Abstract
Introduction
Coronavirus disease 2019 (COVID-19) is an unprecedented pandemic, threatening human health worldwide. The need to produce novel small-molecule inhibitors against the ongoing pandemic has resulted in the use of drugs such as chloroquine, azithromycin, dexamethasone, favipiravir, ribavirin, remdesivir and azithromycin. Moreover, the reports of the clinical trials of these drugs proved to produce detrimental effects on patients with side effects like nephrotoxicity, retinopathy, cardiotoxicity and cardiomyopathy. Recognizing the need for effective and non-harmful therapeutic candidates to combat COVID-19, we aimed to develop promising drugs against SARS-COV-2.
Discussion
In the current investigation, high-throughput virtual screening was performed using the Comprehensive Marine Natural Products Database against five non-structural proteins: Nsp3, Nsp5, Nsp12, Nsp13 and Nsp15. Furthermore, standard precision (SP) docking, extra precision (XP) docking, binding free energy calculation and absorption, distribution, metabolism, excretion and toxicity studies were performed using the Schrӧdinger suite. The top-ranked 5 hits obtained by computational studies exhibited to possess a greater binding affinity with the selected non-structural proteins. Amongst the five hits, CMNPD5804, CMNPD20924 and CMNPD1598 hits were utilized to design a novel molecule (D) that has the capability of interacting with all the key residues in the pocket of the selected non-structural proteins. Furthermore, 200 ns of molecular dynamics simulation studies provided insight into the binding modes of D within the catalytic pocket of selected proteins.
Conclusion
Hence, it is concluded that compound D could be a promising inhibitor against these non-structural proteins. Nevertheless, there is still a need to conduct in vitro and in vivo studies to support our findings.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A new case of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was first exposed in Wuhan, China [1, 2]. It is a deadly virus which disseminates amongst humans and other mammals causing a broad range of infections from common cold to fatal diseases such as respiratory syndrome. SARS-CoV-2 is a round-shaped, single-stranded RNA virus consisting of 30,000 bp of genomic sequence incorporating approximately eleven open reading frames (ORFs) encoding both structural and non-structural proteins intricate in viral life cycle [3,4,5]. The first and foremost protein, playing a key role in the viral infection and adaptive immunity, is spike glycoprotein (SGP). SGPs usually project from the surface of mature virions which play a role in fusion, attachment and entry into the host cell. The virions bind to the angiotensin-converting enzyme 2 (ACE-2) receptor via SGPs and mediate viral infection. Thus, SGPs’ surface location renders it as a direct anti-CoV-2 target for most of the therapeutic interventions. After the entry of virus into the host cell, it releases its RNA into the cell cytoplasm followed by the translation of its replicase gene. The two overlapping ORFs (1a and 1b) of replicase gene translated to polyproteins 1a and 1b, which are subsequently cleaved by main protease/3-chymotrypsin-like protease (Mpro/3CLpro) and papain-like protease (PLpro), respectively. The cleavage of polyproteins results in the formation of 16 non-structural proteins (Nsp1–16). Amongst them, papain-like protease (Nsp3) [6, 7], main protease (Nsp5) [8, 9], RNA-dependent RNA polymerase (RdRp/Nsp12) [10, 11], helicase (Nsp13) [12, 13] and endoribonuclease/XendoU (Nsp15) [14, 15] are considered to be viable anti-CoV-2 targets (Fig. 1) [4, 16]. The details of these five non-structural proteins are represented in Table 1.
Because of the COVID-19 outbreak, pharmaceutical hubs, private and government organisations, institutions and biotechnology firms are under burdened to produce high-quality, safety and low-cost drugs or vaccines against the ongoing unprecedented pandemic. Therefore, drugs such as hydroxychloroquine, ivermectin, azithromycin and various antiviral drugs have been compassionately used for the management of COVID-19 [17]. Although significant data are available from the randomized clinical and preclinical studies, there were many shreds of evidence on the adverse effects of these therapies on virus-infected patients [18]. The major side effects that are encountered after administering the anti-COVID trial drugs are cardiomyopathy, nephrotoxicity, retinopathy, cardiotoxicity and hepatotoxicity [18,19,20]. As a result, the need for safer and more easily available solutions to combat this viral scourge has intensified.
Naturally occurring compounds are now amongst the most common sources of prototypes for antimicrobial and antiviral drugs that continue to be developed today. Thousands of substances derived from aquatic animals are currently being studied in depth for use in medicine, with more than forty now in the medicine market. Several chemical groups of naturally biologically active substances, such as peptides, flavonoids and alkaloids, have been successfully tested against SARS-CoV-2.
