Abstract
The structure of a hairpin loop—in particular its large accessible surface area and its exposed hydrogen-bonding edges—facilitate an inherent possibility for interactions. Just like higher-order RNA macromolecules, pre-microRNAs possess a hairpin loop, and it plays a crucial role in miRNA biogenesis. Upon inspecting the crystal structures of RNAs with various functions, we noticed that, along with a fairly long double helix, the RNAs contained sequentially different hairpin loops comprising four residues. We therefore applied molecular dynamics simulation to analyze six of these previously unexplored tetraloops, along with GNRA (where N is any nucleotide and R is a purine nucleotide) tetraloops, to understand their structural and functional characteristics. A number of analyses quantifying loop stability by examining base–base stacking, base–sugar and base–phosphate hydrogen bonding, and backbone variability were performed. Importantly, we determined the different interbase stacking preferences of the single-stranded unpaired bases of the hairpin loops, which had not previously been quantified in any form. Furthermore, our study indicates that canonical GNRA structural properties are exhibited by some structures containing non-GNRA loop sequences.

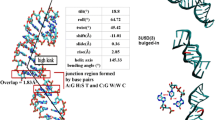
Stacking overlap at loop region




Similar content being viewed by others
References
Mattick JS, Makunin IV (2006) Non-coding RNA. Hum Mol Genet 15:R17–R29. https://doi.org/10.1093/hmg/ddl046
Kaikkonen MU, Lam MTY, Glass CK (2011) Non-coding RNAs as regulators of gene expression and epigenetics. Cardiovasc Res 90:430–440. https://doi.org/10.1093/cvr/cvr097
Phillips T (2014) Small non-coding RNA and gene expression. Nat Educ 1:115
Hendrix DK, Brenner SE, Holbrook SR (2005) RNA structural motifs: building blocks of a modular biomolecule. Q Rev Biophys 38:221–243. https://doi.org/10.1017/S0033583506004215
Gutell RR (2013) Comparative analysis of the higher-order structure of RNA. In: Russell R (ed) Biophysics of RNA folding. Springer, New York, pp 11–22
Petrov AI, Zirbel CL, Leontis NB (2013) Automated classification of RNA 3D motifs and the RNA 3D motif atlas. RNA 19:1327–1340. https://doi.org/10.1261/rna.039438.113
Mortimer SA, Kidwell MA, Doudna JA (2014) Insights into RNA structure and function from genome-wide studies. Nat Rev Genet 15:469–479. https://doi.org/10.1038/nrg3681
Weeks KM (2015) Review toward all RNA structures, concisely. Biopolymers 103:438–448. https://doi.org/10.1002/bip.22601
Parlea LG, Sweeney BA, Hosseini-Asanjan M, et al. (2016) The RNA 3D motif atlas: computational methods for extraction, organization and evaluation of RNA motifs. Methods 103:99–119. https://doi.org/10.1016/j.ymeth.2016.04.025
Cheong C, Cheong H-K (2015) RNA structure: tetraloops. eLS. https://doi.org/10.1002/9780470015902.a0003135.pub2
Woese CR, Winker S, Gutell RR (1990) Architecture of ribosomal RNA: constraints on the sequence of “tetra-loops”. Proc Natl Acad Sci USA 87:8467–8471. https://doi.org/10.1073/pnas.87.21.8467
Groebe DR, Uhlenbeck OC (1988) Characterization of RNA hairpin loop stability. Nucleic Acids Res 16:11725–11735
Kuznetsov SV, Ren C-C, Woodson SA, Ansari A (2008) Loop dependence of the stability and dynamics of nucleic acid hairpins. Nucleic Acids Res 36:1098–1112. https://doi.org/10.1093/nar/gkm1083
Zeng Y, Yi R, Cullen BR (2005) Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J 24:138–148. https://doi.org/10.1038/sj.emboj.7600491
