Abstract
Aetiological diagnosis of gastrointestinal infections is challenging since a wide range of bacteria, parasites and viruses can be causal agents and derived clinical manifestations appear quite similar. Our aim was to evaluate contribution of the novel QIAstat-DxGastrointestinal Panel (GIP) to aetiological diagnosis of gastrointestinal infections and rational antimicrobial prescription in a reference paediatric hospital. Evaluation included comparison of diagnostic yield and agreement of results of QIAstat-Dx GIP and conventional microbiological methods. Parallel testing was performed on stool samples collected prospectively from children admitted to Sant Joan de Deu Barcelona Hospital (Spain) during the period February–March 2019. Influence of the panel test use on antimicrobial prescription was assessed using a pre–post study design. Eighty-six (68.8%) out of 125 specimens were positive by QIAstat-Dx GIP versus 44 (35.2%) positive by a composite of conventional methods (p<0.001). Global agreement of panel test results with rotavirus-adenovirus antigen detection (92.8%) and a two-step antigen/toxin and PCR-based algorithm for toxigenic Clostridioides difficile detection (87.5%) was greater than that with bacterial culture (76.0%) and parasite microscopic identification (64.3%). Panel test results orientated antimicrobial prescription changes in 18 (14.4%) patients, including antimicrobial start in 11 cases initially untreated, targeted antimicrobial prescription in 5 and discontinuation in 2 cases empirically treated. Results showed that QIAstat-Dx GIP significantly expanded aetiological diagnosis of gastrointestinal infections compared to conventional microbiological methods while orientating a more judicious use of antimicrobial drugs in hospitalised children.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Diarrhoea remains an important global health problem that was estimated to cause 1.6 million deaths in 2016 [1]. The disease especially affects paediatric populations [2,3,4] and has been acknowledged as one of the five leading causes of mortality and disability-adjusted life years in children under 5 years of age [5]. Diarrhoea usually occurs in the context of an acute or chronic gastrointestinal infection. Aetiological diagnosis of gastrointestinal infections is challenging since a wide range of bacteria, parasites and viruses can be causal agents [6, 7], and derived clinical manifestations appear quite similar [8].
Traditionally, aetiological diagnosis of gastrointestinal infections has relied on diverse microbiological methods such as stool bacterial culture, parasite microscopic examination, antigen-based detection of specific viruses and nucleic acid amplification for DNA/RNA detection of single highly prevalent pathogens like Clostridioides difficile (C. difficile) or norovirus. In the last years, new rapid syndromic molecular tests that can simultaneously detect and identify pathogenic gastrointestinal bacteria, viruses and parasites have emerged to overcome limitations of conventional microbiological tests [9,10,11,12]. Timely and comprehensive detection of gastrointestinal pathogens is essential to guide targeted antimicrobial treatment, prevent infection transmission and improve clinical outcomes [13].
The QIAstat-Dx Gastrointestinal Panel (Qiagen, Germany), hereafter QIAstat-DxGIP, is a CE-marked molecular panel assay that allows detection of 14 bacteria, 4 parasites and 6 viruses in 1 step in about 1 h. The panel test includes the following targets: C. difficile toxin A/B, enteroaggregative Escherichia coli (EAEC), enteroinvasive E. coli (EIEC)/Shigella, enteropathogenic E. coli (EPEC), enterotoxigenic E. coli (ETEC) lt/st, pathogenic Campylobacter spp., Plesiomonas shigelloides, Salmonella spp., shiga-like toxin producing E. coli (STEC) stx1/stx2, shiga-like toxin producing E. coli (STEC) O:157:H7, Vibrio cholera, Vibrio parahaemolyticus, Vibrio vulnificus, Yersinia enterocolitica, Cyclospora cayetanensis, Cryptosporidium spp., Entamoeba histolytica, Giardia lamblia, adenovirus F40/41, astrovirus, norovirus GI, norovirus GII, rotavirus A and sapovirus (I, II, IV and V). This real-time PCR test performs sample preparation and analysis steps automatically within disposable cartridges and yields results in 70 min, including cycle threshold (Ct) values for detected targets. An internal control producing a positive signal validates the results of the test. Conversely, a negative signal from the internal control invalidates all negative results except for those targets positively identified.
The objective of this study was to assess the contribution of QIAstat-Dx GIP to aetiological diagnosis of gastrointestinal infections and antimicrobial stewardship in a reference paediatric hospital.
