Abstract
Thirteen root extract samples of four goldenrod (Solidago) species present in Europe were investigated by hyphenated high-performance thin-layer chromatography (HPTLC). Only S. virgaurea is native, whereas S. gigantea, S. canadensis, and S. graminifolia have been introduced from North America. The bioactive zones in the Aliivibrio fischeri bioautogram were identified as polyacetylenes, labdane diterpenes, or clerodane diterpenes by HPTLC coupled to high-resolution mass spectrometry, exploiting the two interfaces, heated electrospray ionization, and direct analysis in real time. Principal component analysis of the obtained bioprofiles enabled the discrimination of the Solidago species. Furthermore, chemometrics pointed to the discriminative components, the main bioactive markers of the species: Z,Z-matricaria ester from S. virgaurea, solidagenone from S. canadensis, solidagoic acid A, and a dialdehyde clerodane diterpene from S. gigantea, and Z-dehydromatricaria ester from S. graminifolia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
High-performance thin-layer chromatography (HPTLC) fingerprints of plant extracts can be used and compared to discriminate species, subspecies, varieties, or chemotypes [1,2,3]. Multiple HPTLC fingerprints can be obtained from the same separation by documenting at ultraviolet (UV for absorbance; FLD for fluorescence detection) and visible light (Vis), after performing chemical derivatizations or (bio)assays that visualize the chemical profiles or bioprofiles, respectively. The combination of multi-imaging HPTLC with pattern recognition using different chemometric tools enables the rapid fingerprinting and classification of the samples [4,5,6]. However, the usefulness of image processing for distinguishing samples based on their bioprofiles or biochemical profiles (effect-directed classification) has been demonstrated in only a few cases [7,8,9].
HPTLC combined especially with high-resolution mass spectrometry (HRMS) is an efficient tool for the characterization and identification of the selected biomarker compounds [10, 11]. The most widespread elution head-based coupling interface (elution head of 2 mm × 4 mm) is installed between the pump for eluent delivery and the MS [12, 13]. It enables a targeted MS analysis of zones of interest, while ambient desorption–ionization-based techniques, such as desorption electrospray ionization and direct analysis in real time (DART), allow scanning of the whole plate, however, only from an aliquot of each sample on the surface [14, 15].
The herbaceous perennial goldenrods (Solidago, Asteraceae) with yellow flowers often grow up to 2 m high [16]. The only goldenrod species native in Europe is S. virgaurea (European goldenrod). However, three further invasive, alien goldenrod plants are also widespread in Europe, i. e., S. gigantea (giant goldenrod), S. canadensis (Canadian goldenrod), and S. graminifolia (also known as Euthamia graminifolia, grass-leaved goldenrod). They were introduced about 250 years ago as ornamentals from North America and have become remarkably successful competitive invaders of abandoned fields, forest edges, and river banks in most European countries. Several goldenrods are known as medicinal plants and the aerial part of S. virgaurea and those of S. canadensis and S. gigantea are listed in the European Pharmacopoeia as Latin names Solidaginis virgaureae herba and Solidaginis herba, respectively. Aerial parts are used in traditional medicine in the treatment of urinary complaints and as anti-inflammatory agents [17,18,19]. The decoction of Solidago roots is used by Indians against diseases of the urinary tract, diabetes, fever, pain, and inflammation [20].
The fully flowered plants can be distinguished based on their aerial parts. Only S. canadensis and S. gigantea are very similar, but their distinctive mark is their hairy and bald stems, respectively. Recently, HPTLC profiling of root extracts via post-chromatographic derivatization with vanillin-sulfuric acid reagent has been demonstrated as an efficient discrimination tool, which was confirmed by principal component analysis (PCA). Several antimicrobial root components of these species were identified using a non-target HPTLC–bioassay screening followed by compound isolation and highly targeted characterization. Among them were poly-acetylene matricaria esters from S. virgaurea, dehydromatricaria esters from S. graminifolia, three labdane diterpenes from S. canadensis, and eight clerodane diterpenes from S. gigantea [21,22,23,24].
This study investigated the effect-directed classification of the four Solidago species in Europe based on their HPTLC–A. fischeri bioprofiles from root extracts as well as the assignment and identification of the responsible discriminative bioactive compounds.
2 Experimentals
2.1 Materials
HPTLC plates silica gel 60 F254 and methanol (MS grade) were purchased from Merck (Darmstadt, Germany). Further solvents (analytical grade) were from Th. Geyer (Renningen, Germany) or Sigma-Aldrich (Steinheim, Germany). The bioluminescent marine bacterium Aliivibrio fischeri (DSM 7151) was from Leibniz Institute DSMZ, German Collection of Microorganisms and Cell Cultures (Berlin, Germany). The culture medium was prepared as described [25].