The global history in marine pharmacy attests to the enormous potential of marine species as raw materials for the production of novel medicinal substances. In the course of evolution, aquatic species have evolved several anti-infectious mechanisms and molecules to shield them from assaults of microbes and viruses that inhabit the marine environment. Compounds isolated from aquatic animals that suppress DNA and RNA viruses, including coronaviruses, have been discovered in a variety of structural groups, including polysaccharides, terpenoids, steroids, alkaloids and peptides. The various mechanisms used by these chemical groups to suppress coronaviruses account for their diversity.
Recognizing the uniqueness of their possibilities to treat a variety of diseases, we looked at a subset of its derivatives in this article to learn more about how they function as inhibitors against the above-mentioned viral non-structural proteins using in silico approaches [21,22,23]. Recent studies on the SARS-CoV-2 targets, such as Mpro, PLpro and RdRP, have highlighted the value of structure-based drug design in the search for powerful compounds from a variety of chemical libraries, including azadirachtins, ceramicines, withanolides and nucleotide analog medicines [24,25,26,27]. As a result, the creation of new medications for the control and treatment of this continuing pandemic may benefit from unique alternative evidence based on a molecular docking approach. In this paper, virtual screening and molecular docking were employed to screen a large comprehensive marine database library (https://cmnpd.org/) against non-structural proteins of SARS-CoV-2 such as Nsp3, Nsp5, Nsp12, Nsp13 and Nsp14. Furthermore, pharmacological and toxicological parameters were evaluated for the best hit compounds. The final hits were used for the designing of a novel compound (D), and its potential of inhibiting the Nsps through computational approaches was evaluated.
Materials and methods
Protein crystal structure preparation
The protein X-ray crystal structures of five non-structural proteins (Nsp3, Nsp5, Nsp12, Nsp13 and Nsp14) were retrieved from the protein databank, with accession IDs of 6LU7 [7], 6W9C [8], 6M71 [10], 6XEZ [13] and 6VWW [15], respectively. Protein preparation wizard available in the Glide module was used for the rectification of specific errors like missing loops and hydrogen atoms in the structure of the proteins during crystallographic studies [28,29,30]. Before optimization, water molecules located >3 Å were removed from the protein crystal structure. Furthermore, hydrogen bonds and missing side chain atoms were added and repaired using the module Prime. Minimization of protein was carried out using OPLS3e force field with heavy atom convergence criteria to a root-mean-square deviation (RMSD) of 0.3 Å [31]. Ramachandran plots of all five proteins were assessed to determine stereospecificity and favourable and unfavourable regions in the protein crystal structure. A 3D grid box of 10 Å was created, defining the co-crystallized ligand, active residues and dimensions of the catalytic site (Table 2) [32].
Comprehensive marine database library preparation
Approximately 47,451 compounds from marine origin were downloaded from Comprehensive Marine Natural Products Database (https://www.cmnpd.org). Ligands were prepared using the LigPrep module available in the Maestro interface of Schrödinger suite. Low-energy conformers were segregated, and possible ionization states were generated for the 2D structures of all the ligands at a physiological pH of 7.2 ± 0.2. All the ligands were minimized using an optimized OPLS3e force field by keeping all other options as default.
High-throughput virtual screening and molecular docking studies
High-throughput virtual screening (HTVS) was carried out in the binding pocket of Nsp3, Nsp5, Nsp12, Nsp13 and Nsp15 via the Glide module of the Schrӧdinger suite. Prior to docking, the non-structural proteins were made rigid and the ligands were made flexible. Before the analysis, the physicochemical characteristics of the database library (47,451 compounds) acquired from CMNPD were filtered. Virtual screening is divided into three categories: HTVS, SP and XP docking. These three docking methods were used to take out a possible lead molecule in a short period. Although HTVS quickly screens a huge number of molecules, the sampling techniques were limited, and the results could not be immediately evaluated. As a result, the ligands identified by HTVS were docked using SP, which selects an appropriate binding posture from a vast pool of ligands [33]. Furthermore, the top 10% of SP molecules was chosen for docking in the XP mode and examined using an XP visualizer. Based on their Glide score, Glide E-model and Glide energy, the top 10% of molecules was picked for free energy estimates [34,35,36,37].
Molecular mechanics with generalized Born surface area calculation
Binding free energy (BFE) calculations were carried out for XP-docked complexes using molecular mechanics with generalized Born surface area (MM-GBSA) approach. Docked complexes were minimized using local optimization feature in the module Prime. OPLS3e force field was employed to evaluate the BFE for a set of protein-ligand complexes [31].
where ΔGSA is the difference in surface area energies in the protein and ligand and the complexes, ΔGsolv is the difference between the solvation energies of the complexes and individual proteins and ligands and ΔEMM is the variation between the minimized energy of the protein-ligand complexes.