Materials and methods
Study design and setting
A prospective study was conducted to determine usefulness of QIAstat-Dx GIP and conventional microbiological methods for diagnosis of gastrointestinal infections and antimicrobial stewardship in Sant Joan de Deu Barcelona Hospital (Spain) during February–May 2019. The setting is a 318-bedsize reference university paediatric hospital that attends a population of approximately 300,000 children. Inclusion criteria were as follows: (1) use of fresh loose stool samples collected from patients ≤18 years of age hospitalised in the study site with suspicion of gastrointestinal infection and diarrhoea symptomatology; (2) stool testing by QIAstat-Dx GIP, bacterial culture and rotavirus-adenovirus antigen detection, as well as microscopic examination for identification of parasites and performance of a 2-step antigen/toxin and PCR-based algorithm for toxigenic C. difficile detection in specific patients; and (3) selection of the first stool sample produced by each patient. Impact of the panel test utilisation on antimicrobial prescribing decisions was evaluated using a pre–post study design.
Outcomes sought were diagnostic yield of QIAstat-Dx GIP and conventional pathogen-targeted tests, agreement between tests and effect of the new syndromic test use on antimicrobial prescription changes.
Diagnostic methods and interpretation of results
Stool specimens were collected and routinely processed on demand by bacterial culture to identify E. coli, Campylobacter spp., Plesiomonas shigelloides, Salmonella spp., Vibrio cholera, Vibrio parahaemolyticus, Vibrio vulnificus and Yersinia enterocolitica. Culturing was performed according to standard operational procedures of the study setting. Identification of rotavirus and adenovirus was carried out using the antigen SD Rota/Adeno Rapid test following the manufacturer’s instructions. Parasites including Cryptosporidium spp., Cyclospora cayetanensis, Entamoeba histolytica and Giardia lamblia were identified by microscopy observation of their morphological characteristics. Toxigenic C. difficile detection combined the use of the C. Diff QuikChek Complete test (Abbott, USA) for C. difficile antigen and toxin detection and, if it was the case, the PCR-based GeneXpert® CD assay (Cepheid, USA) for confirmation of C. difficile antigen-positive and toxin-negative results, according to instructions of manufacturers. Microscopy observations for parasites and sequential tests for toxigenic C. difficile infection were only requested for a determinate number of samples at the clinician’s discretion. All samples were also tested by QIAstat-Dx GIP in parallel.
A multi-disciplinary board including a paediatrician, a paediatric infectious diseases specialist and a microbiologist jointly interpreted results of tests and reassessed diagnostic classifications and adequacy of baseline antimicrobial prescriptions.
Statistical analysis
Percent agreement between results by QIAstat-Dx GIP and conventional methods and kappa coefficients were calculated as described elsewhere [14]. Differences in proportions of positivity rates between tests and extent of antimicrobial prescription changes due to QIAstat-Dx GIP results were determined by the Chi-square or the exact test. Statistical significance was set at a p-value of <0.05 and confidence intervals (CI) at 95% level. All statistical analyses were performed using Stata v.15.1 software (Stata Corp.).
Results
Sample testing methods and patient characteristics
A total of 146 stool specimens were collected during the study period. Twenty-one (14.4%) of them were discarded, including 15 samples not tested by culture and/or rotavirus-adenovirus antigen detection, 5 samples that yielded invalid results by QIAstat-Dx GIP (negative for the internal control and for any target) and 1 sample not tested by the panel test. One hundred and twenty-five specimens from 125 patients were finally included in the study and underwent QIAstat-Dx GIP as well as conventional stool culture and rotavirus-adenovirus antigen testing. Additionally, 24 (19.2%) and 14 (11.2%) samples were tested for antigen/toxin- and PCR-based toxigenic C. difficile detection and parasite identification, respectively.
Seventy-three (58.4%) patients were male. Median age of patients was 20.4 months (IQR, 8.0–65.9 months). A majority of participants were recruited in the general paediatric ward (n=81, 64.8%), followed by those staying in haematology (n=12, 9.6%) and oncology (n=9, 7.2%) wards.
Laboratory diagnostic results
Eighty-six (68.8%) out of 125 specimens were found positive by QIAstat-Dx GIP. A single pathogen was detected in 50 (58.1%) specimens whereas co-detection of 2, 3 and 4 pathogens was observed in 26 (30.2%), 8 (9.3%) and 2 (2.3%) samples, respectively. A total of 134 pathogens were identified in the 86 positive samples, rotavirus being the most prevalent species (n=40), followed by toxigenic C. difficile (n=17) and norovirus GII (n=14). QIAstat-Dx GIP identified co-infections in 36 (28.8%) specimens. Fourteen samples positive for toxigenic C. difficile corresponded to children aged less than 2 years. Mean time to result was 10 h (IQR, 4–24 h).
