Abstract
Purpose
The purpose of this study was to determine the incidence of Clostridium difficile infections (CDI) and to characterise the isolates in 14 departments of ten academic hospitals in Slovakia.
Methods
During a one-month study (September 2012) all unformed stool samples were investigated using a rapid test to detect the presence of GDH and toxins A/B. Positive samples were cultured anaerobically and C. difficile isolates were characterised by ribotyping, multiple-locus variable-number tandem repeats analysis, and gyrA, rpoB and ermB investigation.
Results
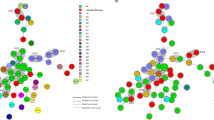
A total of 194 unformed stool samples were investigated and 38 (19.6 %) had a positive rapid test. Of 38 samples, 27 revealed a positive result for GDH and free toxins A/B in the stool, and 11 samples only for the presence of GDH. The mean CDI incidence in 2012 was 5.2 cases per 10,000 patient bed-days. Twenty C. difficile isolates were available for molecular analysis; seventeen belonged to PCR-ribotype 001 (85 %) whereas the remaining three isolates were identified as PCR-ribotypes 017, 078 and 449. MLVA of the PCR-ribotype 001 isolates identified two clonal complexes and a close genetic relatedness between isolates from six different hospitals. Molecular analysis of antibiotic-resistance determinants revealed the presence of ermB gene encoding resistance to the MLSB group of antibiotics in 90 % of isolates, and Thr82Ile amino acid substitution in the gyrA gene associated with resistance to fluoroquinolones in 85 % of isolates.
Conclusions
We conclude that C. difficile PCR-ribotype 001 is the predominant PCR-ribotype in Slovakia with a strong potential for clonal spread and development of multidrug resistance.



Similar content being viewed by others
References
Freeman J, Bauer MP, Baines SD, Corver J, Fawley WN, Goorhuis B, Kuijper EJ, Wilcox MH (2010) The changing epidemiology of Clostridium difficile infections. Clin Microbiol Rev 23(3):529–49
Bauer MP1, Notermans DW, van Benthem BH, Brazier JS, Wilcox MH, Rupnik M, Monnet DL, van Dissel JT, Kuijper EJ, ECDIS Study Group (2011) Clostridium difficile infection in Europe: a hospital-based survey. Lancet 377(9759):63–73. doi:10.1016/S0140-6736(10)61266-4
Davies KA, Longshaw CM, Davis GL, Bouza E, Barbut F, Barna Z, Delmée M, Fitzpatrick F, Ivanova K, Kuijper E, Macovei IS, Mentula S, Mastrantonio P, von Müller L, Oleastro M, Petinaki E, Pituch H, Norén T, Nováková E, Nyč O, Rupnik M, Schmid D, Wilcox MH (2014) Underdiagnosis of Clostridium difficile across Europe: the European, multicentre, prospective, biannual, point-prevalence study of Clostridium difficile infection in hospitalised patients with diarrhoea (EUCLID). Lancet Infect Dis 4(12):1208–19. doi:10.1016/S1473-3099(14)70991-0
Freeman J, Vernon J, Morris K, Nicholson S, Todhunter S, Longshaw C et al (2015) Pan-European Longitudinal Surveillance of antibiotic resistance among prevalent Clostridium difficile Ribotypes’ study group. Clin Microbiol Infect 21(3):248, e9-248.e16
Kuijper EJ, Coignard B, Tüll P, ESCMID Study Group for Clostridium difficile; EU Member States; European Centre for Disease Prevention and Control (2006) Emergence of Clostridium difficile-associated disease in North America and Europe. Clin Microbiol Infect 12(6):2–18, Review
McDonald LC, Coignard B, Dubberke E, Song X, Horan T, Kutty PK; Ad Hoc Clostridium Difficile Surveillance Working Group (2007) Recommendations for surveillance of Clostridium difficile-associated disease. Infect Control Hosp Epidemiol 28(2):140–5
