874091X00701010033
Jensen MA, Fukushima M, Davis RW (2010) DMSO and betaine greatly improve amplification of GC-rich constructs in de novo synthesis. PLoS One 5(6):e11024. https://doi.org/10.1371/journal.pone.0011024
Hardjasa A, Ling M, Ma K, Yu H (2010) Investigating the effects of DMSO on PCR fidelity using a restriction digest-based method. J Exp Microbiol Immunol 14:161–164
Hung T, Mak K, Fong K (1990) A specificity enhancer for polymerase chain reaction. Nucleic Acids Res 18(16):4953. https://doi.org/10.1093/nar/18.16.4953
Kovárová M, Dráber P (2000) New specificity and yield enhancer of polymerase chain reactions. Nucleic Acids Res 28(13):E70. https://doi.org/10.1093/nar/28.13.e70
Onyuok SO, Hu B, Li B, Fan Y, Kering K, Ochola GO et al (2019) Molecular detection and genetic characterization of novel RNA viruses in wild and synanthropic rodents and shrews in Kenya. Front Microbiol 10:2696. https://doi.org/10.3389/fmicb.2019.02696
Latimer E, Zong J-C, Heaggans SY, Richman LK, Hayward GS (2011) Detection and evaluation of novel herpesviruses in routine and pathological samples from Asian and African elephants: identification of two new probosciviruses (EEHV5 and EEHV6) and two new gammaherpesviruses (EGHV3B and EGHV5). Vet Microbiol 147(1):28–41. https://doi.org/10.1016/j.vetmic.2010.05.042
Prepens S, Kreuzer KA, Leendertz F, Nitsche A, Ehlers B (2007) Discovery of herpesviruses in multi-infected primates using locked nucleic acids (LNA) and a bigenic PCR approach. Virol J 4:84. https://doi.org/10.1186/1743-422X-4-84
Shivaprasad HL, Phalen DN (2012) A novel herpesvirus associated with respiratory disease in Bourke's parrots (Neopsephotus bourkii). Avian Pathol 41(6):531–539. https://doi.org/10.1080/03079457.2012.732692
Hughes-Hanks JM, Schommer SK, Mitchell WJ, Shaw DP (2010) Hepatitis and enteritis caused by a novel herpesvirus in two monitor lizards (Varanus spp.). J Vet Diagn Invest 22(2):295–299
Acknowledgements
The authors would like to acknowledge and thank Dr Julian Druce (Victorian Infectious Diseases Reference Laboratory, Victoria, Australia), Prof Timothy Mahony (University of Queensland, Queensland, Australia), Dr. Karina Jones (Murdoch University), and Dr Cathy Shilton (Berrimah Veterinary Laboratory, Northern Territory, Australia) for providing viral samples and/or DNA for testing.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions This study was funded through an Australian Wildlife Society University Student Grant and a JCU Higher Degree Research Enhancement Scheme Grant.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 7561 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okoh, G.R., Lockhart, M., Grimsey, J. et al. Development of subfamily-based consensus PCR assays for the detection of human and animal herpesviruses. Eur J Clin Microbiol Infect Dis 42, 741–746 (2023). https://doi.org/10.1007/s10096-023-04605-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-023-04605-w