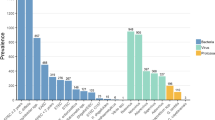
Forty-four (35.2%) samples yielded positive results and 45 pathogens were identified by conventional microbiological methods. Rotavirus-adenovirus was found in 34 (27.2%) out of the 125 samples and toxigenic C. difficile in 4 (16.7%) out of 24 samples, either directly by C. Diff QuikChek Complete antigen and toxin detection test (n=1) or after GeneXpert CD confirmation (n=3). Colonies of Campylobacter (n=4, 3.2%), Salmonella (n=1, 0.8%) and Yersinia (n=1, 0.8%) grew by stool culture. Only one bacterial-viral co-infection by toxigenic C. difficile and rotavirus-adenovirus was identified by the set of traditional diagnostic methods. Positivity rates for pathogen groups and at pathogen-level are detailed in Table 1. Overall distribution of pathogens by QIAstat-Dx GIP and conventional microbiological methods is depicted in Fig. 1. Supplementary Table 1 describes pathogen combinations identified by QIAstat-Dx GIP in co-infected samples.
Overall agreement of QIAstat-Dx GIP results with those of individual conventional techniques ranged from high for rotavirus-adenovirus (92.8%) and toxigenic C. difficile (87.5%) identification, to fair for bacteria identification by stool culture (76.0%), to poor for parasite identification by microscopy (64.3%). Overall, positive and negative percent agreement values and kappa coefficients are detailed in Table 2. There were 13 discrepant results in samples tested by both QIAstat-DX GIP and conventional methods for detection of shared targets, including 5 positives for Campylobacter spp. by QIAstat-DX GIP and negative by stool culture, 4 positives for Giardia lamblia by QIAstat-DX GIP and negative by microscopy, two positives for toxigenic C. difficile by QIAstat-DX GIP and negative by the 2-step algorithm, one negative for toxigenic C. difficile by QIAstat-DX GIP and positive by the 2-step algorithm, and one positive for Salmonella spp. by QIAstat-DX GIP and negative by stool culture.
Contribution of QIAstat-Dx GIP to targeted antimicrobial prescription
More comprehensive aetiological diagnosis of gastrointestinal infections by the panel test led to antimicrobial treatment changes for 18 (14.4%) out of the 125 patients, representing a significant effect (p<0.001) in comparison to baseline prescriptions. Eleven out of 83 patients that had not initially been treated with antimicrobials started receiving them upon identification of bacterial (n=5), parasitic (n=5) and bacterial-parasitic co-infections (n=1), including two patients younger than 2 years of age that were found positive for toxigenic C. difficile by QIAstat-Dx GIP. Among the remainder 42 patients that received antimicrobials empirically on admission, 5 were prescribed a more targeted therapy following identification of bacterial (n=4) and parasitic (n=1) infections, and 2 had antimicrobials discontinued once a viral aetiology had been confirmed. Seven (38.9%) out of the 18 antimicrobial prescription changes affected onco-haematology inpatients, a subgroup with a significantly lower participation weight (16.8%, p=0.03) in relation to the total study population. Details of pathogens identified before and after panel testing and subsequent antimicrobial treatment changes are explained in Table 3.
Discussion
As far as we know, our study is the first to report a significant influence of the use of a syndromic molecular test, the QIAstat-Dx GIP, on antimicrobial prescription practices for children hospitalised with suspected gastrointestinal infections. Interestingly, Beal et al. documented a trend for a lower intensity of antibiotic utilisation that did not reach statistical significance when comparing two groups of hospitalised children tested by another multiplex molecular assay, the FilmArray Gastrointestinal Panel (FA GIP), and stool culture [15]. We speculate that the comparatively higher effect of syndromic molecular testing on antimicrobial prescription practices observed in our study could partly be due to the noticeable number of children with oncology-haematology conditions, since this subgroup was subject to approximately 1.5 times more post-test antimicrobial prescription changes than the rest of participants.
Among inpatients of all ages, Cybulski et al. described that subjects diagnosed by FA GIP were significantly more prone to receive targeted than empirical therapy compared to those diagnosed by culture [16], whereas Axelrad et al. observed a significant reduction in antibiotic prescription within 14 days since stool testing after implementation of FA GIP, in comparison with conventional stool culture and antigen detection of adenovirus, rotavirus and parasites [11]. In contrast, Freeman et al. in their systematic review of syndromic molecular testing for GI did not identify robust evidence of antimicrobial optimisation and other clinical impacts following the introduction of panel tests [17].