Pharmacological and toxicological parameters
After molecular docking studies, SwissADME (http://www.swissadme.ch/) was utilized to estimate the absorption, distribution, metabolism, excretion and toxicity (ADMET) characteristics of the compounds. This module calculates numerous attributes and offers values for descriptors that forecast ligand drug likeness. It also generates functional groups, rings and a variety of other components that are used to predict ligand descriptors [38]. This approach is also used to connect the physicochemical and pharmacokinetic features of the test ligands to support the discovery of small-molecule inhibitors.
Molecular dynamics simulation studies
Molecular dynamics simulation (MDS) studies were performed for the compound D with the selected non-structural proteins using OPLS3e force field. The systems were solvated using TIP4P water in orthorhombic boxes within the Desmond module [39, 40]. Counter ions were added to the systems to achieve neutralization. A tolerance of 1e−09 for long-range interactions and a 9-Å cut-off radius for short-range columbic were applied using smooth particle mesh Ewald method [41]. The MDS was performed for 200 ns each for the selected non-structural protein-ligand complexes under NPT conditions, which were maintained by Martyna-Tobias-Klein (1 bar pressure) and Nose-Hoover thermostat (300 K). A steepest descent method was used to minimize the gradient energy of optimized structures [42,43,44].
Results and discussion
The SARS-CoV-2 infection continues to disseminate like wildfire, demanding urgent attention from scientists across the globe to discover a possible therapeutic against viral infection.
Molecular docking studies, post docking analysis and binding free energy calculation
Molecular docking plays a crucial role in the designing of novel drugs against non-structural proteins of interest. It predicts the experimental binding mode and binding affinity of molecules within the catalytic site of the selected proteins. In the first mode of HTVS, the screening of a marine database library was carried out using the selected non-structural proteins. Nearly 30% of obtained hits was used for SP docking and XP docking analysis. The results of the molecular docking studies along with the 2D structures and IUPAC names of the predicted ligands are depicted in Table 3 and Fig. 2. The compounds having high negative Glide score and which are observed in common with all the proteins were described briefly.
Nsp3 (PDB ID: 6LU7)
From the docking results, it is clear that CMNPD5804 had the most favourable Gscore of −7.804 kcal/mol. The compound also well fitted within the catalytic pocket of Nsp3 and exhibited four hydrogen bonding (HB) interactions with the key amino acid residues. The –OH of the phenolic moiety formed a HB interaction with the Lys102. Furthermore, the –OH present at positions 4 and 8 of the quinoline nucleus formed two HB interactions with the Gln110 and Tyr154, respectively. In addition, the carbonyl group present at position 5 of the quinoline moiety also showed a HB interaction with the protonated Asn151. Furthermore, the compound has been stabilized in the catalytic pocket of Nsp3 by the formation of two π-π stacking interactions between the fused rings of quinoline nucleus and electron cloud of Phe294. Moreover, the quinoline nucleus of the compound was found to be crucial for the binding affinity as evident by its ΔGbind −75.56 kcal/mol and key interactions.
Nsp5 (PDB ID: 6W9C)
From the docking result, it is clear that CMNPD20924 had the most favourable Gscore of −8.928 kcal/mol against Nsp5. The compound also well fitted within the catalytic pocket of Nsp5 and exhibited four HB interactions with the key amino acid residues. The two –OH molecules of the catechol moiety formed two HB interactions with Gly100. Furthermore, the two –OH groups of resorcinol formed a HB interaction with Ser278 and the other –OH present at the meta position of this structure formed two HB interactions with Lys279 and Thr277.
Nsp12 (PDB ID: 6M71)
With the help of docking results, it can be seen that CMNPD20924 had the most suitable Gscore of −9.002 kcal/mol. The compound also interacted well with the catalytic pocket of Nsp12 and showed four HB interactions with the essential amino acid residues. The resorcinol moiety exhibited three HB interactions with Thr394, Arg249 and Arg457. Furthermore, the –OH moieties of the catechol moiety showed two HB interactions with Glu350 and Ser318. In addition, the carbonyl group present at position 2 of the dihydrofuran moiety showed a HB interaction with Phe396. Moreover, the catechol nucleus of the compound exhibited a π-cationic interaction with Arg349.
Nsp13 (PDB ID: 6XEZ)
With the aid of docking outcomes, CMNPD15988 had the most favourable Gscore of −8.569 kcal/mol with Nsp13. The compound also connected most precisely with catalytic residues with four HB interactions and one π-π interaction. The –OH of the imidazole moiety formed a HB interaction with Tyr38. The carbonyl group present at position 2 of the imidazole moiety formed a HB interaction with Asn209. The –NH of the pyrrole moiety and carbonyl group formed two HB interactions with Asp221 and Arg733. Moreover, the compound has been stabilized in the catalytic pocket of Nsp13 by the formation of salt bridge with Mg1003.
