Abstract
The influence of metal ions on the structure of amyloid-\(\beta \) (Aβ) protofibril models was studied through molecular dynamics to explore the molecular mechanisms underlying metal-induced Aβ aggregation relevant in Alzheimer’s disease (AD). The models included 36-, 48-, and 188-mers of the Aβ42 sequence and two disease-modifying variants. Primary structural effects were observed at the N-terminal domain, as it became susceptible to the presence of cations. Specially when β-sheets predominate, this motif orients N-terminal acidic residues toward one single face of the β-sheet, resulting in the formation of an acidic region that attracts cations from the media and promotes the folding of the N-terminal region, with implications in amyloid aggregation. The molecular phenotype of the protofibril models based on Aβ variants shows that the AD-causative D7N mutation promotes the formation of N-terminal β-sheets and accumulates more Zn2+, in contrast to the non-amyloidogenic rodent sequence that hinders the β-sheets and is more selective for Na+ over Zn2+ cations. It is proposed that forming an acidic β-sheet domain and accumulating cations is a plausible molecular mechanism connecting the elevated affinity and concentration of metals in Aβ fibrils to their high content of β-sheet structure at the N-terminal sequence.
Graphic abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1. Introduction
Considered by the World Health Organization (WHO) as a public health priority, Alzheimer’s Disease (AD) accounts for approximately 60–70% of dementia cases worldwide [1, 2]. This neurodegenerative disorder causes deterioration of memory and cognitive ability to perform everyday activities [1, 3]. One of the hallmarks of AD is the presence of fibrillar protein deposits in the brain known as senile or amyloid plaques composed mainly by the amyloid-β peptide (Aβ). It was originally conjectured that the accumulation of Aβ and its aggregation into amyloid fibrils would be causative in neurodegeneration leading to AD [4]. However, later findings point to oligomeric forms of Aβ as the most neurotoxic species [5, 6].
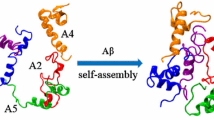
Aβ is produced from the proteolytic cleavage of the amyloid precursor protein (APP) by β- and γ-secretases, yielding Aβ peptides with 40 to 43 amino acid residues [7]. Among the different Aβ variants [8,9,10], Aβ42 is the most abundant species in AD amyloid plaques, it is the most aggregation-prone, and it produces toxic oligomers [10,11,12]. The peptide may be divided into two domains: the hydrophilic N-terminal (residues 1-16) that binds metal ions, such as copper or zinc; and the hydrophobic C-terminal domain (17-42), engaged in β-sheet formation (Fig. 1a) [13, 14]. Aβ displays structural flexibility and sensitivity to environmental factors such as temperature, concentration, pH, and interaction with metal ions or membranes. In non-polar solvents or membranes, Aβ adopts predominantly β structure, [15,16,17] whereas in aqueous solution it may be found with an extended random coil structure (Fig. 1b) [18], with a tendency to aggregate into oligomeric species. Soluble-Aβ oligomers can go from two and up to hundreds of Aβ peptides with unknown structure; however, increasing oligomer size (dimer, trimer, etc.) has been associated with higher contents of β structure with a concomitant increase in toxicity [19, 20]. Larger aggregates form non-crystalline and pleomorphic deposits of insoluble fibrils. Even though high-definition structural characterization of fibrils is difficult, several structural features have been described and are briefly discussed below [21,22,23,24].
Structure of the Aβ protein. a Primary structure. b NMR structure in trifluoroethanol/water solution (PDB: 1z0q). c Solid-state NMR structure of the hydrophobic core of Aβ42 fibrils (PDB: 2beg). d Near-atomic resolution cryo-EM structure of Aβ42 fibrils (PDB: 5oqv). The amino acid type is highlighted in red for acidic, blue for basic, green for polar, and gray for non-polar residues
The structure of amyloid fibrils contains the cross-β motif, consisting of β-sheets formed along the fibril, with peptide strands extending approximately perpendicular to the long fibril axis and orienting inter-strand backbone hydrogen bonds approximately parallel to the same direction [21,22,23]. This arrangement, also known as a filament, may be seen as an array or a layer of protein molecules aligned along the fibril axis (Fig. 1c). In Aβ fibrils, a minimum of two or three of these layers (with two- or three-fold symmetry) are commonly associated in forming a structural unit known as protofibril [25,26,27,28,29]. Lateral association and supercoiling of two or more protofibril subunits would form “mature” fibrils [23].
Each Aβ peptide in the cross-β motif has been proposed to fold into a β-sheet-turn-β-sheet motif [27, 30, 31], although many different folds have been described so far [32,33,34,35,36,37,38,39,40,41,42]. (Fig. 1d) This arrangement allows the formation of an “intra-layer” steric zipper, where side chains from two stacking β-sheets interdigit and confront complementary hydrophobic surfaces [43, 44]. Steric zippers formed in the interface between two Aβ layers are also possible [28], and provide structural stability to the hydrophobic core of protofibrils. Amyloid fibrils are generally observed as twisted structures within a range of crossover lengths and fibril widths [45], although non-twisted topologies have been observed for amyloid fibrils seeded from brain tissue samples [34]. These and other features like Aβ variants, oligomeric forms, intra/interchain contacts, peptide fold, arrangement of steric zippers, the parallel/antiparallel orientation of β-strands, amount of β structure, sensitivity to environmental factors or experimental conditions, variable lateral association/supercoiling, and diverse metal binding modes/sites, all contribute to the characteristic polymorphism observed in Aβ fibrils [46,47,48,49]. Additionally to ssNMR-derived structural models, recent cryo-electron microscopic models have provided greater insights into the diversity of molecular arrangements that the structure of these aggregates may cover [39,40,41,42, 50].
The involvement of Aβ and its oligomeric forms is considered a necessary but insufficient factor responsible for producing the neuropathology in AD, implying that additional cellular and biochemical alterations must exist [51, 52]. Among these, altered metal homeostasis has been implicated as a pathogenic factor in AD [11, 14, 53,54,55,56]. Biometals such as copper and zinc promote aggregation of Aβ and are found concentrated within amyloid plaques [57,58,59,60,61,62,63,64,65]. Moreover, metal ion homeostasis, including zinc, is perturbed in Alzheimer’s disease [66,67,68]; disturbances on metal ion homeostasis in Alzheimer´s disease is also summarized in refs [56] and [69]. During synaptic transmission, copper and zinc are released into the synaptic cleft, where they are found as Cu2+ and Zn2+ ions [66, 71]; these metal ions can coordinate to the N-terminal region of Aβ regardless of its state as monomer, oligomer or fibril. It has been proposed that Cu-Aβ species potentially produce reactive oxygen species (ROS), contributing to oxidative stress and neurodegeneration [12, 72]. Even though it is known that the toxicity of oligomeric Aβ is dependent on its metal content [73,74,75,76,77], and that the affinity of biometals for these species may compete with the affinity for other metalloproteins [14, 78, 79], the molecular mechanisms leading to metal-induced aggregation, metal accumulation in amyloid plaques and metal-enhanced cytotoxicity remain unclear.
To gain a better understanding of the role played by metal interactions in the molecular mechanisms underlying the formation of amyloid fibrils, in this work, we addressed the issue by performing molecular dynamics (MD) simulations on a series of atomistic Aβ42 amyloid protofibril models in the presence of different ion species. A molecular model is proposed to explain the increased metal ion affinity in amyloid oligomers and fibrils, relating it to the increase of β-sheet structure at the N-terminal domain. The proposed model was tested using disease-modifying variants of the Aβ peptide that contain N-terminal substitutions on key residues. For these variants, significant differences in the number of metal cations accumulated as well as in the amount of N-terminal β-sheets were found, suggesting that the formation of N-terminal β-sheets may be considered as a molecular phenotype modulating the metal-interaction behavior of different Aβ sequences and the concomitant formation of fibrils.
2. Computational methods
Amyloid-β (1-42) protofibril models
The structure of the Aβ42 protofibril models used in this work are based on the solid state nuclear magnetic resonance (ssNMR) model 2beg [30] from the Protein Data Bank (PDB), considering also specific quaternary-structure interactions observed in ssNMR measurements of Aβ40 fibrils [28]. The structure of the N-terminal domain was incorporated into the models as detailed in the Supplementary Information (Section SI-1). The starting model (I), a 36-mer arranged as a 2-layered C2-pseudosymmetric protofibril, included an N-terminal (1–16) domain in extended (β) conformation, a non-twisted topology which was spontaneously corrected (twisted) during simulations (see Sect. 3.1), and the same “intra-layer” steric zipper as the original 2beg model along with a new “inter-layer” steric zipper holding the two layers together (see Figs. 2a and b).
Initial structure of the protofibril models I and V. a Building of the 36-strands oligomer (Aβ42)36 (model I). (i) Three chains of Aβ17−42 were extracted from the PDB structure 2beg (blue strands). (ii) The missing residues 1–16 were added to each chain in an extended conformation (red strands). (iii) The three chains were reproduced along the fibril axis to reach 18 monomers (green strands). (iv) The whole oligomer was duplicated, rotated over the fibril axis, and assembled to make a C2-pseudosymmetry structure with two layers of Aβ42 molecules. b Cross-sectional view of two strands from each fibril layer; the amino acid type is highlighted in red for acidic, blue for basic, green for polar, and gray for non-polar residues. c Unit cell of the infinite-length (Aβ42)188 protofibril model V
As described in Section SI-2 in the Supplementary Information, models II–IV and VI–VII (48-mers) were built from the equilibrated structure of model I; model V (188-mer) was built from model III. An inherited feature in these models was the twisting of the protofibril in the equilibrated model I; considering the equilibrium degree of twisting and the C2 pseudosymmetry of the constructs, model V was built using 94 monomers per protofibril layer that allowed to reach a 180° twist and the periodic reproduction of the structure along the fibril axis. We took advantage of this symmetry and adjusted the water box of model V to allow the periodic boundary conditions to represent an infinite-length amyloid protofibril (Fig. 2c). Table 1 summarizes the composition of the models. The protonation state of all the models corresponds to neutral pH with histidine residues protonated at the Nδ of its imidazole ring.