Bauer MP, Kuijper EJ, van Dissel JT, European Society of Clinical Microbiology and Infectious Diseases (2009) European society of clinical Microbiology and Infectious Diseases (ESCMID): treatment guidance document for Clostridium difficile infection (CDI). Clin Microbiol Infect 15(12):1067–79. doi:10.1111/j.1469-0691.2009.03099.x
Debast SB, Bauer MP, Kuijper EJ, European Society of Clinical Microbiology and Infectious Diseases (2014) European Society of Clinical Microbiology and Infectious Diseases: update of the treatment guidance document for Clostridium difficile infection. Clin Microbiol Infect 20(2):1–26. doi:10.1111/1469-0691.12418
Stubbs SL, Brazier JS, O'Neill GL, Duerden BI (1999) PCR targeted to the 16S-23S rRNA gene intergenic spacer region of Clostridium difficile and construction of a library consisting of 116 different PCR ribotypes. J Clin Microbiol 37(2):461–3
Indra A, Huhulescu S, Schneeweis M, Hasenberger P, Kernbichler S, Fiedler A, Wewalka G, Allerberger F, Kuijper EJ (2008) Characterization of Clostridium difficile isolates using capillary gel electrophoresis-based PCR ribotyping. J Med Microbiol 57:1377–82. doi:10.1099/jmm.0.47714-0
Van den Berg RJ, Schaap I, Templeton KE, Klaassen CH, Kuijper EJ (2007) Typing and subtyping of Clostridium difficile isolates by using multiple-locus variable-number tandem-repeat analysis. J Clin Microbiol 45(3):1024–8
Goorhuis A, Legaria MC, van den Berg RJ, Harmanus C, Klaassen CH, Brazier JS, Lumelsky G, Kuijper EJ (2009) Application of multiple-locus variable-number tandem-repeat analysis to determine clonal spread of toxin A-negative Clostridium difficile in a general hospital in Buenos Aires, Argentina. Clin Microbiol Infect 15(12):1080–6. doi:10.1111/j.1469-0691.2009.02759.x
Spigaglia P, Mastrantonio P (2004) Comparative analysis of Clostridium difficile clinical isolates belonging to different genetic lineages and time periods. J Med Microbiol 53(11):1129–36
Dridi L, Tankovic J, Burghoffer B, Barbut F, Petit JC (2002) gyrA and gyrB mutations are implicated in cross-resistance to ciprofloxacin and moxifloxacin in Clostridium difficile. Antimicrob Agents Chemother 46(11):3418–21
Curry SR, Marsh JW, Shutt KA, Muto CA, O'Leary MM, Saul MI, Pasculle AW, Harrison LH (2009) High frequency of rifampin resistance identified in an epidemic Clostridium difficile clone from a large teaching hospital. Clin Infect Dis 48(4):425–9. doi:10.1086/596315
Zaiß NH, Witte W, Nübel U (2010) Fluoroquinolone resistance and Clostridium difficile, Germany. Emerg Infect Dis 16(4):675–677. doi:10.3201/eid1604.090859
Wiuff C, Brown DJ, Mather H, Banks AL, Eastaway A, Coia JE (2011) The epidemiology of Clostridium difficile in Scotland. J Infect 62(4):271–9. doi:10.1016/j.jinf.2011.01.015
Novak A, Spigaglia P, Barbanti F, Goic-Barisic I, Tonkic M (2014) First clinical and microbiological characterization of Clostridium difficile infection in a Croatian University Hospital. Anaerobe 30:18–23. doi:10.1016/j.anaerobe.2014.07.007
Vanek J, Hill K, Collins J, Berrington A, Perry J, Inns T, Gorton R, Magee J, Sails A, Mullan A, Gould FK (2012) Epidemiological survey of Clostridium difficile ribotypes in the North East of England during an 18-month period. J Hosp Infect 81(3):209–12. doi:10.1016/j.jhin.2012.04.017
Alcalá L, Martín A, Marín M, Sánchez-Somolinos M, Catalán P, Peláez T, Bouza E, Spanish Clostridium Difficile Study Group (2012) The undiagnosed cases of Clostridium difficile infection in a whole nation: where is the problem? Clin Microbiol Infect 18(7):E204–13. doi:10.1111/j.1469-0691.2012.03883.x