2.2 Sample preparation
Roots of 13 goldenrod plants of four species (S. virgaurea, S. gigantea, S. canadensis, and S. graminifolia) were collected between 2014 and 2017 from various parts of Hungary (Table 1). Dried and ground (Bosch MKM6000, Stuttgart, Germany) samples were macerated in ethanol (150 mg/mL) for 24 h. The filtered crude extract was used after dilution (1:10 with ethanol).
2.3 HPTLC–bioassay
Extracts (1–5 µL/band) were applied as 6-mm bands with a 9-mm track distance and 8-mm distance from the bottom onto the HPTLC plate (ATS4, CAMAG, Muttenz, Switzerland). HPTLC separation was carried out with n-hexane‒isopropyl acetate‒acetone (16:3:1, V/V) in an unsaturated Twin Trough Chamber (20 cm × 10 cm, CAMAG) up to a migration distance of 70 mm, which took about 20 min [22, 23]. After development, the plate was dried in a cold stream of air, documented with a TLC Visualizer Documentation System (CAMAG). The antibacterial A. fischeri bioassay was performed as described [25]. Shortly, the dried chromatogram was immersed into the cell suspension of the bioluminescent A. fischeri and immediately recorded (50 s exposure time, BioLuminizer CAMAG). Active zones were indicated as dark zones on the bioluminescent background (depicted as greyscale image).
2.4 HPTLC–HRMS
The plates were pre-washed (methanol‒water, 4:1, V/V) and dried (100 °C, 20 min). For HPTLC–HESI–HRMS experiments, the zones were online-eluted with MS-grade methanol at a flow rate of 0.1 mL/min (Ultimate LPG-3400 XRS, Dionex Softron, Germering, Germany) via the TLC–MS Interface (CAMAG) or PlateExpress Interface (Advion, Ithaca, NY, USA), both including an oval elution head (4 mm × 2 mm), into the heated electrospray ionization probe (HESI-II) of the Q Exactive Plus hybrid quadrupole-orbitrap mass spectrometer operated by Xcalibur 3.0.63 software (Thermo Fisher Scientific, Bremen, Germany). The spray voltage was 3.5 kV, the capillary temperature was 270 °C, the sheath gas was 20 arbitrary units, and the auxiliary nitrogen gas 10 arbitrary units. A full scan was recorded in the range of m/z 50–750 with a resolution of 280,000 in both negative and positive ionization modes. The automatic gain control target (AGCT) was 3 × 106, and the maximum injection time (IT) was 100 ms. For HPTLC–DART–HRMS scanning, a modified DART system [14] (IonSense, Saugus, MA, USA) was coupled to HRMS with the following settings: helium gas (99.999%) with flow rate 3.0 L/min and temperature 500 °C; scanning speed 0.2 mm/s; initial needle voltage 4 kV; grid voltage 50 V; positive ionization mode; full scan in the range of m/z 100 − 750 with a resolution of 35,000; AGCT 5 × 104; maximum IT 50 ms.
2.5 Multivariate image analysis of goldenrod root extracts
The open-source rTLC software for multivariate data analysis of planar chromatograms (http://shinyapps.ernaehrung.uni-giessen.de/rtlc/) [6] was applied. The HPTLC bioautogram image after the A. fischeri bioassay was uploaded in the software. Based on unsupervised pattern recognition, PCA was performed for the categorization of the 13 samples. The grey channel signals and hRF 30 − 90 were selected as variables.
3 Results and discussion
The compounds of the 13 root samples from four Solidago species were separated by HPTLC with n-hexane‒isopropyl acetate‒acetone (16:3:1, V/V). After the A. fischeri bioassay, several bioactive zones were observed (Fig. 1). As this mobile phase had already been applied for the separation of S. gigantea [23], S. virgaurea [22], and S. canadensis [22], the bioactive compounds could easily be identified by hRF value comparison. Thus, zones Sc1 (hRF 45) and Sc2 (hRF 53) from S. canadensis were assigned as solidagenone and the mixture of R- and S-presolidagenone, respectively. Zones Sv1 (hRF 83) and Sv2 (hRF 92) from S. virgaurea were identified as Z,Z- and E,Z-matricaria esters, respectively. Six bioactive zones from S. gigantea corresponded to eight clerodane diterpenes. Zone Sg1 (hRF 9) represented kingidiol, Sg2 (hRF 31) an epoxy-hemiacetal, Sg3 (hRF 46) three di-terpenes, a hautriwaic lactone, an alcohol, and a hemiacetal, Sg4 (hRF 51) a dialdehyde, Sg5 (hRF 58) solidagoic acid A, and Sg6 (hRF 37) solidagoic acid B.
HPTLC–A. fischeri bioautograms of 13 root extract samples (Table 1) from the four Solidago species S. gigantea (Sg), S. canadensis (Sc), S. virgaurea (Sv), and S. graminifolia (Sgr), developed with n-hexane‒isopropyl acetate‒acetone (16:3:1, V/V).