The water box for models I-VII included different numbers and types of ions to evaluate their effect on the structure of the fibril. Model I included 3 equivalents (eq) of Na+ cations to neutralize the protein charge (-3 per Aβ42 monomer), while model II was devoid of ions. For models III–VII, the same concentration (0.1 M) of Na+ and Cl− ions was considered, i.e., the number of sodium and chloride ions were adjusted to neutralize the final total charge and to reach a salt concentration of 0.1 M, whereas 1 eq of Zn2+ (a “soft-sphere” cation per Aβ42 monomer with random positions) was added to models IV, VI, and VII. Models VI and VII were intended to assess the stability of N-terminal β-sheets of disease-modifying mutations at the N-terminal region and the effect on the interaction with cations. Model VI contains the familiar AD-promoting Tottori mutation (D7N), whereas model VII corresponds to the non-amyloidogenic Aβ42 rodent sequence (R5G, Y10F, and H13R). These considered the same base structure as models II-IV and the same solvent conditions as model IV (1 eq of Zn2+ cations and 0.1 M of NaCl). For further details, see Supporting Information section SI-3 and Sect. 3.6.
2.2 Molecular dynamics simulations
Equilibration and MD simulations of the models were carried out using the program NAMD2.7[80] and the CHARMM27 force field [81]. Non-bonded interactions were calculated by applying a cutoff of 10 Å and a switching function at 8 Å. Electrostatics were calculated using the particle-mesh Ewald method for periodic boundary conditions [82, 83]. A time step of 2 fs was used to integrate the equations of motion, together with the SHAKE algorithm [84] to keep all bonds to hydrogen atoms rigid. The temperature and the pressure were controlled through Langevin dynamics and the Langevin piston Nosé–Hoover methods as implemented in NAMD [85, 86]. The equilibration protocol consisted of solvent-adaptation, warming up, and density-adaptation stages. For model I, the water molecules were optimized through 3000 conjugate gradient energy-minimization steps and warmed up from 10 to 310 K in a 20 ps simulation, with the protein structure fixed. The system was then released from any structural constraint and minimized for another 3000 steps, followed by a new 100 ps warming simulation and a last NTP run at 310 K and 1 atm for 40 ps. Equilibration of models II-VII followed the same protocol but with longer minimization (5000 steps) and warming up (200–400 ps) runs. The final conditions of the equilibration (310 K, 1 atm) in the NTP ensemble were used in the 100-ns production simulations for all models; in addition, a replica of the simulation of the model I was run for 200 ns to ensure a stationary structure. All structure manipulation, visualization, and trajectory analyses were done using the VMD program. [87] Secondary structure assignments were done using the STRIDE algorithm [88].
Results and discussions
Several interesting features were observed from the simulations performed on the base model I, most very well recognized by experimental or theoretical evidence. In the following sections, several of these features for all the models are analyzed in detail, and their implications on the molecular mechanisms underlying amyloid aggregation and metal-accumulation are discussed.
3.1 Structural features of the amyloid protofibril models
After 200 ns of MD simulations, the whole structure of the protofibril model I remained assembled, and most of the features initially adopted were conserved and stabilized. In Fig. 3a–c, the final structures of this model are depicted, highlighting some of the structural transitions and features observed during the simulations, which include twisting of the protofibril structure, formation of water channels within the hydrophobic core, changes in secondary-structure motifs, folding of the N-terminal domain, and distribution of the cations on the surface of the oligomer. These features are described in detail in the following paragraphs.
Structure of model I after 100 ns of MD simulation at 310 K and 1 atm. a–c show a lateral, superior, and frontal views, respectively, of the structure. Yellow circles represent the final positions of Na+ ions. The positions of the water molecules inside the hydrophobic core are also highlighted. d RMSD Cα vs Time plot considering the whole protofibril model I (red) and only the hydrophobic core (blue) for 200 ns of simulation
Although the length of the production simulations may seem too short in comparison to the time needed for amyloid fibrils to form, our models were built from an “already consolidated” fibril structure; thus, the simulation time comprises exclusively the time needed for the structure to adapt and equilibrate to the simulation conditions. Whereas many reported simulations of Aβ have focused on long simulation runs (up to microsecond scale) or enhanced sampling of small oligomers [89,90,91,92], in this work, we have focused on larger aggregates to deal with structural features more compatible with consolidated protofibrils rather than small soluble oligomers, with the sacrifice of longer simulation times or exhaustive sampling. In other words, we select larger space scales rather than longer simulation runs. The plot of the root-mean-squared deviation for alpha carbons (RMSD Cα) shown in Fig. 3d confirms that the simulation time was long enough to obtain a stationary structure and to observe important structural transitions (The RMSD Cα plots for the protofibril domains of all models are depicted in Figures SI-1 and SI-2 of the Supplementary Information).
The most evident structural transition in the simulation of model I was the twisting of the aggregate over the protofibril axis (see Figure SI-3a). This feature is common in protein structures with cross-β motifs such as Aβ fibrils [21,22,23], and has been reported in several MD studies [93,94,95,96,97,98,99,100]. The average angle formed between each pair of monomers plotted against the simulation time, shown in Figures SI-3b, SI-3c, and SI- 3d, quantifies the degree of twisting for all the models. For model I, the twist transition was almost completed within the first 5 ns of simulation, reaching an average angle of ∼2.2°. Considering this degree of twisting, the length required for a longer aggregate to perform a full 180° twist (i.e., crossover distance) is estimated to be ∼39 nm, corresponding to ∼82 Aβ42 parallel strands separated by ∼4.8 Å along the protofibril axis (see Table 2).
As it can be seen in Table 2 and in Figures SI-3c and SI-3d, models II–IV, VI, and VII (48-mers) reached a lower degree of twisting relative to model I (a 36-mer); therefore, increasing the size of the oligomer reduces the degree of twisting, as it has been already shown in some MD simulations [93]. Comparing only 48-mers with a different ion environment, model II, containing no ions in the solvent, reached the highest level of twisting (2.1°); models III, IV, VI, and VII showed average angles between strands of ∼1.7°–1.9° per strand, although important fluctuations during the simulations were observed. Thus, the ions present in the solvent also seem to affect the morphology of the protofibril models.
Along the simulations, the models showed degrees of twisting (∼1.5°–2.5° per strand) corresponding to fibril crossover distances in the 35–56 nm range. These results are directly comparable to experimental morphologic characterizations of amyloid fibrils. For instance, a distribution of crossover distances for amyloid Aβ40 fibrils grown in phosphate buffer showed a maximum frequency at 50–60 nm [45]; the smallest crossover distances found in that work were ∼30 nm, although even shorter distances (∼25 nm) have been reported [26]. Thus, the degree of twisting obtained in the simulations of the present work is very well aligned with the experimental observations. The capability of reproducing the experimental topologic parameters confirms that the structural features incorporated in the models, all compatible with the definition of the cross-β motif, successfully reproduce the morphology of amyloid protofibrils.
Considering the hydrophobic core region, i.e., sequences 17–42 in both protofibril layers forming steric zippers and stacking four β-sheets (Fig. 1), after production simulations, all the protofibril models conserved the structure of their hydrophobic core assembled and stable, with the two layers of Aβ42 peptides paired (intertwined). A systematic study on the size of single/paired-topology oligomers based on the same experimental structure as the present models concluded that a minimum oligomer size of 10–12 monomers would be necessary to obtain stable paired topologies [93]. Our models behaved consequently, in the sense that the number of monomers considered in our models was expected to keep the assigned topology during all the simulation runs, as they did.
The two types of steric zippers forming the hydrophobic core of the models, i.e., inter-layer and intra-layer steric zippers, in general, conserved the structure of the base models with a few changes. Figure 4 shows the main structural changes observed for model I; a detailed analysis and comparisons can be found in the supporting material (Figures SI-4 and SI-5). For inter-layer steric zippers (blue region in Fig. 4), a compact and dry structure was observed for all the models despite the differences in the protofibril twisting. Although MD simulations have studied different arrangements of the inter-layer steric zipper [93, 94, 98], the general stability observed in all our simulations validates the adopted arrangement. Moreover, this arrangement may explain the reduced aggregation observed for the Met35-sulfoxide Aβ variant [101], as this modification would produce steric hindrance and reduced hydrophobicity within the steric zipper.
Structure of the steric zippers for a middle Aβ strand in the hydrophobic core of the fibril model I. a Initial structure. b Twisted structure after 100 ns of MD simulations. In blue, the steric zipper from the interface between the two fibril layers is highlighted, and in green, the steric zipper inside layer
Another interesting feature observed in Fig. 4b (green region) is the opening of the intra-layer steric zippers near the turn region, also observed as a broader distribution of distances between alpha carbons in Figure SI-5b. This structural change had two main consequences: (1) distortion of the secondary structure close to the turn region, (see Sect. 3.4), and (2) formation of cavities that allowed the incorporation of water molecules into the hydrophobic core (see also Fig. 3c). This later observation has been reported in MD simulations of Aβ amyloid fibrils [93, 94, 98] or in structures with amphipathic steric zippers [102]. The water molecules in these channels tend to form hydrogen bonds with the peptidic backbone, and even some water molecules were found between the fibril strands. These kinds of interactions have been proposed from 2D infrared experiments. [103]
3.2 Structure of the infinite-length (periodic) protofibril model V
Figure 5a) shows the final structure of model V in the context of its unit cell reproduced periodically. As can be observed, the symmetry of the construct allowed the modeling of an infinite-length protofibril that remained assembled along the simulation.