The new syndromic test was able to unveil aetiology of gastrointestinal infections for the majority of study patients and had a nearly two-fold higher detection rate than targeted microbiological assays. Recent studies comparing QIAstat-Dx GIP with FA GIP and laboratory-developed RT-PCR assays reported similar diagnostic performance by the compared tests for common targets [18, 19]. Other comparative studies between multiplex molecular tests have also described similar capabilities to identify gastrointestinal pathogens [20, 21]. We hypothesise that the positive results yielded by QIAstat GIP in 9 samples (5 negative for Campylobacter spp. by stool culture and 4 negative for Giardia lamblia by microscopy observation) may probably be true positives, since real-time PCR tests like QIAstat GIP are widely acknowledged to have higher accuracy than the conventional methods that were used as comparators. A balance of discrepancies was observed between results by QIAstat GIP and the 2-step antigen- and PCR-based method for detection of toxigenic C. difficile in 3 samples (two pairs of positive results by the former method and negative by the latter, one pair of negative results by the former method and positive by the latter). This balance could indicate that both methods have equivalent diagnostic accuracy, as reflected by the high overall agreement of their results (92.8%). Generally, syndromic molecular testing appears to correctly identify gastrointestinal pathogens in concordance with results of conventional microbiological methods while additionally finding a larger number of pathogenic species and co-infections that would otherwise have been missed [11, 16, 17].
Studies on asymptomatic carriers have raised considerable uncertainty about whether or not additional gastrointestinal infection-positive results delivered by panel tests may be clinically important. Presence of C. difficile or its toxin in children under 2 years of age may be confounding since up to 70% of healthy newborns are colonized asymptomatically, a rate that gradually falls in parallel to the establishment of the intestinal microbiota by about that age [22]. Pathogenicity of EPEC and EAEC is also open to debate [23], given that these pathotypes have commonly been found in immunosuppressed cancer patients [24]. So is the identification of gastrointestinal viruses, as noticeable carriage rates of gastrointestinal viruses have been observed in asymptomatic children attending daycare centres [25]. Similarly, identification of Cryptosporidium and Giardia parasites in paediatric stool samples needs careful interpretation in light of non-negligible asymptomatic carriage rates reported among preschool children in a developed country [26] and the lack of a significant association between Giardia and diarrhoea documented in children up to 2 years of age in resource-limited settings [27]. In this regard, QIAstat-Dx GIP reporting of pathogen cycle threshold (Ct) values appears as an interesting feature that might contribute to discern the pathogenic or commensal role of specific gastrointestinal microorganisms. In addition, given the risk of antibiotic overuse or misuse in symptomatic patients, a prudent and cost-effective diagnostic strategy could be to prioritise patients with serious comorbidities or a deteriorating clinical course over milder patients for rapid testing by QIAstat-Dx GIP. Moreover, on-demand test use should be based on efficient pre- and post-analytical operational procedures to ensure that turnaround time for results keeps close to QIAstat-Dx GIP 70-min process time.
Interestingly, GI prevalence (68.8%) and co-infection (28.8%) rates determined by QIAstat-Dx GIP were remarkably higher than those reported in previous syndromic testing studies on paediatric study populations. In particular, Stockmann et al. reported a GI prevalence of 52% and a co-infection rate of 15% among children inpatients and outpatients with median age of 5 years [28], whereas Beckmann et al. determined the same prevalence rate for a subgroup of inpatients and outpatients predominantly aged 1–5 years [29]. We hypothesise that patient demographic and clinical characteristics as well as local epidemiology of GI and seasonal effects may explain these differences, since infants <2 years were significantly more prone to co-infection in our study than older children.
This study has some limitations for generalisation of results. First, it was a single-centre observational study with a relatively small sample size. Second, our study population included 14 infants under 2 years of age positive for toxigenic C. difficile by QIAstat-Dx GIP who might have been carriers and not cases. However, a potential bias in the real contribution of QIAstat-Dx GIP to targeted antimicrobial use in those infants was unlikely, since administration of antibiotics was only started in two of them initially untreated as a consequence of a positive result for C.difficile by the panel test. Third, we could only assess agreement of QIAstat-Dx GIP with conventional microbiological assays for specific targets only covered by the panel test. Four, re-testing of samples yielding discrepant results was not performed.