Nsp15 (PDB ID: 6VWW)
Given that, CMNPD5804 had the stronger Glide score of −8.639 kcal/mol with Nsp15. The compound was well locked with the catalytic pocket of Nsp15 and demonstrated three HB interactions with the amino acid residues. The –OH and –COOH moiety of the quinoline moiety formed two HB interactions with Lys290 and Ser294. Furthermore, the carbonyl group exhibited a HB interaction with Ser294. The phenolic moiety made a π-π interaction with Trp333.
ADMET properties
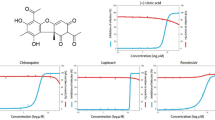
The SwissADME was used to calculate the pharmacological and toxicological (ADMET) characteristics, as well as to theorize on the acceptance of title compounds (CMNPD5804, CMNPD20924, CMNPD15988, CMNPD26614, CMNPD2414) (Table 4). The total amount of overall polar atoms or molecules in a compound is within the acceptable range (<120 Å2) of topological surface area (TSA) (primarily oxygen and nitrogen, including their bonded hydrogen atoms). None of the compounds was permeable through the blood-brain barrier. Amongst these five compounds, only CMNPD20924 was considered as a P-glycoprotein substrate and is constantly pumped out from the brain. Log Kp is a skin penetration indicator that shows the absorption of a substance via the skin. The results show that all drugs are absorbed through the skin. All these compounds had a substantial bioavailability score (BS), indicating that they had good permeability and bioavailability. Pains alert (PA) (PAN assay interference chemicals) were absent in all the compounds, which implicates a specific interaction with the chosen target and no contact with other unintended biological targets.
Design of novel compound
From the docking results, it is clear that the three compounds (CMNPD5804, CMNP15988 and CMNPD20924) exhibited significant interactions with the key amino acid residues of all non-structural proteins. The nuclei responsible for the interactions were isolated and conjoined to generate one molecule (D) (Fig. 3). Furthermore, docking studies and MM-GBSA were also performed to the designed molecule. The results were astonishing, and all the non-structural protein (D) complexes established greater binding affinity when compared with the hits obtained (Table 5, Fig. 4).
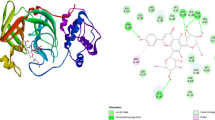
Two-dimensional diagram of a D/Nsp3, b D/Nsp5, c D/Nsp12, d D/Nsp13 and e D/Nsp15 docking complex exposing key interactions in the catalytic pocket of non-structural proteins. Furthermore, MDS studies were performed to get insights into binding modes of the complexes D/6LU7, D/6W9C, D/6M71, D/6XEZ and D/6VWW by analysing the trajectory frames
The complex D/6LU7 exhibited a Gscore of −7.239 kcal/mol and a ΔGbind score of −90.562 kcal/mol with four HB interactions. The –OH groups of resorcinol formed two HB interactions with Thr169 and Gly170, while the carbonyl oxygen linker showed a HB interaction with Lys137. On the other side of the compound, quinoline exhibited two HB interactions with Gln127 (–OH of the carboxyl moiety) and Lys137 (–OH at position 8). Furthermore, the compound was stabilized by the π-cationic interaction between the π-cloud of quinoline and the protonated Lys5 residue.
The complex D/6W9C showed a Gscore of −7.191 kcal/mol and a ΔGbind score of −67.119 kcal/mol and exhibited eight HB interactions. The –OH of resorcinol nucleus had a HB interaction with Gln97 whereas the –NH at position 3 of resorcinol exhibited a HB interaction with Gly100.The pyrrole –NH exhibited a HB interaction with Gln121, while the carbonyl oxygen linker showed a HB interaction with Lys279. The carboxyl moiety at position 5 and –OH at position 8 of the quinoline nucleus established four HB interactions with Gln122, Glu124 and Lys279.
The complex D/6M71 established a Gscore of −6.514 kcal/mol and a ΔGbind score of −98.478 kcal/mol and exhibited six HB interactions. The carboxyl moiety of quinoline nucleus exhibited two HB interactions with Ile266 and Trp268. The –NH of pyrrole exhibited a HB interaction with Phe321. The –OH groups of resorcinol formed two HB interactions with Asn459 and Pro677, while carbonyl oxygen linker showed a HB interaction with Arg349. Moreover, the compound was stabilized by π-π stacking interaction between the π-cloud of quinoline nucleus and the phenol ring of Tyr265.
The complex D/6XEZ exhibited a Gscore of −7.331 kcal/mol and a ΔGbind score of −119.69 kcal/mol and exhibited five HB interactions. The –OH and –COOH on quinoline nucleus exhibited two HB interactions with Thr324 and Pro378, respectively. The –NH of pyrrole nucleus exhibited a HB interaction with Phe326. The carbonyl oxygen of amide linker exhibited a HB interaction with Arg349, while the –OH of resorcinol exhibited a HB interaction with Val675.