Structure of the periodic amyloid protofibril model V. a Perspective view of the final structure (at 100 ns of MD simulation). Yellow and orange spheres represent Na+ and Cl− atom positions, respectively. b Structure of the hydrophobic core after 100 ns of MD simulation (blue) superimposed to the initial structure (green). The final structure looks arched in the unit cell in comparison to the initial structure
The most interesting features observed for model V were: (1) the reduction of the N-terminal β-sheet structure, which seems to be an effect of the size of the oligomer (see Sect. 3.3), and (2) the curvature or arching of the protofibril highlighted in Fig. 5b. This arching occurred during the first 30 ns of simulation (Figure SI-1 b in the Supporting information). At this point, the origin of this structural transition is not clear. Two possible factors may be argued: (1) the need for twisting relaxation since, according to our observations, the longer the aggregate, the smaller the degree of twisting, but in model V, it is restricted due to its periodicity. (2) The need for supercoiling: model V is more compatible with a “protofibril” species that tends to associate laterally with one or more similar structural units to form mature supercoiled fibrils, which may be required to form stable linear constructs. Therefore, the arched structure of model V may be more suitable to intertwine with other protofibrils to be stabilized as a straight mature amyloid fibril. It is important to remark that, to our knowledge, this is the first model of an infinite-length amyloid protofibril considering twisted topology; see, for instance, Refs. [94, 95] for analysis of non-twisted periodic models. This model shows interesting differences in relation to finite-length oligomers, and it is worth further analysis in future work as well as for contrasting with experimental observations.
3.3 Structure of the N-terminal domain of the amyloid protofibril models
Most of the structural effects observed during the simulations, highlighted in Fig. 3, are associated with the N-terminal region in the protofibril model I. After comparing it with the rest of the models, the structure of this region turned out to be highly affected by the presence of different ions in the solvent. Three features at the N-terminal region of model I stand out: the high amount of β-sheet structure, the folding of the amino-termination, and the regioselective accumulation of cations. These features seem to be related to each other and were found to vary quantitatively when considering different models. As described in deeper detail later, this behavior seems to be related to the high incidence of acidic residues at specific positions of the N-terminal tail, which generates an acidic domain when the aggregate shows β-sheet structure in this region. The high incidence of N-terminal β-sheet opposes several experimental reports, however, the observed elevated stability in most simulations deserves careful analysis. The other two features, namely, the folding of the N-terminal tail and its regioselective accumulation of cations, are processes much less analyzed in the literature due to the difficulty of resolving flexible parts of amyloid fibrils. Thus, the present simulations offer an opportunity to analyze features often invisible to experiments on amyloid aggregates. To better rationalize these features, we are defining a model termed the N-terminal acidic β-sheet domain that allows us to explain the observed features; see below. Due to the relevance of the interaction of metal cations with amyloid fibrils, a detailed discussion of these features and the implication of amyloid aggregation is also done in the forthcoming sections.
3.4 Impact of metal ion binding in the structure of Aβ protofibril models
In this and the following sections, those structural transitions and features directly related to the interaction of metal cations with the Aβ42 protofibrils in the simulations, including the effect of modifications in the amino acid sequence of the N-terminal domain of the aggregates, are discussed.
From the high amount of β-sheet structure observed in Model I (see Fig. 6a and b) and the comparison with the rest of the models (Fig. 6c and Figure SI-6 in the Supporting Information), it is clear that the secondary structure of the N-terminal region of the aggregates is the most sensitive to the presence of different ions in the models. The amount of β-sheet structure at the hydrophobic core was very stable, which also included part of the N-terminal domain starting at residue 10, showing consistent structure patterns in all the models and good agreement with several reports [23].
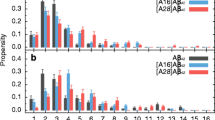
Incidence of secondary structure motifs in Aβ42 protofibril models. a Initial secondary structure assignment of model I. b Average incidence of secondary structure motifs per residue for all strands over the first 100 ns of MD simulation for model I. c Incidence of β-sheet structure for different domains of all the protofibril models
At the N-terminal domain, the amount of β-sheet motif was dependent on several factors; considering models that varied their content of ions, model I showed the highest incidence of β structure at the N-terminal domain (52%), whereas the lowest incidence was observed in model II (devoid of ions) (25%). Relative to model I, the increase in the concentration of Na+ cations and the addition of Cl− anions in model III decreased the β structure to 40%; further addition of 1 eq of Zn2+ cations in model IV decreased even more that motif (34%). Interestingly, with the same ion contents as that in model III, the periodic model V reduced N-terminal β-sheet (34%), indicating that aggregates of smaller size would favor the stability of the N-terminal β-sheet.
Another trend is that the presence of Na+ ions seems necessary to favor the N-terminal β-sheet structure. In contrast, the presence of anions or divalent cations would interfere with the formation of this motif. This Na+-mediated stabilization of secondary structure is in line with a computational study of the influence of alkali ions on the structure and stability of amyloid-β oligomers, concluding that charge compensation and carboxylate bridging explained the stabilizing effect of alkali ions, which was more evident in smaller oligomers [104]. The observed trend is also consistent with experiments reporting the reduction of β-sheet structure by Cu2+ in Aβ42 fibrils [105] or by Cu2+ or Zn2+ in both Aβ40 and Aβ42 [106]. Metal ions affect the final morphology of amyloid fibrils, forming less defined or even non-fibrillar aggregates [63, 77, 106]. Moreover, copper-chelating agents yield less branched and elongated fibrils, as compared to those grown in the presence of the metal [107], which indicates that copper may induce a different aggregation pathway for Aβ.
Although, the structure of the N-terminal region in model I was initially built in extended (β) conformation, this motif was expected to be unstable in a relatively long production simulation, as this region was generally reported as a random coil [30]. However, even after 200 ns of MD simulation, the amount of N-terminal β-sheet structure did not show any decreasing trend during the simulation of model I (see Figure SI-7 in the Supporting Information). On the other hand, several experiments provide evidence of forming N-terminal β-sheets in Aβ aggregates [39, 106, 108,109,110]. Moreover, several structural models have been solved with well-defined N-terminal tails with stable β-sheet motifs [34, 36, 38, 39]. Interestingly, it has been suggested that N-terminal β-sheets may play a role in the amyloid aggregation pathway, favoring protofibril species over mature fibrils [110]. The consolidation/distortion of β-sheets in the N-terminal domain of amyloid oligomers may thus be characteristic of different metastable forms of oligomeric Aβ species resulting from different aggregation pathways.
The interaction of the ions with the protofibril models was analyzed to understand the origin of the regioselective accumulation of cations in the structural features observed at the N-terminal domain of the models. As depicted in Fig. 7a), the Na+ ions saturated the surface of the protofibril model I during the first ∼10 ns of simulation. We quantified the average number of ions in contact with each Aβ42 residue (within 3.5 Å), finding that the residues interacting with Na+ ions (in addition to Glu22 and Ala42) were found at the N-terminal domain and exclusively at odd positions (Fig. 7b). This regioselective interaction may be explained in terms of the β-sheet structure preserved at the N-terminal domain of model I, which keeps the side chains of the acidic residues (Asp1, Glu3, Asp7, and Glu11) on a single face of the N-terminal β-sheet. This produces a region predominantly rich in acidic residues, capable of attracting and concentrating cations from the media, and that can be described as an N-terminal acidic β-sheet domain (Fig. 7c).
Ions on the surface of protofibril models within 3.5 Å from the protein. a Number of sodium ions against time for model I. b Average number of sodium ions per strand and per residue for model I. c Structure of the N-terminal domain of one strand of model IV. Na+ (yellow), Cl− (orange), and Zn2+ (green) ions are shown
Four conditions drive the formation of the acidic β-sheet domain: (1) The conserved β-sheet structure at the N-terminal domain that properly orients acidic side chains; (2) the structure and fold of the hydrophobic core making the peptidic carboxylate of Ala42 reachable to the N-terminal domain; (3) the folding of the N-terminal domain toward the hydrophobic core allowed by a bend-like region at residues Ser8 and Gly9; (4) the Aβ primary sequence by itself, specially at the N-terminal tail. Indeed, the arrangement of the N-terminal acidic residues along with Arg5 and His13, as shown in Fig. 7c, strongly suggests a functional role of the N-terminal sequence of the Aβ peptide. Combining these features to form this acidic region establishes a connection between the accumulation of cations and the formation of the N-terminal β-sheet, a process dependent on the aggregation of the Aβ protein. Additionally, some experimental reports support the formation of the acidic β-sheet domain. For instance, ssNMR measurements have shown the participation of carboxylate groups from glutamate residues and the C-terminus of Aβ40 fibrils in Cu.2+ coordination [111]. Moreover, a kink or bend-like structure, consistent with the observed bending at Ser8 and Gly9, has been reported. [18, 108]
In the case of model IV incorporating Zn2+ cations, despite the reduction of the N-terminal β-sheet, the regioselective accumulation of cations was also observed (Fig. 8). Actually, the ion-density isosurfaces in Fig. 8a) show that the specific interaction of Zn2+ affected the secondary structure and fold of the N-terminal domain. Figure 8b shows that both Na+ and Zn2+ cations seem to compete for the acidic β-sheet domain of the model. Although Na+ ions saturated the protofibril surface within 20 ns, the number of interacting Zn2+ ions slowly raised during 80 ns of this simulation, displacing the necessary amount of sodium.