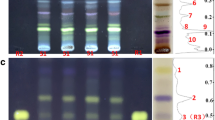
Based on the latest literature [24], dehydromatricaria esters were proposed to be the main bioactive zones Sgr1 (hRF 75) and Sgr2 (hRF 87) of S. graminifolia. To verify these preliminary assignments, HPTLC–HRMS experiments were carried out. The HPTLC–HESI–HRMS analysis of compound zones Sgr1 and Sgr2 provided very similar HRMS spectra in the positive ionization mode. For zone Sgr1 (Fig. 2), the sodium adduct ions were dominant at m/z 195.0412, corresponding to the molecular formula C11H8O2Na+ (calculated m/z 195.0422). Besides, the sodium adduct of the dimer at m/z 367.0952 (C22H16O4Na+, calculated m/z 367.0946) and the protonated molecule at m/z 173.0593 (C11H9O2+, calculated m/z 173.0603) were detected.
Similarly, the same mass signals were recorded for both compound zones by HPTLC–DART+–HRMS scanning (Fig. 3), namely the protonated molecule at m/z 173.0601 for Sgr1 and m/z 173.0602 for Sgr2. These results confirmed the preliminary assignments of zones Sgr1 and Sgr2 as Z- and E-dehydromatricaria esters, respectively.
The open-source rTLC software was used to perform PCA on the signals obtained from the 13 separated root extracts in the HPTLC–A. fischeri bioautogram (Fig. 4). It was evaluated whether it was possible to distinguish the four Solidago species and to point to the most discriminative bioactive compounds according to the loading plot. The first three PCs accounted for 99.33% of the total variance, in which PC1, PC2, and PC3 referred to 95.59%, 2.71%, and 1.03%, respectively. PC2 showed the best separation among the species, while PC3 enabled their discrimination as well. Thus, the samples were divided into four distinct groups, confirming that PCA allowed the classification of the S. canadensis, S. gigantea, S. graminifolia, and S. virgaurea species based on their root extracts collected in wintertime from their persistent rhizomes. PC2 and PC3 were highly influenced by the compounds Sv1, Sgr1, Sc1, Sg5, and Sg4, which were considered as the most important antibacterial compounds for the discrimination of the goldenrod species.
4 Conclusions
The HPTLC–A. fischeri bioautograms of 13 goldenrod root extracts combined with chemometrics allowed us to distinguish the four Solidago species present in Europe. PCA and HPTLC–HRMS revealed the main biomarkers of the species responsible for their distinction. HPTLC–HRMS was shown as a straightforward hyphenation for the characterization and identification of the bioactive compounds.
References
Birk CD, Provensi G, Gosmann G, Reginatto FH, Schenkel EP (2005) TLC fingerprint of flavonoids and saponins from Passiflora species. J Liq Chromatogr Relat Technol 28:2285–2291. https://doi.org/10.1081/JLC-200064212
Shawky E, El Sohafy SM (2020) Untargeted and targeted chemical profiling for efficacy-directed discrimination of Hedera helix L. subspecies using HPTLC‒image analysis and HPTLC/MS. Ind Crops Prod 145:111980. https://doi.org/10.1016/j.indcrop.2019.111980
Baj T, Sieniawska E, Ludwiczuk A, Widelski J, Kiełtyka-Dadasiewicz A, Skalicka-Woźniak K, Głowniak K (2017) Thin-layer chromatography–fingerprint, antioxidant activity, and gas chromatography–mass spectrometry profiling of several Origanum L. species. JPC - J Planar Chromatogr - Mod TLC 30:386–391. https://doi.org/10.1556/1006.2017.30.5.7
Shawky E, Abou El Kheir RM (2018) Rapid discrimination of different Apiaceae species based on HPTLC fingerprints and targeted flavonoids determination using multivariate image analysis. Phytochem Anal 29:452–462. https://doi.org/10.1002/pca.2749
Ristivojević PM, Morlock GE (2016) The influence of preprocessing methods on multivariate image analysis in high-performance thin-layer chromatography fingerprinting. JPC - J Planar Chromatogr - Mod TLC 29:310–317. https://doi.org/10.1556/1006.2016.29.4.10
Fichou D, Ristivojević P, Morlock GE (2016) Proof-of-principle of rTLC, an open-source software developed for image evaluation and multivariate analysis of planar chromatograms. Anal Chem 88:12494–12501. https://doi.org/10.1021/acs.analchem.6b04017