Lund E, Dahlberg JE (2006) Substrate selectivity of exportin 5 and Dicer in the biogenesis of microRNAs. Cold Spring Harb Symp Quant Biol 71:59–66. https://doi.org/10.1101/sqb.2006.71.050
Michlewski G, Guil S, Semple CA, Cáceres JF (2008) Posttranscriptional regulation of miRNAs harboring conserved terminal loops. Mol Cell 32:383–393. https://doi.org/10.1016/j.molcel.2008.10.013
Mirihana Arachchilage G, Dassanayake AC, Basu S (2015) A potassium ion-dependent RNA structural switch regulates human pre-miRNA 92b maturation. Chem Biol 22:262–272. https://doi.org/10.1016/j.chembiol.2014.12.013
Bikard D, Loot C, Baharoglu Z, Mazel D (2010) Folded DNA in action: hairpin formation and biological functions in prokaryotes. Microbiol Mol Biol Rev 74:570–588. https://doi.org/10.1128/MMBR.00026-10
Lee JC, Gutell RR (2014) Helix capping in RNA structure. PLoS One 9:e93664. https://doi.org/10.1371/journal.pone.0093664
Fujita Y, Tanaka T, Furuta H, Ikawa Y (2012) Functional roles of a tetraloop/receptor interacting module in a cyclic di-GMP riboswitch. J Biosci Bioeng 113:141–145. https://doi.org/10.1016/j.jbiosc.2011.10.004
Fiore JL, Nesbitt DJ (2013) An RNA folding motif: GNRA tetraloop–receptor interactions. Q Rev Biophys 46:223–264. https://doi.org/10.1017/S0033583513000048
Thapar R, Denmon AP, Nikonowicz EP (2014) Recognition modes of RNA tetraloops and tetraloop-like motifs by RNA-binding proteins. Wiley Interdisc Rev RNA 5:49–67. https://doi.org/10.1002/wrna.1196
Kralovicova J, Patel A, Searle M, Vorechovsky I (2015) The role of short RNA loops in recognition of a single-hairpin exon derived from a mammalian-wide interspersed repeat. RNA Biol 12:54–69. https://doi.org/10.1080/15476286.2015.1017207
Uhlenbeck OC (1990) Tetraloops and RNA folding. Nature 346:613–614. https://doi.org/10.1038/346613a0
Bevilacqua PC, Blose JM (2008) Structures, kinetics, thermodynamics, and biological functions of RNA hairpins. Annu Rev Phys Chem 59:79–103. https://doi.org/10.1146/annurev.physchem.59.032607.093743
Banáš P, Hollas D, Zgarbová M, et al. (2010) Performance of molecular mechanics force fields for RNA simulations: stability of UUCG and GNRA hairpins. J Chem Theory Comput 6:3836–3849
Aviv T, Lin Z, Ben-Ari G, et al. (2006) Sequence-specific recognition of RNA hairpins by the SAM domain of Vts1p. Nat Struct Mol Biol 13:168–176. https://doi.org/10.1038/nsmb1053
Sahu B, Khade PK, Joseph S (2012) Functional replacement of two highly conserved tetraloops in the bacterial ribosome. Biochemistry 51:7618–7626. https://doi.org/10.1021/bi300930r
Deng N-JJ, Cieplak P (2010) Free energy profile of RNA hairpins: a molecular dynamics simulation study. Biophys J 98:627–636. https://doi.org/10.1016/j.bpj.2009.10.040
Chakraborty D, Collepardo-Guevara R, Wales DJ (2014) Energy landscapes, folding mechanisms, and kinetics of RNA tetraloop hairpins. J Am Chem Soc 136:18052–18061. https://doi.org/10.1021/ja5100756
Miner JC, Chen AA, García AE (2016) Free-energy landscape of a hyperstable RNA tetraloop. Proc Natl Acad Sci USA 113:6665–6670. https://doi.org/10.1073/pnas.1603154113
Nivón LG, Shakhnovich EI (2004) All-atom Monte Carlo simulation of GCAA RNA folding. J Mol Biol 344:29–45. https://doi.org/10.1016/j.jmb.2004.09.041
Chen AA, Garcia AE (2013) High-resolution reversible folding of hyperstable RNA tetraloops using molecular dynamics simulations. Proc Natl Acad Sci USA 110:16820–16825. https://doi.org/10.1073/pnas.1309392110