Interaction of ions with fibril model IV. a Ion density isosurface (isovalue = 0.07, considering the last 20 ns of simulation) for Zn2+ (green) and Na+ (yellow) ions over the final fibril structure (blue β-strands). b Average number of ions per fibril strand at less than 3.5 Å. Yellow, orange and green spheres represent Na+, Cl− and Zn2+ ions, respectively
The formation of the acidic β-sheet domain implies folding or bending of the N-terminal domain toward the hydrophobic core. We quantified this folding for each model as the distribution of distances between N- and C-terminal alpha carbons through the radial pair distribution function g(r) shown in Figure SI-8. For the models varying the content of ions (I–IV), a maximum close to 30 Å was observed except for model II, where the maximum was shifted up to 43 Å. This suggests that the absence of ions in model II produced more extended (less folded) strands at the N-terminal domain, probably due to the repulsion of negative charges from the acidic residues at the acidic β-sheet domain. The difference in the folding of the models containing cations relative to model II shows the effect of the cations in shielding the negative charge at the acidic β-sheet domain.
3.5 Implications on the N-terminal acidic β-sheet domain in metal binding
The described molecular mechanisms at the N-terminal domain of the protofibril models have important implications in the coordination chemistry of amyloid fibrils, since the residues forming the acidic β-sheet domain may also participate in metal coordination. The involvement of Aβ N-terminal residues such as Asp1, His6, Glu11, His13, and His14 in metal coordination either as monomers or oligomers has been described [111,112,113,114,115,116,117,118,119]. Considering that histidine residues are known to be effective anchoring ligands, specially in multi-histidine peptides forming macro-chelate rings [120], metal coordination involving histidine together with aspartate and/or glutamate residues will impose structural restrictions that would enhance the observed reduction of the N-terminal β-sheet, without necessarily affecting the hydrophobic core of the fibril. Copper coordination to N-terminal residues of the protofibril was already studied in models of the peptide Aβ1-17 in complexes with Cu2+ through all-electron density functional theory (DFT) calculations from one of the strands of model I [121]. Although several binding modes were found upon copper coordination, the extended conformation from residues 11 to 17 was almost unaffected (see Figure SI-9 in the Supplementary Information). This supports the fact that the effects of the metal ions on the amyloid fibril structure originated at the N-terminal domain, which is also compatible with the recently proposed N-terminal hypothesis for AD [122].
3.6 Molecular phenotype of Aβ42 variants: effect of N-terminal mutations on metal binding and β-sheet stability
Together with the turn region, the N-terminal domain has been shown to contain the major structural differences in Aβ oligomers relative to mature fibrils; moreover, both regions contain most of the Aβ mutations relevant in AD [123]. Thus, these regions are of particular interest. Considering also the formation of the acidic β-sheet domain and its tendency to accumulate cations, we aimed to investigate the effect of mutations in the N-terminal domain on the metal-accumulation properties of this region considering known disease-modifying variants of Aβ. Particularly, mutations in residues forming the acidic β-sheet domain are expected to have an important effect in the binding of ions and the stability of the N-terminal β-sheet structure according to the molecular mechanisms observed on the protofibril models described in previous sections.
As already argued, the participation of acidic residues (Asp1, Glu3, Asp7, and Glu11) and the bending of Ser8 and Gly9 are key to forming the acidic β-sheet domain. Residues Arg5 and His13 have interesting implications too. Arg5 was observed to assist in folding the N-terminal domain and consolidating the N-terminal β-sheet, whereas His13 would have a role in metal coordination. These residues, along with others forming the acidic β-sheet domain, are known to be relevant in disease-modifying natural variants of the Aβ sequence. For instance, the rodent-Aβ sequence (H13R, R5G, and Y10F substitutions), which rarely forms amyloid plaques (i.e., a protective variant) [124], was shown to form amyloid plaques with reduced metal binding abilities relative to human Aβ [125]. These observations may be related to the highlighted roles of Arg5 and His13. Moreover, His13 is a crucial residue in the Zn2+-induced aggregation of Aβ [126]. In the context of the present analysis, the regioselective accumulation of cations may be proposed to be a preamble favoring Zn2+ coordination to His13 in the acidic β-sheet domain, which affects the aggregation pathway and thus would be altered in the rodent sequence, explaining the behavior of this variant. Interestingly, isothermal titration calorimetry (ITC) measurements on rat Aβ1-16 peptide in the presence of Zn2+ may provide insights into such role for His13; [127] in those experiments only residues His6 and His14 of the rat sequence were shown to be involved in Zn2+ coordination, forming peptide dimers complexed with zinc in an arrangement that could hinder peptide aggregation [127]. In contrast, metal binding to His13 in human Aβ would promote coordination to Asp and Glu residues within the acidic β-sheet domain. Other substitutions found in the acidic β-sheet domain, such as D7N, known as the Tottori familial AD mutation [128], are known to modify the aggregation properties of this (AD-promoting) Aβ variant [129, 130]. From the analysis performed on models I-V, specially regarding the regioselective accumulation of cations at the acidic β-sheet domain and the elevated stability of the N-terminal β-sheet, we expected that mutation of residue Asp7 to asparagine in the Tottori variant would modify the metal-accumulation behavior in the studied models. We also chose the rodent-Aβ sequence as a second variant associated with an opposite phenotype relative to the Tottori to gain further insights into the roles of N-terminal residues.
To evaluate such roles, we built models VI and VII based on the Tottori and rodent sequences of the Aβ42 protein, respectively, which were subjected to the same simulation conditions and ion contents as model IV, acting as reference (human Aβ sequence). Table 3 shows the time-averaged incidence of N-terminal tail β-sheet and the number of cations in the surface of the protofibril models, whereas Figures SI-10 and SI-11, show the detailed time evolution of each property.
From Table 3, it can be observed that the substitutions at the N-terminus of the Aβ sequence produced important differences in the amount of N-terminal β-sheet structure, the folding of the N-terminal tail, and the number of cations accumulated at the acidic β-sheet domain. The AD-promoting D7N variant (model VI) maximized the N-terminal β-sheet amount, whereas the aggregation-preventive rodent sequence (VII) minimized it. The amount of N-terminal β-sheet may thus be considered a molecular phenotype for the aggregates of these variants. These observations are consistent with experimental measurements of secondary structure for the Tottori variant that, additionally, were associated with the increased toxicity of these aggregates [130]. Moreover, replica-exchange molecular dynamics simulations on D7H, D7N, and H6R mutants of the Aβ peptide showed an increase in the amount of β structure and enhanced the turn at Ser8 and Gly9. [131] The distortion of the N-terminal β-sheet in the rodent sequence may be directly associated with its reduced aggregation, whereas the reduced metal load may be associated with the H13R substitution located in the acidic β-sheet domain. The N-terminal β-sheet distortion in model VII resulted in collapsed strands, as reflected by the distributions of N-ter to C-ter distances (see Figure SI-8).
The average amount of each ion in contact with the acidic β-sheet domain (within 3.5 Å) shows that among the three models in Table 3, both the amount of Zn2+ and Na+ cations varied while the amount of Cl− remained almost the same. Zn2+ cations were more crowded in model IV; for model VI, the amounts of Zn2+ and Na+ decreased relative to IV, whereas in model VII, Zn2+ decreased and Na+ increased. The Zn2+/Na+ ratio reveals that the proportion of these cations is almost the same for models IV and VI and that model VII became selective for Na+ cations. Therefore, although reducing the negative charge in the D7N variant reduces the number of both cations, the selectivity for each cation depends on the Aβ sequence and probably on the amount of the N-terminal β-sheet motif.
The results regarding the amount of N-terminal β-sheet are consistent (in terms of the associated phenotypes) with the effects on the N-terminal domain of the A2T and A2V mutations studied in Aβ42 dimers [132], the AD-protective A2T variant diminishes N-terminal contacts, whereas the AD-causative A2V variant and the wild-type sequence promotes them. From the present study, the AD-protective rodent sequence reduces the N-terminal β-sheet (i.e., diminishes N-terminal contacts); the human Aβ sequence and the AD-causative D7N variant increase the N-terminal β-sheet. In our models, the AD-preventive variant also loads less Zn2+ than the AD-causative. The D7N and other charge-modifying variants of the Aβ sequence may have a direct effect on the solubility of the aggregates, which would be added to the putative charge-reducing effect upon zinc coordination to acidic Aβ residues [11]. An increased hydrophobicity may have an effect on the aggregation behavior of amyloid aggregates as well as in the interaction with biological membranes. These effects may have consequences in the neurotoxicity of amyloid oligomers and metal-mediated neurotoxicity.
4. Conclusions
The time evolution of the structure and other properties of the different Aβ42 protofibril models in response to different simulation conditions described here in the presence of metal ions may be interpreted in the context of the mechanisms of metal accumulation in amyloid plaques and on metal-induced aggregation and cytotoxicity. The regioselective accumulation of cations through the formation of the proposed N-terminal acidic β-sheet domain provides by itself a molecular mechanism that directly resembles the observed accumulation and co-localization of metals within amyloid plaques in brain tissue samples, together with the increased metal affinity for oligomeric forms. The described fold of the N-terminal domain produces confinement of the N-terminal hydrophilic residues together with the attracted cations, which has the potential of increasing the hydrophobicity of the aggregate, an effect further enhanced by the reduction of the Aβ charge that metal coordination to acidic residues would produce. The increase in hydrophobicity has implications in Aβ aggregation and in the interaction with membranes (i.e., affecting their permeability); both consequences may be related to the neurotoxic properties of amyloid aggregates, which are in agreement with the enhanced amyloid aggregation and toxicity promoted by metals. It was also suggested that the distortion of the N-terminal β-sheet structure by metal ions might be related to the change in the pathway of Aβ aggregation. Since the consolidation of β-sheet structure requires oligomerization in Aβ, and the N-terminal β-sheet tends to attract cations, amyloid aggregation may respond to metal dyshomeostasis. These effects may be connected to the high β-sheet content characteristic in amyloid fibrils. The molecular mechanisms here described and contrasted with experimental reports enhance our understanding of the molecular-level processes relevant to the aggregation of Aβ peptides that form AD amyloid plaques, and aid in the comprehension of the role that metal ions such as zinc may play in such processes, particularly, the role that the Aβ N-terminal sequence and its interaction with metals plays.