In conclusion, QIAstat-DxGIP use significantly improved aetiological diagnosis of gastrointestinal infections in hospitalised children in comparison with conventional microbiological methods. The contribution of this syndromic test to orientate specific antimicrobial prescription was also remarkable, particularly in onco-haematology paediatric inpatients. Further studies from multi-centre and experimental approaches are needed to confirm clinical impact of the new test.
Data availability
The dataset generated during the current study is available from the corresponding author on reasonable request.
References
Alemu Abajobir A, Abbafati C, Abbas KM et al (2017) Global, regional, and national age-sex specific mortality for 264 causes of death, 1980–2016: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Infect Dis 17:909–948
Liu L, Johnson HL, Cousens S et al (2012) Global, regional, and national causes of child mortality: an updated systematic analysis for 2010 with time trends since 2000. Lancet 379(9832):2151–2161
Boschi-Pinto C, Velebit L, Shibuya K (2008) Estimating child mortality due to diarrhoea in developing countries. Bull World Health Organ 86(9):710–717
Guerrant RL, Kosek M, Lima AAM et al (2002) Updating the DALYs for diarrhoeal disease. Trends Parasitol 18(5):191–193
Troeger C, Blacker BF, Khalil IA et al (2018) Estimates of the global, regional, and national morbidity, mortality, and aetiologies of diarrhoea in 195 countries: a systematic analysis for the Global Burden of Disease Study 2016. Lancet Infect Dis 18(11):1211–1228
Scallan E, Hoekstra RM, Angulo FJ et al (2011) Foodborne illness acquired in the United States-major pathogens. Emerg Infect Dis 17(1):7–15
Savioli L, Smith H, Thompson A (2006) Giardia and Cryptosporidium join the Neglected Diseases Initiative. Trends Parasitol 22(5):203–208
Thielman NM, Guerrant RL (2004) Acute infectious diarrhea. N Engl J Med 350(1):38–47
Freeman K, Tsertsvadze A, Taylor-Phillips S et al (2017) Agreement between gastrointestinal panel testing and standard microbiology methods for detecting pathogens in suspected infectious gastroenteritis: test evaluation and meta-analysis in the absence of a reference standard. PLoS One 12(3):e0173196
Binnicker MJ (2015) Multiplex molecular panels for diagnosis of gastrointestinal infection: performance, result interpretation, and cost-effectiveness. J Clin Microbiol 53(12):3723–3728
Axelrad JE, Freedberg DE, Whittier S, et al (2019) Impact of gastrointestinal panel implementation on health care utilization and outcomes. J Clin Microbiol 57(3):e01775–18
Zhang H, Morrison S, Tang YW (2015 Jun) Multiplex polymerase chain reaction tests for detection of pathogens associated with gastroenteritis. Clin Lab Med 35(2):461–486
Buss SN, Leber A, Chapin K et al (2015) Multicenter evaluation of the biofire filmarray gastrointestinal panel for etiologic diagnosis of infectious gastroenteritis. J Clin Microbiol 53(3):915–925
Cicchetti DV, Feinstein AR (1990) High agreement but low kappa: II. Resolving the paradoxes. J Clin Epidemiol 43(6):551–558
Beal SG, Tremblay EE, Toffel S et al (2018) A gastrointestinal PCR panel improves clinical management and lowers health care costs. J Clin Microbiol 56(1):1–25
Cybulski RJJ, Bateman AC, Bourassa L et al (2018) Clinical impact of a multiplex gastrointestinal polymerase chain reaction panel in patients with acute gastroenteritis. Clin Infect Dis 67(11):1688–1696
Freeman K, Mistry H, Tsertsvadze A et al (2017) Multiplex tests to identify gastrointestinal bacteria, viruses and parasites in people with suspected infectious gastroenteritis: a systematic review and economic analysis. Health Technol Assess 21(23):1–188
Hannet I, Engsbro AL, Pareja J, et al (2019) Multicenter evaluation of the new QIAstat-Dx Gastrointestinal Panel for the rapid syndromic testing of acute gastroenteritis. Eur J Clin Microbiol Infect Dis 38(11):2103–2112
Boers SA, Peters CJA, Wessels E et al (2020) Performance of the QIAstat-Dx Gastrointestinal Panel for diagnosing infectious gastroenteritis. J Clin Microbiol 58(3):e01737–e01719
Yalamanchili H, Dandachi D, Okhuysen PC (2018) Use and interpretation of enteropathogen multiplex nucleic acid amplification tests in patients with suspected infectious diarrhea. Gastroenterol Hepatol 14(11):646–652