Bisaria N, Herschlag D (2015) Probing the kinetic and thermodynamic consequences of the tetraloop/tetraloop receptor monovalent ion-binding site in P4–P6 RNA by smFRET. Biochem Soc Trans 43:172–178. https://doi.org/10.1042/BST20140268
Furukawa A, Maejima T, Matsumura S, Ikawa Y (2016) Characterization of an RNA receptor motif that recognizes a GCGA tetraloop. Biosci Biotechnol Biochem 80:1386–9. doi:https://doi.org/10.1080/09168451.2016.1156483
Kozomara A, Griffiths-Jones S (2014) miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res 42:D68–D73. https://doi.org/10.1093/nar/gkt1181
Castilla-Llorente V, Nicastro G, Ramos A (2013) Terminal loop-mediated regulation of miRNA biogenesis: selectivity and mechanisms. Biochem Soc Trans 41:861–865. https://doi.org/10.1042/BST20130058
Kundu S, Mukherjee S, Bhattacharyya D (2012) Effect of temperature on DNA double helix: an insight from molecular dynamics simulation. J Biosci 37:445–455. https://doi.org/10.1007/s12038-012-9215-5
Mukherjee S, Bhattacharyya D (2013) Influence of divalent magnesium ion on DNA: molecular dynamics simulation studies. J Biomol Struct Dyn 31:896–912. https://doi.org/10.1080/07391102.2012.713780
Ben Imeddourene A, Xu X, Zargarian L, et al. (2016) The intrinsic mechanics of B-DNA in solution characterized by NMR. Nucleic Acids Res 44:3432–3447. https://doi.org/10.1093/nar/gkw084
Colizzi F, Bussi G (2012) RNA unwinding from reweighted pulling simulations. J Am Chem Soc 134:5173–5179. https://doi.org/10.1021/ja210531q
Zgarbová M, Otyepka M, Šponer J, et al. (2014) Base pair fraying in molecular dynamics simulations of DNA and RNA. J Chem Theory Comput 10:3177–3189. https://doi.org/10.1021/ct500120v
Xu X, Yu T, Chen S-J (2016) Understanding the kinetic mechanism of RNA single base pair formation. Proc Natl Acad Sci USA 113:116–121. https://doi.org/10.1073/pnas.1517511113
Melchers WJG, Zoll J, Tessari M, et al. (2006) A GCUA tetranucleotide loop found in the poliovirus oriL by in vivo SELEX (un)expectedly forms a YNMG-like structure: extending the YNMG family with GYYA. RNA 12:1671–1682. https://doi.org/10.1261/rna.113106
Du Z, Yu J, Andino R, James TL (2003) Extending the family of UNCG-like tetraloop motifs: NMR structure of a CACG tetraloop from coxsackievirus B3. Biochemistry 42:4373–4383. https://doi.org/10.1021/bi027314e
Bottaro S, Lindorff-Larsen K (2017) Mapping the universe of RNA tetraloop folds. Biophys J 113:257–267. https://doi.org/10.1016/j.bpj.2017.06.011
Pingali PK, Halder S, Mukherjee D, et al. (2014) Analysis of stacking overlap in nucleic acid structures: algorithm and application. J Comput Aided Mol Des 28:851–867. https://doi.org/10.1007/s10822-014-9767-6
Das J, Mukherjee S, Mitra A, Bhattacharyya D (2006) Non-canonical base pairs and higher order structures in nucleic acids: crystal structure database analysis. J Biomol Struct Dyn 24:149–161. https://doi.org/10.1080/07391102.2006.10507108
Ray SS, Halder S, Kaypee S, Bhattacharyya D (2012) HD-RNAS: an automated hierarchical database of RNA structures. Front Genet 3:59. https://doi.org/10.3389/fgene.2012.00059
Kührová P, Best RB, Bottaro S, et al (2016) Computer folding of RNA tetraloops: identification of key force field deficiencies. J Chem Theory Comput 12:4534–4548. doi:https://doi.org/10.1021/acs.jctc.6b00300