Data availability
The data encompassing input/output files, scripts, raw analysis data, and visualizations generated throughout this study are openly accessible upon direct inquiry to the authors. We welcome requests for any supplementary materials utilized in our computational investigations.
References
Dementia: a public health priority (2012) https://www.who.int/publications/i/item/dementia-a-public-health-priority. Accessed 27 Dec 2022
Towards a dementia plan: a WHO guide (2018) https://www.who.int/publications/i/item/9789241514132. Accessed 27 Dec 2022
Goedert M, Spillantini MG (2006) A century of Alzheimer’s disease. Science 314(5800):777–781. https://doi.org/10.1126/science.1132814
Hardy JA, Higgins GA (1992) Alzheimer’s disease: the amyloid cascade hypothesis. Science 256(5054):184–185. https://doi.org/10.1126/science.1566067
Walsh DM, Klyubin I, Fadeeva JV, Cullen WK, Anwyl R, Wolfe MS, Rowan MJ, Selkoe DJ (2002) Naturally secreted oligomers of amyloid β protein potently inhibit hippocampal long-term potentiation in vivo. Nature 416(6880):535–539. https://doi.org/10.1038/416535a
Cleary JP, Walsh DM, Hofmeister JJ, Shankar GM, Kuskowski MA, Selkoe DJ, Ashe KH (2005) Natural oligomers of the amyloid-beta protein specifically disrupt cognitive function. Nat Neurosci 8(1):79–84. https://doi.org/10.1038/nn1372
O’Brien RJ, Wong PC (2011) Amyloid precursor protein processing and Alzheimer’s disease. Annu Rev Neurosci 34:185–204. https://doi.org/10.1146/annurev-neuro-061010-113613
Masters CL, Simms G, Weinman NA, Multhaup G, McDonald BL, Beyreuther K (1985) Amyloid plaque core protein in Alzheimer disease and down syndrome. Proc Natl Acad Sci 82(12):4245–4249. https://doi.org/10.1073/pnas.82.12.4245
Mori H, Takio K, Ogawara M, Selkoe DJ (1992) Mass spectrometry of purified amyloid beta protein in Alzheimer’s disease. J Biol Chem 267(24):17082–17086. https://doi.org/10.1016/s0021-9258(18)41896-0
Jarrett JT, Berger EP, Lansbury PT Jr (1993) The carboxy terminus of the beta amyloid protein is critical for the seeding of amyloid formation: implications for the pathogenesis of Alzheimer’s disease. Biochemistry 32(18):4693–4697. https://doi.org/10.1021/bi00069a001
Kepp KP (2012) Bioinorganic chemistry of Alzheimer’s disease. Chem Rev 112(10):5193–5239. https://doi.org/10.1021/cr300009x
Huang X, Cuajungco MP, Atwood CS, Hartshorn MA, Tyndall JD, Hanson GR, Stokes KC, Leopold M, Multhaup G, Goldstein LE, Scarpa RC, Saunders AJ, Lim J, Moir RD, Glabe C, Bowden EF, Masters CL, Fairlie DP, Tanzi RE, Bush AI (1999) Cu(II) potentiation of alzheimer abeta neurotoxicity. Correlation with cell-free hydrogen peroxide production and metal reduction. J Biol Chem 274(52):37111–37116. https://doi.org/10.1074/jbc.274.52.37111
Glenner GG, Wong CW (1984) Alzheimer’s disease: initial report of the purification and characterization of a novel cerebrovascular amyloid protein. Biochem Biophys Res Commun 120(3):885–890. https://doi.org/10.1016/s0006-291x(84)80190-4
Tõugu V, Tiiman A, Palumaa P (2011) Interactions of Zn(II) and Cu(II) ions with Alzheimer’s amyloid-beta peptide. Metal ion binding, contribution to fibrillization and toxicity. Metallomics Integr Biometal Sci 3(3):250–261. https://doi.org/10.1039/c0mt00073f
Sticht H, Bayer P, Willbold D, Dames S, Hilbich C, Beyreuther K, Frank RW, Rösch P (1995) Structure of amyloid A4-(1–40)-peptide of Alzheimer’s disease. Eur J Biochem 233(1):293–298. https://doi.org/10.1111/j.1432-1033.1995.293_1.x
Coles M, Bicknell W, Watson AA, Fairlie DP, Craik DJ (1998) Solution structure of amyloid beta-peptide(1–40) in a water-micelle environment. Is the membrane-spanning domain where we think it is? Biochemistry 37(31):11064–11077. https://doi.org/10.1021/bi972979f
Crescenzi O, Tomaselli S, Guerrini R, Salvadori S, D’Ursi AM, Temussi PA, Picone D (2002) Solution structure of the Alzheimer amyloid beta-peptide (1–42) in an apolar microenvironment. Similarity with a virus fusion domain. Eur J Biochem 269(22):5642–5648. https://doi.org/10.1046/j.1432-1033.2002.03271.x
Hou L, Shao H, Zhang Y, Li H, Menon NK, Neuhaus EB, Brewer JM, Byeon I-JL, Ray DG, Vitek MP, Iwashita T, Makula RA, Przybyla AB, Zagorski MG (2004) Solution NMR studies of the Aβ(1–40) and Aβ(1–42) peptides establish that the Met35 oxidation state affects the mechanism of amyloid formation. J Am Chem Soc 126(7):1992–2005. https://doi.org/10.1021/ja036813f
Simmons LK, May PC, Tomaselli KJ, Rydel RE, Fuson KS, Brigham EF, Wright S, Lieberburg I, Becker GW, Brems DN (1994) Secondary structure of amyloid beta peptide correlates with neurotoxic activity in vitro. Mol Pharmacol 45(3):373–379
Ono K, Condron MM, Teplow DB (2009) Structure–neurotoxicity relationships of amyloid β-protein oligomers. Proc Natl Acad Sci 106(35):14745–14750. https://doi.org/10.1073/pnas.0905127106
Makin OS, Serpell LC (2005) Structures for amyloid fibrils. FEBS J 272(23):5950–5961. https://doi.org/10.1111/j.1742-4658.2005.05025.x
Fändrich M (2007) On the structural definition of amyloid fibrils and other polypeptide aggregates. Cell Mol Life Sci CMLS 64(16):2066–2078. https://doi.org/10.1007/s00018-007-7110-2
Fändrich M, Schmidt M, Grigorieff N (2011) Recent progress in understanding Alzheimer’s β-amyloid structures. Trends Biochem Sci 36(6):338–345. https://doi.org/10.1016/j.tibs.2011.02.002
Gallardo R, Ranson NA, Radford SE (2020) Amyloid structures: much more than just a cross-β fold. Curr Opin Struct Biol 60:7–16. https://doi.org/10.1016/j.sbi.2019.09.001
Tycko R (2011) Solid-state NMR studies of amyloid fibril structure. Annu Rev Phys Chem 62:279–299. https://doi.org/10.1146/annurev-physchem-032210-103539
Goldsbury CS, Wirtz S, Müller SA, Sunderji S, Wicki P, Aebi U, Frey P (2000) Studies on the in vitro assembly of a beta 1–40: implications for the search for a beta fibril formation inhibitors. J Struct Biol 130(2–3):217–231. https://doi.org/10.1006/jsbi.2000.4259
Antzutkin ON, Leapman RD, Balbach JJ, Tycko R (2002) Supramolecular structural constraints on Alzheimer’s β-amyloid fibrils from electron microscopy and solid-state nuclear magnetic resonance. Biochemistry 41(51):15436–15450. https://doi.org/10.1021/bi0204185
Petkova AT, Leapman RD, Guo Z, Yau WM, Mattson MP, Tycko R (2005) Self-propagating, molecular-level polymorphism in Alzheimer’s beta-amyloid fibrils. Science 307(5707):262–265. https://doi.org/10.1126/science.1105850
Paravastu AK, Leapman RD, Yau WM, Tycko R (2008) Molecular structural basis for polymorphism in Alzheimer’s beta-amyloid fibrils. Proc Natl Acad Sci USA 105(47):18349–18354. https://doi.org/10.1073/pnas.0806270105
Lührs T, Ritter C, Adrian M, Riek-Loher D, Bohrmann B, Döbeli H, Schubert D, Riek R (2005) 3D structure of Alzheimer’s amyloid-beta(1–42) fibrils. Proc Natl Acad Sci USA 102(48):17342–17347. https://doi.org/10.1073/pnas.0506723102
Petkova AT, Yau WM, Tycko R (2006) Experimental constraints on quaternary structure in Alzheimer’s beta-amyloid fibrils. Biochemistry 45(2):498–512. https://doi.org/10.1021/bi051952q
Xiao Y, Ma B, McElheny D, Parthasarathy S, Long F, Hoshi M, Nussinov R, Ishii Y (2015) Aβ(1–42) fibril structure illuminates self-recognition and replication of amyloid in Alzheimer’s disease. Nat Struct Mol Biol 22(6):499–505. https://doi.org/10.1038/nsmb.2991
Qiang W, Yau WM, Luo Y, Mattson MP, Tycko R (2012) Antiparallel β-sheet architecture in Iowa-mutant β-amyloid fibrils. Proc Natl Acad Sci USA 109(12):4443–4448. https://doi.org/10.1073/pnas.1111305109
Lu JX, Qiang W, Yau WM, Schwieters CD, Meredith SC, Tycko R (2013) Molecular structure of β-amyloid fibrils in Alzheimer’s disease brain tissue. Cell 154(6):1257–1268. https://doi.org/10.1016/j.cell.2013.08.035
Sgourakis NG, Yau WM, Qiang W (2015) Modeling an in-register, parallel “iowa” aβ fibril structure using solid-state NMR data from labeled samples with rosetta. Structure 23(1):216–227. https://doi.org/10.1016/j.str.2014.10.022
