Abstract
Epidermal growth factor receptor (EGFR) is highlighted as a target for anticancer treatment. Several EGFR inhibitors were approved in cancer treatment. Comparatively, 5D-QSAR is a new methodology which considers an ensemble of different induced-fit models. Based on 1H-pyrazole derivatives as EGFR inhibitors, a 5D-QSAR was studied in which the method of quasi-atomistic receptor surface modeling was used. The presented QSAR model showed contributions of the hydrogen bond acceptor, and hydrophobic and salt bridge fields to the activity. The QSAR model was statistically validated and also externally validated applying 19 compounds (test set) which were not included in the model generation process. The scramble tests were performed to further verify the robustness. Apart from exploration of the binding of 1H-pyrazole derivatives to the EGFR, the 5D-QSAR model can be helpful to design of new EGFR inhibitors.
Graphical Abstract
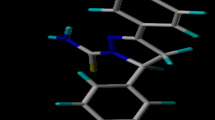
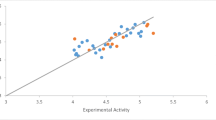
The five-dimensional quantitative structure–activity relationship (5D-QSAR) of 1H-pyrazole derivatives as EGFR inhibitors with quasi-atomistic receptor surface modeling approach is described.







Similar content being viewed by others
Data availability
All data concerning the results of this study is shown in this manuscript, and raw material may be sent on request.
Code availability
All the software used in the study are available online for use. No additional coding is done.
References
He Y, Harrington BS, Hooper JD (2016) New crossroads for potential therapeutic intervention in cancer-intersections between CDCP1, EGFR family members and downstream signaling pathways. Oncoscience 3:5–8
Yarden Y, Sliwkowski MX (2001) Untangling the ErbB signalling network. Nat rev Mol Cell Bio 2:127–137
Hsu JL, Hung MC (2016) The role of HER2, EGFR, and other receptor tyrosine kinases in breast cancer. Cancer Metastasis Rev 35:575–588
Huo LF, Wang YN, Xia WY, Hsu SC, Lai CC, Li LY, Chang WC, Wang Y, Hsu MC, Yu YL (2010) RNA helicase A is a DNA-binding partner for EGFR-mediated transcriptional activation in the nucleus. Proc Natl Acad ScI USA 107:16125–16130
Xu MJ, Johnson DE, Grandis JR (2017) EGFR-targeted therapies in the post-genomic era. Cancer Metastasis Rev 36:463–473
Hossam M, Lasheen DS, Abouzid KAM (2016) Covalent EGFR inhibitors: binding mechanisms, synthetic approaches, and clinical profiles. Arch Pharm 349:573–593
Liu F, Tang B, Liu H, Li L, Liu G, Cheng Y, Xu Y, Chen WW, Huang Y (2016) 4-Anilinoquinazoline derivatives with epidermal growth factor receptor inhibitor activity. Anti-Cancer Agents in Med Chem 16:1652–1664
Burotto M, Chiou VL, Lee JM, Kohn EC (2014) The MAPK pathway across different malignancies: a new perspective. Cancer 120:3446–3456
Kubinyi H (1997) QSAR and 3D QSAR in drug design Part 2: applications and problems. Drug Discov Today 2:538–546
Kubinyi H (1997) QSAR and 3D QSAR in drug design Part 1: methodology. Drug Discov Today 2:457–467
Vedani A, Dobler M (2002) Multidimensional QSAR: moving from three- to five-dimensional concepts. Quant Struct-Act Relat 21:382–390
Roy K, Das RN (2014) A review on principles, theory and practices of 2D-QSAR. Curr Drug Metab 15:346–379
Damale MG, Harke SN, Kalam Khan FA, Shinde DB, Sangshetti JN (2014) Recent advances in multidimensional QSAR (4D–6D): a critical review. Mini Rev Med Chem 14:35–55
Vedani A, Dobler M, Lill MA (2005) Combining protein modeling and 6D-QSAR. Simulating the binding of structurally diverse ligands to the estrogen receptor. J Med Chem 48:3700–3703
Oberdorf C, Schmidt TJ, Wunsch B (2010) 5D-QSAR for spirocyclic sigma1 receptor ligands by Quasar receptor surface modeling. Eur J Med Chem 45:3116–3124
Roy K, Kar S, Das RN (2015) Understanding the basics of QSAR for applications in pharmaceutical sciences and risk assesment. Newer QSAR techniques. Academic Press, Elsevier, pp 319–356
Lv PC, Li HQ, Sun J, Zhou Y, Zhu HL (2010) Synthesis and biological evaluation of pyrazole derivatives containing thiourea skeleton as anticancer agents. Biorg Med Chem 18:4606–4614
Hassinen T, Perkyl M (2001) New energy terms for reduced protein models implemented in an off-lattice force field. J Comput Chem 22:1229–1242
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera—a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Patil R, Sawant S (2015) Molecular dynamics guided receptor independent 4D QSAR studies of substituted coumarins as anticancer agents. Curr Comput Aided Drug Des 11:39–50
Berendsen HJC, Postma JV, van Gunsteren WF, DiNola A, Haak JR (1984) Molecular dynamics with coupling to an external bath. J Chem Phys 81:3684–3690
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N⋅log (N) method for Ewald sums in large systems. J Chem Phys 98:10089–10092
Vedani A, Dobler M. User and reference manual Quasar 5.0. Biograf Laboratory 3R, Basel, Switzerland,
Pourbasheer E, Aalizadeh R, Shiri HM, Banaei A, Ganjali MR (2015) 2D and 3D-QSAR analysis of pyrazole-thiazolinone derivatives as EGFR kinase by COMFA and COMSIA. Curr Comput Aided Drug Des 11:292–303
Funding
The project was supported by the Undergraduate Innovation and Entrepreneurship Foundation (2021CX242, 2020CX289), the Nanchang University Teaching Reform Foundation (NCUJGLX-19–130, NCUJGLX-19–124), and the Jiangxi Province Science Foundation (20171BAB205104).
Author information
Authors and Affiliations
Contributions
DQ: methods, writing and result analysis. XZ: problem selection, writing and result analysis. TZ: data analysis. BC: data analysis. BY: data analysis. GT: methods, project management, result analysis, manuscript editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Qin, D., Zeng, X., Zhao, T. et al. 5D-QSAR studies of 1H-pyrazole derivatives as EGFR inhibitors. J Mol Model 28, 379 (2022). https://doi.org/10.1007/s00894-022-05370-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-022-05370-x