The complex D/6VWW displayed a Gscore of −8.274 and a ΔGbind score of −67.771 and five HB interactions. The –OH group of resorcinol exhibited two HB interactions with Ser294 and Leu346. The carbonyl oxygen of amide linker exhibited a HB interaction with Lys290. Furthermore, nitrogen of quinoline nucleus exhibited a HB interaction with His235, while the –OH group present at position 8 of the quinoline nucleus formed a HB interaction with Asp240. The –NH of pyrrole nucleus exhibited a HB interaction with Thr341 while the carbonyl oxygen of amide linker exhibited a HB interaction with Lys290. The compound was stabilized by the formation of the π-cationic interaction between the π-cloud of pyrrole and the protonated Lys290 residue. In addition, ADMET properties of compound D using SwissADME revealed that the compound D possesses significant druggable properties with TSA, complying with the recommended range (<120 Å2). Furthermore, it exhibited non-permeability through the BBB and also non-substrate for poly-glycoprotein, indicating it cannot be pumped out of the brain or lumen of gastrointestinal tract. Compound D is also an inhibitor for only two cytochromes: CYP1A2 and CYP2C9. From the value of log Kp (−6.80 cm/s), it is clear that the compound D is permeated through the skin. Furthermore, it also exhibited zero PA, indicating that the interaction with the desired target is a specifically bypassing interaction with other targets.
RMSD (Fig. 5a–e) of initial structures of D/6LU7, D/6W9C, D/6M71 and D/6XEZ increased during equilibration and converged till 15 ns of a 200-ns MDS study, while the complex D/6VWW exhibited equilibration till 120 ns which may be attributed to the structural alterations of the protein residues.
After equilibrium, the RMSD values of D/6LU7 (Cα: 1.47–2.62 Å; BB: 1.47–2.17 Å; HA: 1.89–2.82 Å), D/6W9C (Cα: 1.86–2.81 Å; BB: 1.91–2.80 Å; HA: 2.35–3.18 Å), D/6M71 (Cα: 2.69–3.10 Å; BB: 2.61–3.01 Å; HA: 2.88–3.36 Å), D/6XEZ (Cα: 2.53–3.32 Å; BB: 2.52–3.31 Å; HA: 2.93–3.65 Å) and D/6VWW (Cα: 2.34–4.63 Å; BB: 2.35–4.61 Å; HA: 2.72–4.87 Å) indicated minimal fluctuations in the protein structure.
Furthermore, root-mean-square fluctuation (RMSF) of protein residues is presented in Fig. 6a–e. The D/6LU7 complex established lower RMSF values (<2 Å) at the ligand contacts Arg4 to Thr26, Gln69 to Gln74, Ala116 to Gln127, Lys137 to Gly143 and Glu166 to His172, indicating lower fluctuations of the residues. Furthermore, no ligand contacts were observed after His172 except Leu286, Glu288 and Glu290, whereas the complex D/6W9C exhibited lower RMSF values (<2 Å) at the ligand contacts Tyr95 to Thr102, Leu120 to Leu125, Ala139 to Ala144 and Thr277 to Glu280, indicating minimal fluctuations of the residues. In addition, no ligand contacts were observed between Arg3 and Lys94. The complex D/6M71 also exhibited lower RMSF values with the contacting residues Thr248 to Ser225, Pro264 to Trp268, Val315 to Gly327, Tyr456 to Thr462 and Tyr674 to Gly679, indicating minimal residual fluctuations. No ligand contacts were found between Thr680 and Leu931. Furthermore, the complex D/6XEZ exhibited lower RMSF values (<2 Å) with the contacting residues Tyr265 to Tyr273, Phe321 to Lys332, Pro378 to Ser397, Arg457 to Leu460 and Leu663 to Met666. Though significant fluctuations were found for some residues, they are not in contact with the ligand. The other complex D/6VWW was found to possess lower RMSF values (<2 Å) with the ligand contacting residues Glu229 to His250, Gly287 to Ser294, Val314 to Ser316 and Trp333 to Gln347, indicating minimal residual fluctuations and greater stability of protein-ligand interactions. Besides, no ligand contacts were observed between Leu3 and Leu228.