Krutova M, Matejkova J, Nyc O (2013) First results of molecular typing of Clostridium difficile in the Czech Republic. Zprávy CEM (SZÚ, Praha) 22(12):399–401
Arvand M, Vollandt D, Bettge-Weller G, Harmanus C, Kuijper EJ, Clostridium difficile study group Hesse (2014) Increased incidence of Clostridium difficile PCR ribotype 027 in Hesse, vol 19. Euro Surveill, Germany, 2011 to 2013
Arvand M, Hauri AM, Zaiss NH, Witte W, Bettge-Weller G (2009) Clostridium difficile ribotypes 001, 017, and 027 are associated with lethal C. difficile infection in Hesse, vol 14. Euro Surveill, Germany, 45
Goorhuis A, Bakker D, Cover J, Debast SB, Harmanus C, Notermans DW, Bergwerff AA, Dekker FW, Kuijper EJ (2008) Emergence of Clostridium difficile infection due to a new hypervirulent strain, polymerase chain reaction ribotype 078. Clin Infect Dis 47:1162–70
Bakker D, Corver J, Harmanus C, Goorhuis A, Keessen EC, Fawley WN, Wilcox MH, Kuijper EJ (2010) Relatedness of human and animal Clostridium difficile PCR ribotype 078 isolates determined on the basis of multilocus variable number tandem-repeat analysis and tetracycline resistance. J Clin Microbiol 48:3744–3749
Knetsch C, Connor T, Mutreja A, van Dorp S, Sanders I, Browne H, Harris D, Lipman L, Keessen E, Corver J, Kuijper E, Lawley T (2014) Whole genome sequencing reveals potential spread of Clostridium difficile between humans and farm animals in the Netherlands, 2002 to 2011. Euro Surveill 19(45):20954
Peláez T, Alcalá L, Blanco JL, Álvarez-Pérez S, Marín M, Martín-López A, Catalán P, Reigadas E, García ME, Bouza E (2013) Characterization of swine isolates of Clostridium difficile in Spain: a potential source of epidemic multidrug resistant strains? Anaerobe 22:45–9. doi:10.1016/j.anaerobe.2013.05.009
Spigaglia P, Drigo I, Barbanti F, Mastrantonio P, Bano L, Bacchin C, Puiatti C, Tonon E, Berto G, Agnoletti F (2015) Antibiotic resistance patterns and PCR-ribotyping of Clostridium difficile strains isolated from swine and dogs in Italy. Anaerobe 31:42–6. doi:10.1016/j.anaerobe.2014.10.003
Janezic S, Zidaric V, Pardon B, Indra A, Kokotovic B, Blanco JL, Seyboldt C, Diaz CR, Poxton IR, Perreten V, Drigo I, Jiraskova A, Ocepek M, Weese JS, Songer JG, Wilcox MH, Rupnik M (2014) International Clostridium difficile animal strain collection and large diversity of animal associated strains. BMC Microbiol 14:173. doi:10.1186/1471-2180-14-173
Schneeberg A, Neubauer H, Schmoock G, Baier S, Harlizius J, Nienhoff H, Brase K, Zimmermann S, Seyboldt C (2013) Clostridium difficile genotypes in piglet populations in Germany. J Clin Microbiol 51(11):3796–803. doi:10.1128/JCM.01440-13
Taori SK, Wroe A, Hardie A, Gibb AP, Poxton IR (2014) A prospective study of community-associated Clostridium difficile infections: the role of antibiotics and co-infections. J Infect 69(2):134–44. doi:10.1016/j.jinf.2014.04.002
Sharp SE, Ruden LO, Pohl JC, Hatcher PA, Jayne LM, Ivie WM (2010) Evaluation of the C.Diff Quik Chek Complete Assay, a new glutamate dehydrogenase and A/B toxin combination lateral flow assay for use in rapid, simple diagnosis of Clostridium difficile disease. J Clin Microbiol 48(6):2082–6. doi:10.1128/JCM.00129-10
Eastwood K, Else P, Charlett A, Wilcox M (2009) Comparison of nine commercially available Clostridium difficile toxin detection assays, a real-time PCR assay for C. difficile tcdB, and a glutamate dehydrogenase detection assay to cytotoxin testing and cytotoxigenic culture methods. J Clin Microbiol 47(10):3211–7. doi:10.1128/JCM.01082-09