Yoo J, Park J, Lee HK et al (2019) Comparative evaluation of Seegene Allplex Gastrointestinal, Luminex xTAG Gastrointestinal Pathogen Panel, and BD MAX enteric assays for detection of gastrointestinal pathogens in clinical stool specimens. Arch Pathol Lab Med 143(8):999–1005
Shane AL, Mody RK, Crump JA et al (2017) 2017 Infectious Diseases Society of America Clinical Practice Guidelines for the Diagnosis and Management of Infectious Diarrhea. Clin Infect Dis 65(12):e45–e80
Schuetz AN (2019) Emerging agents of gastroenteritis: Aeromonas, Plesiomonas, and the diarrheagenic pathotypes of Escherichia coli. Semin Diagn Pathol 36(3):187–192
Chao AW, Bhatti M, DuPont HL et al (2017) Clinical features and molecular epidemiology of diarrheagenic Escherichia coli pathotypes identified by fecal gastrointestinal multiplex nucleic acid amplification in patients with cancer and diarrhea. Diagn Microbiol Infect Dis 89(3):235–240
Hebbelstrup Jensen B, Jokelainen P et al (2019) Children attending day care centers are a year-round reservoir of gastrointestinal viruses. Sci Rep 9(1):1–8
Davies AP, Campbell B, Evans MR et al (2009) Asymptomatic carriage of protozoan parasites in children in day care centers in the United Kingdom. Pediatr Infect Dis J 28(9):838–840
Miller M, Acosta AM, Chavez CB et al (2014) The MAL-ED study: a multinational and multidisciplinary approach to understand the relationship between enteric pathogens, malnutrition, gut physiology, physical growth, cognitive development, and immune responses in infants and children up to 2 years of age in resource-poor environments. Clin Infect Dis 59(Suppl 4):S193–S206
Stockmann C, Pavia AT, Graham B et al (2017) Detection of 23 gastrointestinal pathogens among children who present with diarrhea. J Pediatric Infect Dis Soc 6(3):231–238
Beckmann C, Heininger U, Marti H et al (2014) Gastrointestinal pathogens detected by multiplex nucleic acid amplification testing in stools of pediatric patients and patients returning from the tropics. Infection 42(6):961–970
Acknowledgements
The authors would like to thank Fundación Godia for contracting SGG.
Declarations
The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Ethics approval
The study was approved by the Ethics Committee of Hospital Sant Joan de Deu Barcelona Hospital prior to start.
Consent to participate
Not applicable
Consent for publication
All the authors give their consent for publication of the study.
Conflict of interest
CMA reports research grants to her institution from Pfizer, Biofire Diagnostics, Roche Diagnostics, BioMérieux, Alere and Genomica SAU, and compensation fees from Qiagen, Roche Diagnostics and BioMérieux for scientific presentations in satellite symposiums outside the submitted work; PB reports compensation fees from Roche Diagnostics for scientific presentations in satellite symposiums outside the submitted work. The rest of authors report no conflict of interest.
Funding
The study was partially funded by Qiagen and CIBER de Epidemiología y Salud Pública CIBERESP.
Author information
Authors and Affiliations
Contributions
MCF and SGG performed laboratory tests, acquired test result data, analysed and interpreted data, prepared the first draft manuscript and revised and approved the final version of the manuscript; SS, CL and MFS acquired clinical data, analysed and interpreted data and revised and approved the final version of the manuscript; MK acquired test result data, analysed and interpreted data and revised and approved the final version of the manuscript; AG analysed and interpreted data and revised and approved the final version of the manuscript; CMA and PB defined the study design, analysed and interpreted data and revised and approved the final version of the manuscript.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Table 1
Pathogen combinations identified by QIAstat-Dx GIP in co-infected samples. Values expressed as No. (%). Abbreviations: C. difficile, Clostridioides difficile; EPEC, enteropathogeni cEscherichia coli; EAEC, enteroaggregative E. coli; ETEC, enterotoxigenic E. coli; G. lamblia, Giardia lamblia (DOCX 13 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Castany-Feixas, M., Simo, S., Garcia-Garcia, S. et al. Rapid molecular syndromic testing for aetiological diagnosis of gastrointestinal infections and targeted antimicrobial prescription: experience from a reference paediatric hospital in Spain. Eur J Clin Microbiol Infect Dis 40, 2153–2160 (2021). https://doi.org/10.1007/s10096-021-04266-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-021-04266-7