Analysis of MD trajectory of the compound D exposed hydrogen bonding, π-π stacking and π-cationic interactions binding pocket residues of the proteins Nsp3, Nsp5, Nsp12, Nsp13 and Nsp15, depicted in Supplementary Figs. S1a–e and S2a–e. All the complexes exhibited similar binding modes of interactions as speculated by the molecular docking study. The complex D/6LU7 established five hydrogen bonding interactions with Glu14 (15% and 25% MD trajectory), Gln19 (13% MD trajectory), Ty118 (43% MD trajectory), Asn119 (11% MD trajectory), Ser123 (13% MD trajectory; water-bridge interaction) and Ser139 (35% MD trajectory). Furthermore, the complex was stabilized by forming the π-π interaction with Phe140, whereas the complex D/6W9C exhibited stable hydrogen bonding interactions with the polar residues of the proteins Gln97 (31% MD trajectory), Thr102 (77% MD trajectory), Gln121 (36% MD trajectory) and Glu124 (71% MD trajectory) and the hydrophilic residue Tyr95 (48% MD trajectory; water-bridge interaction). Furthermore, the complex D/6M71 also exhibited significant hydrogen bonding interactions with most of hydrophilic residues. The –NH of the pyrrole linker established a stable hydrogen boding interaction with Phe321 (99% MD trajectory). The hydroxy groups of the resorcinol and –COOH moiety of quinoline established four hydrogen bonding interactions with Asn459 (85% MD trajectory), Glu350 (68% MD trajectory), Leu460 (25% MD trajectory; water-bridge interaction) and Ile266 (63% MD trajectory). Moreover, the complex was also stabilized by forming the π-π stacking interaction between the electron cloud of quinoline and the aromatic ring of Tyr265.
The complex D/6VWW established moderately favourable hydrogen bonding interactions with the residues Glu234 (38% MD trajectory), Asp240 (35% MD trajectory), Cys334 (23% MD trajectory), Glu340 (11% MD trajectory) and Pro344 (10% MD trajectory). Furthermore, the complex was stabilized by forming π-π stacking and π-cationic interactions with His234, His235 and Lys335, respectively.
The ligand properties of the compound D (Supplementary Fig. S3a–e) with respect to the selected non-structural proteins revealed that the compound D established an RMSD of 0.75–2.83 Å (6LU7), 0.41–1.93 Å (6W9C), 0.66–0.69 Å (6M71), 0.71–1.96 Å (6XEZ) and 0.81–2.79 Å (6VWW), indicating less conformational changes and increased stability of compound D during the MD study. Furthermore, the radius of gyration was found to be in the range of 5.04–5.71 Å (6LU7), 4.83–5.81 Å (6W9C), 5.51–5.66 Å (6M71), 5.36–5.76 Å (6XEZ) and 4.83–5.81 Å (6VWW). In addition, polar surface area, solvent accessible surface area and molar surface area values indicated the stabilization of MD study.
MM-GBSA was performed for the MD complexes D/6LU7, D/6W9C, D/6M71, D/6XEZ and D/6VWW, and the results are depicted in Table 6 and Supplementary Fig. S4a–e. The complex D exhibited significant binding energy with the selected non-structural proteins in the range of −33.28 to −67.66 kcal/mol. ΔGcou energy term was found to be favourable for 6LU7, 6W9C and 6XEZ and unfavourable against 6M71 and 6VWW. Besides, ΔGcov and ΔGlip were found to be moderately favourable for the selected non-structural proteins. From the results, it is clear that ΔGvdW (−30.94 to −81.10 kcal/mol) and ΔGb were found to be the key driving forces for the inhibitory action of compound D against the selected non-structural proteins.
ΔGb is the binding free energy, ΔGcou is the coulombic interaction energy, ΔGcov is the covalent energy, ΔGlip is the lipophilic-solvation energy and ΔGvdW is the van der Waals energy
Conclusion
In summary, HTVS, SP docking, XP docking and binding free energy calculation studies were performed using the Comprehensive Marine Natural Products Database against five non-structural proteins: Nsp3, Nsp5, Nsp12, Nsp13 and Nsp15. Amongst the top hits, CMNPD5804, CMNPD20924 and CMNPD1598 compounds were utilized to design a novel molecule (D) which has the capability of interacting with all the key residues in the pocket of the selected non-structural proteins. The compound D exhibited greater binding affinity with the values of Gscore (−7.234 kcal/mol) and Gbind (−90.562 kcal/mol) for Nsp3, Gscore (−7.191 kcal/mol) and Gbind (−67.119 kcal/mol) for Nsp5, Gscore (−6.514 kcal/mol) and Gbind (−90.478 kcal/mol) for Nsp12, Gscore (−7.331 kcal/mol) and Gbind (−119.69 kcal/mol) for Nsp13 and Gscore (−8.274kcal/mol) and Gbind (−67.771kcal/mol) for Nsp15. Furthermore, MDS studies for the complexes D/6LU7, D/6W9C, D/6M71, D/6XEZ and D/6VWW revealed the greater stability of the compound D within the binding pocket of the selected non-structural proteins. These results indicated that compound D could be a promising inhibitor against these non-structural proteins. However, still the study requires further in vitro and in vivo studies to support our findings.
Data availability
Not applicable.