Schütz AK, Vagt T, Huber M, Ovchinnikova OY, Cadalbert R, Wall J, Güntert P, Böckmann A, Glockshuber R, Meier BH (2015) Atomic-resolution three-dimensional structure of amyloid β fibrils bearing the Osaka mutation. Angewandte Chemie (International ed. in English) 54(1):331–335. https://doi.org/10.1002/anie.201408598
Colvin MT, Silvers R, Ni QZ, Can TV, Sergeyev I, Rosay M, Donovan KJ, Michael B, Wall J, Linse S, Griffin RG (2016) Atomic resolution structure of monomorphic Aβ42 amyloid fibrils. J Am Chem Soc 138(30):9663–9674. https://doi.org/10.1021/jacs.6b05129
Wälti MA, Ravotti F, Arai H, Glabe CG, Wall JS, Böckmann A, Güntert P, Meier BH, Riek R (2016) Atomic-resolution structure of a disease-relevant Aβ(1–42) amyloid fibril. Proc Natl Acad Sci USA 113(34):E4976–E4984. https://doi.org/10.1073/pnas.1600749113
Gremer L, Schölzel D, Schenk C, Reinartz E, Labahn J, Ravelli RBG, Tusche M, Lopez-Iglesias C, Hoyer W, Heise H, Willbold D, Schröder GF (2017) Fibril structure of amyloid-β(1–42) by cryo-electron microscopy. Science 358(6359):116–119. https://doi.org/10.1126/science.aao2825
Schmidt M, Rohou A, Lasker K, Yadav JK, Schiene-Fischer C, Fändrich M, Grigorieff N (2015) Peptide dimer structure in an Aβ(1–42) fibril visualized with cryo-EM. Proc Natl Acad Sci 112(38):11858–11863. https://doi.org/10.1073/pnas.1503455112
Kollmer M, Close W, Funk L, Rasmussen J, Bsoul A, Schierhorn A, Schmidt M, Sigurdson CJ, Jucker M, Fändrich M (2019) Cryo-EM structure and polymorphism of Aβ amyloid fibrils purified from Alzheimer’s brain tissue. Nat Commun 10(1):4760. https://doi.org/10.1038/s41467-019-12683-8
Ghosh U, Thurber KR, Yau WM, Tycko R (2021) Molecular structure of a prevalent amyloid-β fibril polymorph from Alzheimer’s disease brain tissue. Proc Natl Acad Sci USA 118(4):e2023089118. https://doi.org/10.1073/pnas.2023089118
Nelson R, Sawaya MR, Balbirnie M, Madsen AØ, Riekel C, Grothe R, Eisenberg D (2005) Structure of the cross-beta spine of amyloid-like fibrils. Nature 435(7043):773–778. https://doi.org/10.1038/nature03680
Sawaya MR, Sambashivan S, Nelson R, Ivanova MI, Sievers SA, Apostol MI, Thompson MJ, Balbirnie M, Wiltzius JJ, McFarlane HT, Madsen AØ, Riekel C, Eisenberg D (2007) Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature 447(7143):453–457. https://doi.org/10.1038/nature05695
Meinhardt J, Sachse C, Hortschansky P, Grigorieff N, Fändrich M (2009) Abeta(1–40) fibril polymorphism implies diverse interaction patterns in amyloid fibrils. J Mol Biol 386(3):869–877. https://doi.org/10.1016/j.jmb.2008.11.005
Fändrich M, Meinhardt J, Grigorieff N (2009) Structural polymorphism of Alzheimer Abeta and other amyloid fibrils. Prion 3(2):89–93. https://doi.org/10.4161/pri.3.2.8859
Miller Y, Ma B, Nussinov R (2010) Polymorphism in Alzheimer Abeta amyloid organization reflects conformational selection in a rugged energy landscape. Chem Rev 110(8):4820–4838. https://doi.org/10.1021/cr900377t
Willbold D, Strodel B, Schröder GF, Hoyer W, Heise H (2021) Amyloid-type protein aggregation and prion-like properties of amyloids. Chem Rev 121(13):8285–8307. https://doi.org/10.1021/acs.chemrev.1c00196
Al Adem K, Lee S (2023) Structural polymorphism and cytotoxicity of brain-derived β-amyloid extracts. Protein Sci A Publ Protein Soc 32(5):e4639. https://doi.org/10.1002/pro.4639
Fitzpatrick AWP, Falcon B, He S, Murzin AG, Murshudov G, Garringer HJ, Crowther RA, Ghetti B, Goedert M, Scheres SHW (2017) Cryo-EM structures of tau filaments from Alzheimer’s disease. Nature 547(7662):185–190. https://doi.org/10.1038/nature23002
Musiek ES, Holtzman DM (2015) Three dimensions of the amyloid hypothesis: time, space and “wingmen.” Nat Neurosci 18(6):800–806. https://doi.org/10.1038/nn.4018
Iliyasu MO, Musa SA, Oladele SB, Iliya AI (2023) Amyloid-beta aggregation implicates multiple pathways in Alzheimer’s disease: understanding the mechanisms. Front Neurosci 17:1081938. https://doi.org/10.3389/fnins.2023.1081938
Lin CJ, Huang HC, Jiang ZF (2010) Cu(II) interaction with amyloid-beta peptide: a review of neuroactive mechanisms in AD brains. Brain Res Bull 82(5–6):235–242. https://doi.org/10.1016/j.brainresbull.2010.06.003
Kozlowski H, Luczkowski M, Remelli M, Valensin D (2012) Copper, zinc and iron in neurodegenerative diseases (Alzheimer’s, Parkinson’s and prion diseases). Coord Chem Rev 256(19–20):2129–2141. https://doi.org/10.1016/j.ccr.2012.03.013
Mezzaroba L, Alfieri DF, Colado Simão AN, Vissoci Reiche EM (2019) The role of zinc, copper, manganese and iron in neurodegenerative diseases. Neurotoxicology 74:230–241. https://doi.org/10.1016/j.neuro.2019.07.007
Garza-Lombó C, Posadas Y, Quintanar L, Gonsebatt ME, Franco R (2018) Neurotoxicity linked to dysfunctional metal ion homeostasis and xenobiotic metal exposure: redox signaling and oxidative stress. Antioxid Redox Signal 28(18):1669–1703. https://doi.org/10.1089/ars.2017.7272
Lovell MA, Robertson JD, Teesdale WJ, Campbell JL, Markesbery WR (1998) Copper, iron and zinc in Alzheimer’s disease senile plaques. J Neurol Sci 158(1):47–52. https://doi.org/10.1016/s0022-510x(98)00092-6
Wang H, Wang M, Wang B, Li M, Chen H, Yu X, Yang K, Chai Z, Zhao Y, Feng W (2012) Immunogold labeling and X-ray fluorescence microscopy reveal enrichment ratios of Cu and Zn, metabolism of APP and amyloid-β plaque formation in a mouse model of Alzheimer's disease. Metallomics Integr Biometal Sci 4(10):1113–1118. https://doi.org/10.1039/c2mt20056b
Miller LM, Wang Q, Telivala TP, Smith RJ, Lanzirotti A, Miklossy J (2006) Synchrotron-based infrared and X-ray imaging shows focalized accumulation of Cu and Zn co-localized with beta-amyloid deposits in Alzheimer’s disease. J Struct Biol 155(1):30–37. https://doi.org/10.1016/j.jsb.2005.09.004
James SA, Churches QI, de Jonge MD, Birchall IE, Streltsov V, McColl G, Adlard PA, Hare DJ (2017) Iron, copper, and zinc concentration in Aβ plaques in the APP/PS1 mouse model of alzheimer’s disease correlates with metal levels in the surrounding neuropil. ACS Chem Neurosci 8(3):629–637. https://doi.org/10.1021/acschemneuro.6b00362
Dong J, Atwood CS, Anderson VE, Siedlak SL, Smith MA, Perry G, Carey PR (2003) Metal binding and oxidation of amyloid-beta within isolated senile plaque cores: Raman microscopic evidence. Biochemistry 42(10):2768–2773. https://doi.org/10.1021/bi0272151
Huang X, Atwood CS, Moir RD, Hartshorn MA, Tanzi RE, Bush AI (2004) Trace metal contamination initiates the apparent auto-aggregation, amyloidosis, and oligomerization of Alzheimer’s Abeta peptides. J Biol Inorganic Chem JBIC A Publ Soc Biol Inorg Chem 9(8):954–960. https://doi.org/10.1007/s00775-004-0602-8
Smith DP, Ciccotosto GD, Tew DJ, Fodero-Tavoletti MT, Johanssen T, Masters CL, Barnham KJ, Cappai R (2007) Concentration dependent Cu2+ induced aggregation and dityrosine formation of the Alzheimer’s disease amyloid-beta peptide. Biochemistry 46(10):2881–2891. https://doi.org/10.1021/bi0620961
Sarell CJ, Wilkinson SR, Viles JH (2010) Substoichiometric levels of Cu2+ ions accelerate the kinetics of fiber formation and promote cell toxicity of amyloid-{beta} from Alzheimer disease. J Biol Chem 285(53):41533–41540. https://doi.org/10.1074/jbc.M110.171355
Weibull MGM, Simonsen S, Oksbjerg CR, Tiwari MK, Hemmingsen L (2019) Effects of Cu(II) on the aggregation of amyloid-β. J Biol Inorg Chem JBIC A Publ Soc Biol Inorg Chem 24(8):1197–1215. https://doi.org/10.1007/s00775-019-01727-5
Deibel MA, Ehmann WD, Markesbery WR (1996) Copper, iron, and zinc imbalances in severely degenerated brain regions in Alzheimer’s disease: possible relation to oxidative stress. J Neurol Sci 143(1–2):137–142. https://doi.org/10.1016/s0022-510x(96)00203-1
Schrag M, Crofton A, Zabel M, Jiffry A, Kirsch D, Dickson A, Mao XW, Vinters HV, Domaille DW, Chang CJ, Kirsch W (2011) Effect of cerebral amyloid angiopathy on brain iron, copper, and zinc in Alzheimer’s disease. J Alzheimer’s Dis JAD 24(1):137–149. https://doi.org/10.3233/JAD-2010-101503