Planche TD, Davies KA, Coen PG, Finney JM, Monahan IM, Morris KA, O'Connor L, Oakley SJ, Pope CF, Wren MW, Shetty NP, Crook DW, Wilcox MH (2013) Differences in outcome according to Clostridium difficile testing method: a prospective multicentre diagnostic validation study of C. difficile infection. Lancet Infect Dis 13(11):936–45. doi:10.1016/S1473-3099(13)70200-7
Wilcox MH (2012) Overcoming barriers to effective recognition and diagnosis of Clostridium difficile infection. Clin Microbiol Infect 18(Suppl 6):13–20. doi:10.1111/1469-0691.12057
Spigaglia P, Barbanti F, Mastrantonio P, European Study Group on Clostridium difficile (ESGCD) (2011) Multidrug resistance in European Clostridium difficile clinical isolates. Antimicrob Chemother 66(10):2227–34. doi:10.1093/jac/dkr292
Solomon K, Fanning S, McDermott S, Murray S, Scott L, Martin A, Skally M, Burns K, Kuijper E, Fitzpatrick F, Fenelon L, Kyne L (2011) PCR ribotype prevalence and molecular basis of macrolide-lincosamide-streptogramin B (MLSB) and fluoroquinolone resistance in Irish clinical Clostridium difficile isolates. J Antimicrob Chemother 66(9):1976–82. doi:10.1093/jac/dkr275
van den Berg RJ, Claas EC, Oyib DH, Klaassen CH, Dijkshoorn L, Brazier JS, Kuijper EJ (2004) Characterization of toxin A-negative, toxin B-positive Clostridium difficile isolates from outbreaks in different countries by amplified fragment length polymorphism and PCR ribotyping. J Clin Microbiol 42(3):1035–41
Pituch H, Brazier JS, Obuch-Woszczatynski P, Wultanska D, Meisel-Mikolajczyk F, Luczak M (2006) Prevalence and association of PCR ribotypes of Clostridium difficile isolated from symptomatic patients from Warsaw with macrolide-lincosamide-streptogramin B (MLSB) type resistance. J Med Microbiol 55(Pt 2):207–13
Ackermann G, Degner A, Cohen SH, Silva J Jr, Rodloff AC (2003) Prevalence and association of macrolide-lincosamide-streptogramin B (MLS(B)) resistance with resistance to moxifloxacin in Clostridium difficile. J Antimicrob Chemother 51(3):599–603
Spigaglia P, Barbanti F, Mastrantonio P, Brazier JS, Barbut F, Delmée M, Kuijper E, Poxton IR, European Study Group on Clostridium difficile (ESGCD) (2008) Fluoroquinolone resistance in Clostridium difficile isolates from a prospective study of C. difficile infections in Europe. J Med Microbiol 57(Pt 6):784–9. doi:10.1099/jmm.0.47738-0
Drudy D, Kyne L, O'Mahony R, Fanning S (2007) gyrA mutations in fluoroquinolone-resistant Clostridium difficile PCR-027. Emerg Infect Dis 13(3):504–5
O'Connor JR, Galang MA, Sambol SP, Hecht DW, Vedantam G, Gerding DN, Johnson S (2008) Rifampin and rifaximin resistance in clinical isolates of Clostridium difficile. Antimicrob Agents Chemother 52(8):2813–7. doi:10.1128/AAC.00342-08
Acknowledgments
We would like to thank ECDIS-net and the European Study Group of Clostridium difficile infections (ESGCD, ESCMID) for their professional support.
We would like to thank Dr. James Partridge for proofreading the text.
Ethical statement
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
For this type of study formal consent was not required.
Funding
Supported by the Ministry of Health, Czech Republic Internal Grant Agency NT/14209-3 and MH CZ – DRO, University Hospital Motol, Prague, Czech Republic 00064203.
Conflict of interest
The authors declare that they have no competing interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nyc, O., Krutova, M., Liskova, A. et al. The emergence of Clostridium difficile PCR-ribotype 001 in Slovakia. Eur J Clin Microbiol Infect Dis 34, 1701–1708 (2015). https://doi.org/10.1007/s10096-015-2407-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-015-2407-9