Code availability
Not applicable.
References
Hui DS, Azhar EI, Madani TA, Ntoumi F, Kock R, Dar O et al (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 91:264–266
World Health Organization (2020) Addressing human rights as key to the COVID-19: response. World Health Organization. https://apps.who.int/iris/bitstream/handle/10665/331811/WHO-2019-nCoV-SRH-Rights-2020.1-eng.pdf. Accessed 11 May 2023
Anjorin AA (2020) The coronavirus disease 2019 (COVID-19) pandemic: a review and an update on cases in Africa. Asian Pac J Trop Med 13(5):199
Yoshimoto FK (2020) The proteins of severe acute respiratory syndrome coronavirus-2 (SARS CoV-2 or n-COV19), the cause of COVID-19. Protein J 39(3):198–216
Alazmi M, Motwalli O (2021) In silico virtual screening, characterization, docking and molecular dynamics studies of crucial SARS-CoV-2 proteins. J Biomol Struct Dyn 39(17):6761–6771
Báez-Santos YM, John SES, Mesecar AD (2015) The SARS-coronavirus papain-like protease: structure, function and inhibition by designed antiviral compounds. Antiviral Res 115:21–38
Jin Z, Du X, Xu Y, Deng Y, Liu M, Zhao Y et al (2020) Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 582(7811):289–293
Osipiuk J, Azizi S-A, Dvorkin S, Endres M, Jedrzejczak R, Jones KA et al (2021) Structure of papain-like protease from SARS-CoV-2 and its complexes with non-covalent inhibitors. Nat Commun 12(1):1–9
Wu C, Liu Y, Yang Y, Zhang P, Zhong W, Wang Y et al (2020) Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharmaceutica Sinica B 10(5):766–788
Gao Y, Yan L, Huang Y, Liu F, Zhao Y, Cao L et al (2020) Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 368(6492):779–782
Snijder E, Decroly E, Ziebuhr J (2016) The nonstructural proteins directing coronavirus RNA synthesis and processing. Adv Virus Res 96:59–126
Jang K-J, Jeong S, Kang DY, Sp N, Yang YM, Kim D-E (2020) A high ATP concentration enhances the cooperative translocation of the SARS coronavirus helicase nsP13 in the unwinding of duplex RNA. Sci Rep 10(1):1–13
Chen J, Malone B, Llewellyn E, Grasso M, Shelton PM, Olinares PDB et al (2020) Structural basis for helicase-polymerase coupling in the SARS-CoV-2 replication-transcription complex. Cell 182(6):1560–1573.e13
Deng X, Hackbart M, Mettelman RC, O’Brien A, Mielech AM, Yi G et al (2017) Coronavirus nonstructural protein 15 mediates evasion of dsRNA sensors and limits apoptosis in macrophages. Proc Natl Acad Sci 114(21):E4251–E4E60
Kim Y, Jedrzejczak R, Maltseva NI, Wilamowski M, Endres M, Godzik A et al (2020) Crystal structure of Nsp15 endoribonuclease NendoU from SARS-CoV-2. Protein Sci 29(7):1596–1605
Muhammed Y (2020) Molecular targets for COVID-19 drug development: enlightening Nigerians about the pandemic and future treatment. Biosafety and Health 2(04):210–216
McKee DL, Sternberg A, Stange U, Laufer S, Naujokat C (2020) Candidate drugs against SARS-CoV-2 and COVID-19. Pharmacol Res 157:104859
Khan Z, Karataş Y, Rahman H (2020) Anti COVID-19 drugs: need for more clinical evidence and global action. Adv Ther 37(6):2575–2579
Aggarwal G, Henry BM, Aggarwal S, Bangalore S (2020) Cardiovascular safety of potential drugs for the treatment of coronavirus disease 2019. Am J Cardiol 128:147–150
Javorac D, Grahovac L, Manić L, Stojilković N, Anđelković M, Bulat Z et al (2020) An overview of the safety assessment of medicines currently used in the COVID-19 disease treatment. Food Chem Toxicol 144:111639
Lokhande KB, Ballav S, Yadav RS, Swamy KV, Basu S (2020) Probing intermolecular interactions and binding stability of kaempferol, quercetin and resveratrol derivatives with PPAR-γ: docking, molecular dynamics and MM/GBSA approach to reveal potent PPAR-γ agonist against cancer. J Biomol Struct Dyn 40(3):971–981
Lokhande KB, Ballav S, Thosar N, Swamy KV, Basu S (2020) Exploring conformational changes of PPAR-Ɣ complexed with novel kaempferol, quercetin, and resveratrol derivatives to understand binding mode assessment: a small-molecule checkmate to cancer therapy. J Mol Model 26(9):1–12
Lokhande KB, Nagar S, Swamy KV (2019) Molecular interaction studies of deguelin and its derivatives with cyclin D1 and cyclin E in cancer cell signaling pathway: the computational approach. Sci Rep 9(1):1–13
