Abstract
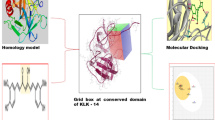
Recent biochemical experiments have revealed that a variety of proteases play important roles in cancer invasion and metastasis. Among these proteases, urokinase-type plasminogen activator (uPA) is particularly important, since its specific binding to the receptor (uPAR) existing on the surface of a cancer cell is considered to be a trigger for cancer invasion. It is thus expected that the blocking of the binding can inhibit cancer invasion in the cancer patients and improve their prognosis dramatically. To develop a potent inhibitor for the binding, many types of peptides of amino acids were produced and their effect on the cancer invasion was investigated in the previous biochemical experiments. On the other hand, our previous ab initio molecular simulations have clarified that some amino acid residues of uPA play important roles in the specific binding between uPA and uPAR. In the present study, we propose some peptides composed of these important residues and investigate the specific interactions and the binding affinity between uPAR and the peptides at an electronic level, using ab initio molecular simulations. Base on the results simulated, we elucidate which peptide can bind more strongly to uPAR and propose a novel potent peptide which can inhibit the binding between uPAR and uPA efficiently.








Similar content being viewed by others
References
Kobayashi H, Gotoh J, Hirashima Y, Fujie M, Sugino D, Terao T (1995) Inhibitory effect of a conjugate between human urokinase and urinary trypsin inhibitor on tumor cell invasion in vitro. J Biol Chem 270:8361–8366
Ruoslahti E (1996) How cancer spreads. Sci Am 9:48–55
Baker EA, Leaper DJ (2003) The plasminogen activator and matrix metalloproteinase systems in colorectal cancer: relationship to tumour pathology. Eur J Cancer 39:981–988
Skelly MM, Troy A, Duffy MJ, Mulcahy HE, Duggan C, Connell TG, O’Donoghue DP, Sheahan K (1997) Urokinase-type plasminogen activator in colorectal cancer: relationship with clinicopathological features and patient outcome. Clin Cancer Res 3:1837–1840
Herszènyi L, Plebani M, Carraro P, De Paoli M, Roveroni G, Cardin R, Tulassay Z, Naccarato R, Farinati F (1999) The role of cysteine and serine proteases in colorectal carcinoma. Cancer 86:1135–1142
Stephens RW, Nielsen HJ, Christensen IJ, Thorlacius-Ussing O, Sorensen S, Dano K, Brunner N (1999) Plasma urokinase receptor levels in patients with colorectal cancer: relationship to prognosis. J Natl Cancer Inst 91:869–874
Nagase K, Kobayashi H, Nagata S, Yamamoto H, Natsume T, Dedachi K, Tsukamoto T, Kurita N (2007) Analysis of cancer invasion mechanism by cellular simulation. J Comp Aid Chem 8:75–84
Kobayashi H, Terao T, Sugino D, Okushima M (1997) Cancerous metastasis inhibitor. US Patent, WO97/25422
Sugiura H, Tsumura N, Kobayashi H, Kurita N (2004) Electronic properties for medicine inhibiting cancer metastasis (1): molecular orbital calculations for physiological substance bikunin. J Comput Aided Chem 5:62–69
Tsumura N, Sugiura H, Kobayashi H, Kurita N (2004) Electronic properties for medicine inhibiting cancer metastasis (2): molecular orbital calculations for urokinase. J Comput Aided Chem 5:70–76
Appella E, Robinson EA, Ullrich SJ, Stoppelli MP, Corti A, Cassani G, Blasi F (1987) The receptor-binding sequence of urokinase. A biological function for the growth-factor module of proteases. J Biol Chem 262:4437–4440
Bdeir K, Kuo A, Sachais BS, Rux AH, Bdeir Y, Mazar A, Higazi AA, Cines DB (2003) The kringle stabilizes urokinase binding to the urokinase receptor. Blood 102:3600–3608