Schrag M, Mueller C, Oyoyo U, Smith MA, Kirsch WM (2011) Iron, zinc and copper in the Alzheimer’s disease brain: a quantitative meta-analysis. Some insight on the influence of citation bias on scientific opinion. Progr Neurobiol 94(3):296–306. https://doi.org/10.1016/j.pneurobio.2011.05.001
Posadas Y, López-Guerrero VE, Arcos-López T, Sayler RI, Sánchez-López C, Segovia J, Perez-Cruz C, Quintanar L (2023) The role of d-block metal ions in neurodegenerative diseases. Compr Inorg Chem III, Third Edn 1–10:575–628. https://doi.org/10.1016/B978-0-12-823144-9.00115-1
Hung YH, Bush AI, Cherny RA (2010) Copper in the brain and Alzheimer’s disease. J Biol Inorg Chem JBIC A Publ Soc Biol Inorg Chem 15(1):61–76. https://doi.org/10.1007/s00775-009-0600-y
Frederickson CJ, Koh JY, Bush AI (2005) The neurobiology of zinc in health and disease. Nat Rev Neurosci 6(6):449–462. https://doi.org/10.1038/nrn1671
Smith DG, Cappai R, Barnham KJ (2007) The redox chemistry of the Alzheimer’s disease amyloid beta peptide. Biochem Biophys Acta 1768(8):1976–1990. https://doi.org/10.1016/j.bbamem.2007.02.002
Meloni G, Faller P, Vasák M (2007) Redox silencing of copper in metal-linked neurodegenerative disorders. J Biol Chem 282(22):16068–16078. https://doi.org/10.1074/jbc.m701357200
Rózga M, Bal W (2010) The Cu(II)/Abeta/human serum albumin model of control mechanism for copper-related amyloid neurotoxicity. Chem Res Toxicol 23(2):298–308. https://doi.org/10.1021/tx900358j
Perrone L, Mothes E, Vignes M, Mockel A, Figueroa C, Miquel MC, Maddelein ML, Faller P (2010) Copper transfer from Cu-Abeta to human serum albumin inhibits aggregation, radical production and reduces Abeta toxicity. Chembiochem A Eur J Chem Biol 11(1):110–118. https://doi.org/10.1002/cbic.200900474
Viles JH (2012) Metal ions and amyloid fiber formation in neurodegenerative diseases. Copper, zinc and iron in Alzheimer’s, Parkinson’s and prion diseases. Coord Chem Rev 256(19–20):2271–2284. https://doi.org/10.1016/j.ccr.2012.05.003
Solomonov I, Korkotian E, Born B, Feldman Y, Bitler A, Rahimi F, Li H, Bitan G, Sagi I (2012) Zn2+-Aβ40 complexes form metastable quasi-spherical oligomers that are cytotoxic to cultured hippocampal neurons. J Biol Chem 287(24):20555–20564. https://doi.org/10.1074/jbc.M112.344036
Sarell CJ, Syme CD, Rigby SE, Viles JH (2009) Copper(II) binding to amyloid-beta fibrils of Alzheimer’s disease reveals a picomolar affinity: stoichiometry and coordination geometry are independent of Abeta oligomeric form. Biochemistry 48(20):4388–4402. https://doi.org/10.1021/bi900254n
Jiang D, Zhang L, Grant GP, Dudzik CG, Chen S, Patel S, Hao Y, Millhauser GL, Zhou F (2013) The elevated copper binding strength of amyloid-β aggregates allows the sequestration of copper from albumin: a pathway to accumulation of copper in senile plaques. Biochemistry 52(3):547–556. https://doi.org/10.1021/bi301053h
Phillips JC, Braun R, Wang W, Gumbart J, Tajkhorshid E, Villa E, Chipot C, Skeel RD, Kalé L, Schulten K (2005) Scalable molecular dynamics with NAMD. J Comput Chem 26(16):1781–1802. https://doi.org/10.1002/jcc.20289
MacKerell AD, Bashford D, Bellott M, Dunbrack RL, Evanseck JD, Field MJ, Fischer S, Gao J, Guo H, Ha S, Joseph-McCarthy D, Kuchnir L, Kuczera K, Lau FT, Mattos C, Michnick S, Ngo T, Nguyen DT, Prodhom B, Reiher WE et al (1998) All-atom empirical potential for molecular modeling and dynamics studies of proteins. J Phys Chem B 102(18):3586–3616. https://doi.org/10.1021/jp973084f
Darden T, York D, Pedersen L (1993) Particle mesh Ewald: an N log(N) method for Ewald sums in large systems. J Chem Phys 98(12):10089–10092. https://doi.org/10.1063/1.464397
Batcho PF, Case DA, Schlick T (2001) Optimized particle-mesh Ewald/multiple-time step integration for molecular dynamics simulations. J Chem Phys 115(9):4003–4018. https://doi.org/10.1063/1.1389854
Ryckaert J-P, Ciccotti G, Berendsen HJC (1977) Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. J Comput Phys 23(3):327–341. https://doi.org/10.1016/0021-9991(77)90098-5
Martyna GJ, Tobias DJ, Klein ML (1994) Constant pressure molecular dynamics algorithms. J Chem Phys 101(5):4177–4189. https://doi.org/10.1063/1.467468
Feller SE, Zhang Y, Pastor RW, Brooks BR (1995) Constant pressure molecular dynamics simulation: the Langevin piston method. J Chem Phys 103(11):4613–4621. https://doi.org/10.1063/1.470648
Humphrey W, Dalke A, Schulten K (1996) VMD: visual molecular dynamics. J Mol Graph 14(1):33–38. https://doi.org/10.1016/0263-7855(96)00018-5
Frishman D, Argos P (1995) Knowledge-based protein secondary structure assignment. Proteins 23(4):566–579. https://doi.org/10.1002/prot.340230412
Nasica-Labouze J, Nguyen PH, Sterpone F, Berthoumieu O, Buchete NV, Coté S, De Simone A, Doig AJ, Faller P, Garcia A, Laio A, Li MS, Melchionna S, Mousseau N, Mu Y, Paravastu A, Pasquali S, Rosenman DJ, Strodel B, Tarus B et al (2015) Amyloid β protein and Alzheimer’s disease: when computer simulations complement experimental studies. Chem Rev 115(9):3518–3563. https://doi.org/10.1021/cr500638n
Ilie IM, Caflisch A (2019) Simulation studies of amyloidogenic polypeptides and their aggregates. Chem Rev 119(12):6956–6993. https://doi.org/10.1021/acs.chemrev.8b00731
Rahman A, Saikia B, Gogoi CR, Baruah A (2022) Advances in the understanding of protein misfolding and aggregation through molecular dynamics simulation. Prog Biophys Mol Biol 175:31–48. https://doi.org/10.1016/j.pbiomolbio.2022.08.007
Söldner CA, Sticht H, Horn AHC (2021) Molecular simulations and Alzheimer’s disease. Syst Med 2:54–70. https://doi.org/10.1016/b978-0-12-801238-3.11541-7
Kahler A, Sticht H, Horn AH (2013) Conformational stability of fibrillar amyloid-beta oligomers via protofilament pair formation—a systematic computational study. PLoS ONE 8(7):e70521. https://doi.org/10.1371/journal.pone.0070521
Buchete NV, Tycko R, Hummer G (2005) Molecular dynamics simulations of Alzheimer’s beta-amyloid protofilaments. J Mol Biol 353(4):804–821. https://doi.org/10.1016/j.jmb.2005.08.066
Buchete NV, Hummer G (2007) Structure and dynamics of parallel beta-sheets, hydrophobic core, and loops in Alzheimer’s A beta fibrils. Biophys J 92(9):3032–3039. https://doi.org/10.1529/biophysj.106.100404
Esposito L, Pedone C, Vitagliano L (2006) Molecular dynamics analyses of cross-beta-spine steric zipper models: beta-sheet twisting and aggregation. Proc Natl Acad Sci USA 103(31):11533–11538. https://doi.org/10.1073/pnas.0602345103
Röhrig UF, Laio A, Tantalo N, Parrinello M, Petronzio R (2006) Stability and structure of oligomers of the Alzheimer peptide Abeta16-22: from the dimer to the 32-mer. Biophys J 91(9):3217–3229. https://doi.org/10.1529/biophysj.106.088542
Zheng J, Jang H, Ma B, Tsai CJ, Nussinov R (2007) Modeling the Alzheimer Abeta17-42 fibril architecture: tight intermolecular sheet-sheet association and intramolecular hydrated cavities. Biophys J 93(9):3046–3057. https://doi.org/10.1529/biophysj.107.110700
Masman MF, Eisel UL, Csizmadia IG, Penke B, Enriz RD, Marrink SJ, Luiten PG (2009) In silico study of full-length amyloid beta 1–42 tri- and penta-oligomers in solution. J Phys Chem B 113(34):11710–11719. https://doi.org/10.1021/jp901057w
Skeby KK, Sørensen J, Schiøtt B (2013) Identification of a common binding mode for imaging agents to amyloid fibrils from molecular dynamics simulations. J Am Chem Soc 135(40):15114–15128. https://doi.org/10.1021/ja405530p
Hou L, Kang I, Marchant RE, Zagorski MG (2002) Methionine 35 oxidation reduces fibril assembly of the amyloid abeta-(1–42) peptide of Alzheimer’s disease. J Biol Chem 277(43):40173–40176. https://doi.org/10.1074/jbc.C200338200
Thirumalai D, Reddy G, Straub JE (2012) Role of water in protein aggregation and amyloid polymorphism. Acc Chem Res 45(1):83–92. https://doi.org/10.1021/ar2000869
Kim YS, Liu L, Axelsen PH, Hochstrasser RM (2009) 2D IR provides evidence for mobile water molecules in beta-amyloid fibrils. Proc Natl Acad Sci USA 106(42):17751–17756. https://doi.org/10.1073/pnas.0909888106