Dhankhar P, Dalal V, Singh V, Tomar S, Kumar P (2022) Computational guided identification of novel potent inhibitors of N-terminal domain of nucleocapsid protein of severe acute respiratory syndrome coronavirus 2. J Biomol Struct Dyn 40(9):4084–4099
Kumar KA, Sharma M, Dalal V, Singh V, Tomar S, Kumar P (2021) Multifunctional inhibitors of SARS-CoV-2 by MM/PBSA, essential dynamics, and molecular dynamic investigations. J Mol Graph Model 107:107969
Dhankhar P, Dalal V, Kumar V (2021) Screening of severe acute respiratory syndrome coronavirus 2 RNA-dependent RNA polymerase inhibitors using computational approach. J Comput Biol 28(12):1228–1247
Kumari R, Kumar V, Dhankhar P, Dalal V (2022) Promising antivirals for PLpro of SARS-CoV-2 using virtual screening, molecular docking, dynamics, and MMPBSA. J Biomol Struct Dyn:1–17
Iwaloye O, Elekofehinti OO, Momoh AI, Babatomiwa K, Ariyo EO (2020) In silico molecular studies of natural compounds as possible anti-Alzheimer’s agents: ligand-based design. Network Modeling Analysis in Health Informatics and Bioinformatics 9(1):1–14
Iwaloye O, Elekofehinti OO, Oluwarotimi EA, Fadipe TM (2020) Insight into glycogen synthase kinase-3β inhibitory activity of phyto-constituents from Melissa officinalis: in silico studies. In Silico Pharmacology 8(1):1–13
Jacobson MP, Pincus DL, Rapp CS, Day TJ, Honig B, Shaw DE et al (2004) A hierarchical approach to all-atom protein loop prediction. Proteins: Structure, Function, and Bioinformatics 55(2):351–367
Roos K, Wu C, Damm W, Reboul M, Stevenson JM, Lu C et al (2019) OPLS3e: extending force field coverage for drug-like small molecules. J Chem Theory Comput 15(3):1863–1874
Yang H, Sun L, Wang Z, Li W, Liu G, Tang Y (2018) ADMETopt: a web server for ADMET optimization in drug design via scaffold hopping. J Chem Inf Model 58(10):2051–2056
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47(7):1739–1749
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA et al (2006) Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein–ligand complexes. J Med Chem 49(21):6177–6196
Halgren TA, Murphy RB, Friesner RA, Beard HS, Frye LL, Pollard WT et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem 47(7):1750–1759
Sastry GM, Adzhigirey M, Day T, Annabhimoju R, Sherman W (2013) Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Comput Aided Mol Des 27(3):221–234
Shukla R, Singh TR (2019) Virtual screening, pharmacokinetics, molecular dynamics and binding free energy analysis for small natural molecules against cyclin-dependent kinase 5 for Alzheimer’s disease. J Biomol Struct Dyn 38(1):248–262
Duffy EM, Jorgensen WL (2000) Prediction of properties from simulations: free energies of solvation in hexadecane, octanol, and water. J Am Chem Soc 122(12):2878–2888
Lawrence C, Skinner J (2003) Flexible TIP4P model for molecular dynamics simulation of liquid water. Chem Phys Lett 372(5-6):842–847
Jorgensen WL, Madura JD (1985) Temperature and size dependence for Monte Carlo simulations of TIP4P water. Mol Phys 56(6):1381–1392
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995) A smooth particle mesh Ewald method. J Chem Phys 103(19):8577–8593
Martyna GJ, Klein ML, Tuckerman M (1992) Nosé–Hoover chains: the canonical ensemble via continuous dynamics. J Chem Phys 97(4):2635–2643
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101(5):4177–4189
Martyna GJ, Tuckerman ME, Tobias DJ, Klein ML (1996) Explicit reversible integrators for extended systems dynamics. Mol Phys 87(5):1117–1157
Acknowledgements
The authors thank Lalji Baldaniya for his continuous support throughout the study.
Author information
Authors and Affiliations
Contributions
Simran Patel: writing of original draft and data curation; Haydara Hasan: writing of original draft and methodology; Divyesh Umraliya: formal analysis; Bharat Kumar Reddy Sanapalli: writing including reviewing and editing, and conceptualization; Vidyasrilekha Yele: writing including reviewing and editing, software, supervision and conceptualization.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 3.96 MB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patel, S., Hasan, H., Umraliya, D. et al. Marine drugs as putative inhibitors against non-structural proteins of SARS-CoV-2: an in silico study. J Mol Model 29, 176 (2023). https://doi.org/10.1007/s00894-023-05574-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-023-05574-9