Barinka C, Parry G, Callahan J, Shaw DE, Kuo A, Bdeir K, Cines DB, Mazar A, Lubkowski J (2006) Structural basis of interaction between urokinase-type plasminogen activator and its receptor. J Mol Biol 363:482–495
GmbH RD, Burgle M, Graeff H, Kessler H, Magdolen V, Konig B, Koppitz M, Riemer C, Schmitt M, Weidle U (1998) Inhibitors for urokinase receptor. WO98/46731
Kasumi T, Araki K, Mizushima T, Kobayashi H, Kurita N (2013) A proposal of potent inhibitor for cancer metastasis blocking the pocket of urokinase receptor: ab initio molecular simulations. J Phys: Conf Ser 433:012034
Tsuji S, Kasumi T, Nagase K, Yoshikawa E, Kobayashi H, Kurita N (2011) The effects of amino-acid mutations on specific interactions between urokinase-type plasminogen activator and its receptor: ab initio molecular orbital calculations. J Mol Graph Model 29:975–984
Kasumi T, Araki K, Ohyama T, Tsuji S, Yoshikawa E, Kobayashi H, Kurita N (2013) The effects of vitronectin on specific interactions between urokinase-type plasminogen activator and its receptor: ab initio molecular orbital calculations. Mol Simul 39:769–779
Huai Q, Zhou A, Lin L, Mazar AP, Callahan J, Shaw DE, Furie BC, Huang M (2008) Crystal structures of two human vitronectin, urokinase and urokinase receptor complexes. J Nat Struct Mol Biol 15:422–423
Schwede T, Kopp J, Gues N, Peitsch MC (2003) SWISS-MODEL: an automated protein homology-modeling server. Nucleic Acids Res 31:3381–3385
Li H, Robertson AD, Jensen JH (2005) Very fast empirical prediction and interpretation of protein pKa values. Proteins 61:704–721
Delphine CB, Rogers DM, Jensen JH (2008) Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins 73:765–783
Olsson MHM, Søndergard CR, Rostkowski M, Jensen JH (2011) PROPKA3: Consistent treatment of internal and surface residues in empirical pKa predictions. J Chem Theory Comput 7:525–537
Case DA, Cheatham TE III, Darden T, Gohlke H, Luo R, Merz KM Jr, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Wang J, Cieplak P, Kollman PA (2000) How well does a restrained electrostatic potential (RESP) model perform in calculating conformational energies of organic and biological molecules? J Comput Chem 21:1049–1074
Jorgensen WL, Chandrasekhar J, Madura J, Klein ML (1983) Comparison of simple potential functions for simulating liquid water. J Chem Phys 79:926–935
Frisch MJ et al. (2009) Gaussian09. Gaussian Inc., Wallingford, CT
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791
Kitaura K, Sawai T, Asada T, Nakano T, Uebayashi M (1999) Pair interaction molecular orbital method: An approximate computational method for molecular interactions. Chem Phys Lett 312:319–324
Ito M, Fukuzawa K, Ishikawa T, Mochizuki Y, Nakano T, Tanaka S (2008) Ab initio fragment molecular orbital study of molecular interactions in liganded retinoid X receptor: specification of residues associated with ligand inducible information transmission. J Phys Chem B112:12081–12094
Mochizuki Y, Nakano T, Koikegami S, Tanimori S, Abe Y, Nagashima U, Kitaura K (2004) A parallelized integral-direct MP2 method with fragment molecular orbital scheme. Theor Chem Acc 112:442–452
Mochizuki Y, Koikegami S, Nakano T, Amari S, Kitaura K (2004) Large scale MP2 calculations with fragment molecular orbital scheme. Chem Phys Lett 396:473–479
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper belongs to Topical Collection 9th European Conference on Computational Chemistry (EuCo-CC9)
Rights and permissions
About this article
Cite this article
Mizushima, T., Sugimoto, T., Kasumi, T. et al. Ab initio molecular simulations for proposing novel peptide inhibitors blocking the ligand-binding pocket of urokinase receptor. J Mol Model 20, 2292 (2014). https://doi.org/10.1007/s00894-014-2292-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00894-014-2292-7