Allnér O, Nilsson L, Villa A (2013) Loop–loop interaction in an adenine-sensing riboswitch: a molecular dynamics study. RNA 19:916–926. https://doi.org/10.1261/rna.037549.112.1
Ochieng PO, White NA, Feig M, Hoogstraten CG (2016) Intrinsic base-pair rearrangement in the hairpin ribozyme directs RNA conformational sampling and tertiary interface formation. J Phys Chem B 120:10885–10898. https://doi.org/10.1021/acs.jpcb.6b05606
Halder S, Bhattacharyya D (2012) Structural variations of single and tandem mismatches in RNA duplexes: a joint MD simulation and crystal structure database analysis. J Phys Chem B 116:11845–11856. https://doi.org/10.1021/jp305628v
Bergonzo C, Henriksen NM, Roe DR, Cheatham TE (2015) Highly sampled tetranucleotide and tetraloop motifs enable evaluation of common RNA force fields. RNA 21:1578–1590. https://doi.org/10.1261/rna.051102.115
MacKerell AD, Banavali NK (2000) All-atom empirical force field for nucleic acids: II. Application to molecular dynamics simulations of DNA and RNA in solution. J Comput Chem 21:105–120. https://doi.org/10.1002/(SICI)1096-987X(20000130)21:2<105::AID-JCC3>3.0.CO;2-P
Denning EJ, Priyakumar UD, Nilsson L, Mackerell AD (2011) Impact of 2′-hydroxyl sampling on the conformational properties of RNA: update of the CHARMM all-atom additive force field for RNA. J Comput Chem 32:1929–1943. https://doi.org/10.1002/jcc.21777
Huang J, MacKerell AD (2013) CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J Comput Chem 34:2135–2145. https://doi.org/10.1002/jcc.23354
Pérez A, Marchán I, Svozil D, et al. (2007) Refinement of the AMBER force field for nucleic acids: improving the description of alpha/gamma conformers. Biophys J 92:3817–3829. https://doi.org/10.1529/biophysj.106.097782
Zgarbová M, Otyepka M, Šponer J, et al. (2011) Refinement of the Cornell et al. nucleic acids force field based on reference quantum chemical calculations of glycosidic torsion profiles. J Chem Theory Comput 7:2886–2902. https://doi.org/10.1021/ct200162x
Berman HM, Olson WK, Beveridge DL, et al. (1992) The nucleic acid database. A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys J 63:751–759. https://doi.org/10.1016/S0006-3495(92)81649-1
Jorgensen WL, Chandrasekhar J, Madura JD, et al. (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926. https://doi.org/10.1063/1.445869
Kalé L, Skeel R, Bhandarkar M, et al. (1999) NAMD2: greater scalability for parallel molecular dynamics. J Comput Phys 151:283–312. https://doi.org/10.1006/jcph.1999.6201
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N·log(N) method for Ewald sums in large systems. J Chem Phys 98:10089. https://doi.org/10.1063/1.464397
Brooks BR, Bruccoleri RE, Olafson BD, et al. (1983) CHARMM: a program for macromolecular energy, minimization, and dynamics calculations. J Comput Chem 4:187–217. https://doi.org/10.1002/jcc.540040211
Feller SE, Zhang Y, Pastor RW, Brooks BR (1995) Constant pressure molecular dynamics simulation: the Langevin piston method. J Chem Phys 103:4613. https://doi.org/10.1063/1.470648
Ryckaert JP, Ciccotti G, Berendsen HJC (1977) Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23:327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Abraham MJ, Murtola T, Schulz R, et al. (2015) GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1:19–25. https://doi.org/10.1016/j.softx.2015.06.001
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14:33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Bansal M, Bhattacharyya D, Ravi B (1995) NUPARM and NUCGEN: software for analysis and generation of sequence dependent nucleic acid structures. Bioinformatics 11:281–287. https://doi.org/10.1093/bioinformatics/11.3.281