Huraskin D, Horn AHC (2019) Alkali ion influence on structure and stability of fibrillar amyloid-β oligomers. J Mol Model 25(2):37. https://doi.org/10.1007/s00894-018-3920-4
House E, Mold M, Collingwood J, Baldwin A, Goodwin S, Exley C (2009) Copper abolishes the beta-sheet secondary structure of preformed amyloid fibrils of amyloid-beta(42). J Alzheimer’s Dis JAD 18(4):811–817. https://doi.org/10.3233/JAD-2009-1235
Olofsson A, Lindhagen-Persson M, Vestling M, Sauer-Eriksson AE, Ohman A (2009) Quenched hydrogen/deuterium exchange NMR characterization of amyloid-beta peptide aggregates formed in the presence of Cu2+ or Zn2+. FEBS J 276(15):4051–4060. https://doi.org/10.1111/j.1742-4658.2009.07113.x
Márquez M, Blancas-Mejía LM, Campos A, Rojas L, Castañeda-Hernández G, Quintanar L (2014) A bifunctional non-natural tetrapeptide modulates amyloid-beta peptide aggregation in the presence of Cu(ii). Metallomics Integr Biometal Sci 6(12):2189–2192. https://doi.org/10.1039/c4mt00257a
Bertini I, Gonnelli L, Luchinat C, Mao J, Nesi A (2011) A new structural model of Aβ40 fibrils. J Am Chem Soc 133(40):16013–16022. https://doi.org/10.1021/ja2035859
Pan J, Han J, Borchers CH, Konermann L (2011) Conformer-specific hydrogen exchange analysis of Aβ(1–42) oligomers by top-down electron capture dissociation mass spectrometry. Anal Chem 83(13):5386–5393. https://doi.org/10.1021/ac200906v
Haupt C, Leppert J, Rönicke R, Meinhardt J, Yadav JK, Ramachandran R, Ohlenschläger O, Reymann KG, Görlach M, Fändrich M (2012) Structural basis of β-amyloid-dependent synaptic dysfunctions. Angewandte Chemie (International ed. in English) 51(7):1576–1579. https://doi.org/10.1002/anie.201105638
Parthasarathy S, Long F, Miller Y, Xiao Y, McElheny D, Thurber K, Ma B, Nussinov R, Ishii Y (2011) Molecular-level examination of Cu2+ binding structure for amyloid fibrils of 40-residue Alzheimer’s β by solid-state NMR spectroscopy. J Am Chem Soc 133(10):3390–3400. https://doi.org/10.1021/ja1072178
Faller P, Hureau C (2009) Bioinorganic chemistry of copper and zinc ions coordinated to amyloid-β peptide. Dalton Trans 7:1080–1094. https://doi.org/10.1039/b813398k
Drew SC, Barnham KJ (2011) The heterogeneous nature of Cu2+ interactions with Alzheimer’s amyloid-β peptide. Acc Chem Res 44(11):1146–1155. https://doi.org/10.1021/ar200014u
Gunderson WA, Hernández-Guzmán J, Karr JW, Sun L, Szalai VA, Warncke K (2012) Local structure and global patterning of Cu2+ binding in fibrillar amyloid-β [Aβ(1–40)] protein. J Am Chem Soc 134(44):18330–18337. https://doi.org/10.1021/ja306946q
Hureau C (2012) Coordination of redox active metal ions to the amyloid precursor protein and to amyloid-β peptides involved in Alzheimer disease. Part 1: an overview. Coord Chem Rev 256(19–20):2164–2174. https://doi.org/10.1016/j.ccr.2012.03.037
Faller P, Hureau C, La Penna G (2014) Metal ions and intrinsically disordered proteins and peptides: from Cu/Zn amyloid-β to general principles. Acc Chem Res 47(8):2252–2259. https://doi.org/10.1021/ar400293h
Drew SC, Noble CJ, Masters CL, Hanson GR, Barnham KJ (2009) Pleomorphic copper coordination by Alzheimer’s disease amyloid-beta peptide. J Am Chem Soc 131(3):1195–1207. https://doi.org/10.1021/ja808073b
Zirah S, Kozin SA, Mazur AK, Blond A, Cheminant M, Ségalas-Milazzo I, Debey P, Rebuffat S (2006) Structural changes of region 1–16 of the Alzheimer disease amyloid beta-peptide upon zinc binding and in vitro aging. J Biol Chem 281(4):2151–2161. https://doi.org/10.1074/jbc.M504454200
Hureau C, Coppel Y, Dorlet P, Solari PL, Sayen S, Guillon E, Sabater L, Faller P (2009) Deprotonation of the Asp1-Ala2 peptide bond induces modification of the dynamic copper(II) environment in the amyloid-beta peptide near physiological pH. Angewandte Chemie (International ed. in English) 48(50):9522–9525. https://doi.org/10.1002/anie.200904512
Sóvágó I, Osz K (2006) Metal ion selectivity of oligopeptides. Dalton transactions (Cambridge, England: 2003) (32):3841–3854. https://doi.org/10.1039/b607515k
Gomez-Castro CZ, Vela A, Quintanar L, Grande-Aztatzi R, Mineva T, Goursot A (2014) Insights into the oxygen-based ligand of the low pH component of the Cu(2+)-amyloid-β complex. J Phys Chem B 118(34):10052–10064. https://doi.org/10.1021/jp5047529
Murray B, Sharma B, Belfort G (2017) N-terminal hypothesis for Alzheimer’s disease. ACS Chem Neurosci 8(3):432–434. https://doi.org/10.1021/acschemneuro.7b00037
Sarkar B, Mithu VS, Chandra B, Mandal A, Chandrakesan M, Bhowmik D, Madhu PK, Maiti S (2014) Significant structural differences between transient amyloid-β oligomers and less-toxic fibrils in regions known to harbor familial Alzheimer’s mutations. Angewandte Chemie (International ed. in English) 53(27):6888–6892. https://doi.org/10.1002/anie.201402636
Johnstone EM, Chaney MO, Norris FH, Pascual R, Little SP (1991) Conservation of the sequence of the Alzheimer’s disease amyloid peptide in dog, polar bear and five other mammals by cross-species polymerase chain reaction analysis. Brain Res Mol Brain Res 10(4):299–305. https://doi.org/10.1016/0169-328x(91)90088-f
Atwood CS, Moir RD, Huang X, Scarpa RC, Bacarra NM, Romano DM, Hartshorn MA, Tanzi RE, Bush AI (1998) Dramatic aggregation of Alzheimer abeta by Cu(II) is induced by conditions representing physiological acidosis. J Biol Chem 273(21):12817–12826. https://doi.org/10.1074/jbc.273.21.12817
Liu ST, Howlett G, Barrow CJ (1999) Histidine-13 is a crucial residue in the zinc ion-induced aggregation of the A beta peptide of Alzheimer’s disease. Biochemistry 38(29):9373–9378. https://doi.org/10.1021/bi990205o
Istrate AN, Tsvetkov PO, Mantsyzov AB, Kulikova AA, Kozin SA, Makarov AA, Polshakov VI (2012) NMR solution structure of rat aβ(1–16): toward understanding the mechanism of rats’ resistance to Alzheimer’s disease. Biophys J 102(1):136–143. https://doi.org/10.1016/j.bpj.2011.11.4006
Wakutani Y, Watanabe K, Adachi Y, Wada-Isoe K, Urakami K, Ninomiya H, Saido TC, Hashimoto T, Iwatsubo T, Nakashima K (2004) Novel amyloid precursor protein gene missense mutation (D678N) in probable familial Alzheimer’s disease. J Neurol Neurosurg Psychiatry 75(7):1039–1042. https://doi.org/10.1136/jnnp.2003.010611
Hori Y, Hashimoto T, Wakutani Y, Urakami K, Nakashima K, Condron MM, Tsubuki S, Saido TC, Teplow DB, Iwatsubo T (2007) The Tottori (D7N) and English (H6R) familial Alzheimer disease mutations accelerate Abeta fibril formation without increasing protofibril formation. J Biol Chem 282(7):4916–4923. https://doi.org/10.1074/jbc.M608220200
Ono K, Condron MM, Teplow DB (2010) Effects of the English (H6R) and Tottori (D7N) familial Alzheimer disease mutations on amyloid beta-protein assembly and toxicity. J Biol Chem 285(30):23186–23197. https://doi.org/10.1074/jbc.M109.086496
Xu L, Chen Y, Wang X (2014) Dual effects of familial Alzheimer’s disease mutations (D7H, D7N, and H6R) on amyloid β peptide: correlation dynamics and zinc binding. Proteins 82(12):3286–3297. https://doi.org/10.1002/prot.24669
Sharma B, Ranganathan SV, Belfort G (2018) Weaker N-terminal interactions for the protective over the causative Aβ peptide dimer mutants. ACS Chem Neurosci 9(6):1247–1253. https://doi.org/10.1021/acschemneuro.7b00412
Acknowledgements
To Conahcyt, grant number 221134 to LQ and PhD scholarship to CZGC. The authors gratefully acknowledge the computing time granted by LANCAD and Conahcyt on the supercomputer Yoltla/Miztli/Xiuhcoatl at LSVP UAM-Iztapalapa/DGTIC UNAM/CGSTIC Cinvestav.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gómez-Castro, C.Z., Quintanar, L. & Vela, A. An N-terminal acidic β-sheet domain is responsible for the metal-accumulation properties of amyloid-β protofibrils: a molecular dynamics study. J Biol Inorg Chem 29, 407–425 (2024). https://doi.org/10.1007/s00775-024-02061-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00775-024-02061-1