Abstract
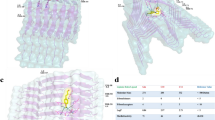
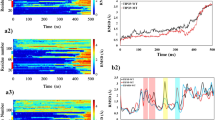
The α-synuclein fibrils are a pathological hallmark of Parkinson’s disease (PD) and are abundant in the brains of PD patients. These amyloid fibrils can aggregate into distinct polymorphism under different physical conditions. Therefore, these different fibril polymorph formations should be considered in drug design studies targeting amyloid fibrils. Recently, the atomic structures of two small fibril segments of α-synuclein, named NACore (68–78) and SubNACore (69–77), have been crystallized. These segments are critical for cytotoxicity and fibril formation. Therefore, elucidation of interface interactions between pair sheets of the NACore and SubNACore is significant for the clarification of the mechanism of fibril formation in PD. In this context, molecular dynamics (MD) simulation technique is a convenient tool to investigate interface interactions of these segments at the atomic level. However, the accuracy of these simulations depends on the utilized force fields. Therefore, we have tested the dependence of interface interactions and stabilities of these small amyloid fibrils on various force fields. From the results of triple long (100 ns) MD simulations, we inferred for the stability investigations of the NACore and SubNACore that CHARMM27 and GROMOS53A6 are the most convenient force fields whereas AMBER99SB-ILDN is the most unfavorable one. Consequently, it is expected that our findings will guide the selection of the appropriate force field for simulations between these segments and possible inhibitors of this disease.






Similar content being viewed by others
References
Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R (1997). Science 276:2045
Singleton A, Farrer M, Johnson J, Singleton A, Hague S, Kachergus J, Hulihan M, Peuralinna T, Dutra A, Nussbaum R (2003). Science 302:841
Nasica-Labouze J, Nguyen PH, Sterpone F, Berthoumieu O, Buchete N-V, Cote S, De Simone A, Doig AJ, Faller P, Garcia A (2015). Chem Rev 115:3518
Bedrood S, Li Y, Isas JM, Hegde BG, Baxa U, Haworth IS, Langen R (2012). J Biol Chem 287:5235
Anguiano M, Nowak RJ, Lansbury PT (2002). Biochemistry 41:11338
Scherzinger E, Sittler A, Schweiger K, Heiser V, Lurz R, Hasenbank R, Bates GP, Lehrach H, Wanker EE (1999). Proc Natl Acad Sci 96:4604
Spillantini MG, Schmidt ML, M-Y Lee V, Trojanowski JQ, Jakes R, Goedert M (1997). Nature 388:839
Xu L, Nussinov R, Ma B (2016). Eur J Med Chem 121:841
Rodriguez JA, Ivanova MI, Sawaya MR, Cascio D, Reyes FE, Shi D, Sangwan S, Guenther EL, Johnson LM, Zhang M (2015). Nature 525:486
Bodles AM, Guthrie DJ, Greer B, Irvine GB (2001). J Neurochem 78:384
Bisaglia M, Trolio A, Bellanda M, Bergantino E, Bubacco L, Mammi S (2006). Protein Sci 15:1408
Periquet M, Fulga T, Myllykangas L, Schlossmacher MG, Feany MB (2007). J Neurosci 27:3338
Volles MJ, Lansbury PT (2007). J Mol Biol 366:1510
Vamvaca K, Volles MJ, Lansbury Jr PT (2009). J Mol Biol 389:413
EI-Agnaf O, Irvine G (2002). Biochem Soc T 30:559
Duan Y, Wu C, Chowdhury S, Lee MC, Xiong G, Zhang W, Yang R, Cieplak P, Luo R, Lee T (2003). J Comput Chem 24:1999
Hornak V, Abel R, Okur A, Strockbine B, Roitberg A, Simmerling C (2006). Proteins 65:712
Lindorff-Larsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE (2010). Proteins 78:1950
Best RB, Zhu X, Shim J, Lopes PE, Mittal J, Feig M, MacKerell Jr AD (2012). J Chem Theory Comput 8:3257
MacKerell Jr AD, Bashford D, Bellott M, Dunbrack Jr RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S (1998). J Phys Chem B 102:3586
Oostenbrink C, Villa A, Mark AE, Van Gunsteren WF (2004). J Comput Chem 25:1656
Schmid N, Eichenberger AP, Choutko A, Riniker S, Winger M, Mark AE, van Gunsteren WF (2011). Eur Biophys J 40:843
Kaminski GA, Friesner RA, Tirado-Rives J, Jorgensen WL (2001). J Phys Chem B 105:6474
Pronk S, Páll S, Schulz R, Larsson P, Bjelkmar P, Apostolov R, Shirts MR, Smith JC, Kasson PM, Van Der Spoel D (2013). Bioinformatics 29:845
Berendsen HJ, Postma JP, van Gunsteren WF, Hermans J (1981) Intermolecular Forces. Springer, Dordrecht, p 331
Jorgensen WL, Chandrasekhar J, Madura JD, Impey RW, Klein ML (1983). J Chem Phys 79:926
Darden T, York D, Pedersen L (1993). J Chem Phys 98:10089
Essmann U, Perera L, Berkowitz ML, Darden T, Lee H, Pedersen LG (1995). J Chem Phys 103:8577
Hess B, Bekker H, Berendsen HJ, Fraaije JG (1997). J Comput Chem 18:1463
Miyamoto S, Kollman PA (1992). J Comput Chem 13:952
Bussi G, Donadio D, Parrinello M (2007). J Chem Phys 126:014101
Parrinello M, Rahman A (1981). J Appl Phys 52:7182
Kumari R, Kumar R, Consortium OSDD, Lynn A (2014). J Chem Inf Model 54:1951
Alred EJ, Scheele EG, Berhanu WM, Hansmann UH (2014). J Chem Phys 141:175101
Berhanu WM, Hansmann UH (2013). Proteins 81:1542
Spiliotopoulos D, Spitaleri A, Musco G (2012). PLoS One 7:e46902
DeLano WL (2002) The PyMol Molecular Graphics System. De Lano Scientific L.L.C., San Carlos, http://pymol.org
Carballo-Pacheco M, Strodel B (2017). Protein Sci 26:174
Funding
This work was supported by the Scientific Research Fund of Zonguldak Bülent Ecevit University with project number 2015-22794455-03, and the numerical calculations reported in this paper were partially performed at TUBITAK ULAKBIM, High Performance and Grid Computing Center (TRUBA resources).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 148 kb).
Rights and permissions
About this article
Cite this article
Alıcı, H. Structural analyses and force fields comparison for NACore (68–78) and SubNACore (69–77) fibril segments of Parkinson’s disease. J Mol Model 26, 132 (2020). https://doi.org/10.1007/s00894-020-04379-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-020-04379-4