Mukherjee S, Bansal M, Bhattacharyya D (2006) Conformational specificity of non-canonical base pairs and higher order structures in nucleic acids: crystal structure database analysis. J Comput Aided Mol Des 20:629–645. https://doi.org/10.1007/s10822-006-9083-x
R Development Core Team (2015) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Schrodinger, LLC (2013) The PyMOL molecular graphics system, version 1.6. Schrodinger, LLC, New York
Duarte CM, Pyle AM (1998) Stepping through an RNA structure: a novel approach to conformational analysis. J Mol Biol 284:1465–1478. https://doi.org/10.1006/jmbi.1998.2233
Ray A, Panigrahi S, Bhattacharyya D (2016) A comparison of four different conformations adopted by human telomeric G-quadruplex using computer simulations. Biopolymers 105:83–99. https://doi.org/10.1002/bip.22751
Zhang Y, Zhao X, Mu Y (2009) Conformational transition map of an RNA GCAA tetraloop explored by replica-exchange molecular dynamics simulation. J Chem Theory Comput 5:1146–1154
DePaul AJ, Thompson EJ, Patel SS, et al. (2010) Equilibrium conformational dynamics in an RNA tetraloop from massively parallel molecular dynamics. Nucleic Acids Res 38:4856–4867. https://doi.org/10.1093/nar/gkq134
Condon DE, Kennedy SD, Mort BC, et al. (2015) Stacking in RNA: NMR of four tetramers benchmark molecular dynamics. J Chem Theory Comput 11:2729–2742. https://doi.org/10.1021/ct501025q
Sokoloski JE, Godfrey SA, Dombrowski SE, Bevilacqua PC (2011) Prevalence of syn nucleobases in the active sites of functional RNAs. RNA 17:1775–1787. https://doi.org/10.1261/rna.2759911
Olson WK, Bansal M, Burley SK, et al. (2001) A standard reference frame for the description of nucleic acid base-pair geometry. J Mol Biol 313:229–237. https://doi.org/10.1006/jmbi.2001.4987
Olson WK, Esguerra M, Xin Y, Lu X-J (2009) New information content in RNA base pairing deduced from quantitative analysis of high-resolution structures. Methods 47:177–186. https://doi.org/10.1016/j.ymeth.2008.12.003
Kailasam S, Bhattacharyya D, Bansal M (2014) Sequence dependent variations in RNA duplex are related to non-canonical hydrogen bond interactions in dinucleotide steps. BMC Res Notes 7:83. https://doi.org/10.1186/1756-0500-7-83
Sorin EJ, Engelhardt MA, Herschlag D, Pande VS (2002) RNA simulations: probing hairpin unfolding and the dynamics of a GNRA tetraloop. J Mol Biol 317:493–506. https://doi.org/10.1006/jmbi.2002.5447
Zhang X, Zeng Y (2010) The terminal loop region controls microRNA processing by Drosha and Dicer. Nucleic Acids Res 38:7689–7697. https://doi.org/10.1093/nar/gkq645
Acknowledgements
We would like to thank the Bioinformatics Resources and Applications Facility (BRAF), C-DAC, Pune for providing computation facilities.
Funding
This work was supported by the Department of Atomic Energy, India through the BARD project and the Department of Biotechnology, Govt. of India.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
ESM 1
(PDF 2394 kb)
Rights and permissions
About this article
Cite this article
Mukherjee, D., Bhattacharyya, D. Intrinsic structural variability in GNRA-like tetraloops: insight from molecular dynamics simulation. J Mol Model 23, 300 (2017). https://doi.org/10.1007/s00894-017-3470-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-017-3470-1