Simion IM, Casoni D, Sârbu C (2018) Characterization and classification of medicinal plants according to their antioxidant profile estimated by thin layer chromatography assisted by chemometric expertise. J Liq Chromatogr Relat Technol 41:342–348. https://doi.org/10.1080/10826076.2018.1448692
Ristivojević PM, Morlock GE (2018) Effect-directed classification of biological, biochemical and chemical profiles of 50 German beers. Food Chem 260:344–353. https://doi.org/10.1016/j.foodchem.2018.03.127
Ristivojević PM, Tahir A, Malfent F, Opsenica DM, Rollinger JM (2019) High-performance thin-layer chromatography/bioautography and liquid chromatography–mass spectrometry hyphenated with chemometrics for the quality assessment of Morus alba samples. J Chromatogr A 1594:190–198. https://doi.org/10.1016/j.chroma.2019.02.006
Morlock G, Schwack W (2010) Coupling of planar chromatography to mass spectrometry. TrAC Trends Anal Chem 29:1157–1171. https://doi.org/10.1016/j.trac.2010.07.010
Morlock GE (2021) High-performance thin-layer chromatography combined with effect-directed assays and high-resolution mass spectrometry as an emerging hyphenated technology: a tutorial review. Anal Chim Acta 1180:338644. https://doi.org/10.1016/j.aca.2021.338644
Luftmann H, Aranda M, Morlock GE (2007) Automated interface for hyphenation of planar chromatography with mass spectrometry. Rapid Commun Mass Spectrom 21:3772–3776. https://doi.org/10.1002/rcm.3276
Morlock GE, Brett N (2015) Correct assignment of lipophilic dye mixtures? A case study for high-performance thin-layer chromatography–mass spectrometry and performance data for the TLC–MS Interface. J Chromatogr A 1390:103–111. https://doi.org/10.1016/j.chroma.2015.02.011
Häbe TT, Morlock GE (2016) Improved desorption/ionization and ion transmission in surface scanning by direct analysis in real time mass spectrometry. Rapid Commun Mass Spectrom 30:321–332. https://doi.org/10.1002/rcm.7434
Van Berkel GJ, Kertesz V (2006) Automated sampling and imaging of analytes separated on thin-layer chromatography plates using desorption electrospray ionization mass spectrometry. Anal Chem 78:4938–4944. https://doi.org/10.1021/ac060690a
Szymura M, Szymura TH (2016) Interactions between alien goldenrods (Solidago and Euthamia species) and comparison with native species in Central Europe. Flora - Morphol Distrib Funct Ecol Plants 218:51–61. https://doi.org/10.1016/j.flora.2015.11.009
Fursenco C, Calalb T, Uncu L, Dinu M, Ancuceanu R (2020) Solidago virgaurea L.: a review of its ethnomedicinal uses, phytochemistry, and pharmacological activities. Biomolecules 10:1619. https://doi.org/10.3390/biom10121619
Toiu A, Vlase L, Vodnar DC, Gheldiu A-M, Oniga I (2019) Solidago graminifolia L. Salisb. (Asteraceae) as a valuable source of bioactive polyphenols: HPLC profile, in vitro antioxidant and antimicrobial potential. Molecules 24:2666. https://doi.org/10.3390/molecules24142666
Kołodziej B (2011) Antibacterial and antimutagenic activity of extracts aboveground parts of three Solidago species: Solidago virgaurea L., Solidago canadensis L. and Solidago gigantea Ait. J Med Plants Res 5:6770–6779. https://doi.org/10.5897/JMPR11.1098
Wangensteen H, Phan TT, Rise F, Halvorsen TG, Malterud KE (2012) New labdane diterpenes from Solidago canadensis. Nat Prod Res 26:1348–1354. https://doi.org/10.1080/14786419.2011.565005
Móricz ÁM, Ott PG, Häbe TT, Darcsi A, Böszörményi A, Alberti Á, Krüzselyi D, Csontos P, Béni S, Morlock GE (2016) Effect-directed discovery of bioactive compounds followed by highly targeted characterization, isolation and identification, exemplarily shown for Solidago virgaurea. Anal Chem 88:8202–8209. https://doi.org/10.1021/acs.analchem.6b02007
Móricz ÁM, Jamshidi-Aidji M, Krüzselyi D, Darcsi A, Böszörményi A, Csontos P, Béni S, Ott PG, Morlock GE (2020) Distinction and valorization of 30 root extracts of five goldenrod (Solidago) species. J Chromatogr A 1611:460602. https://doi.org/10.1016/j.chroma.2019.460602
Móricz ÁM, Krüzselyi D, Ott PG, Garádi Z, Béni S, Morlock GE, Bakonyi J (2021) Bioactive clerodane diterpenes of giant goldenrod (Solidago gigantea Ait.) root extract. J Chromatogr A 1635:461727. https://doi.org/10.1016/j.chroma.2020.461727
Krüzselyi D, Bakonyi J, Ott PG, Darcsi A, Csontos P, Morlock GE, Móricz ÁM (2021) Goldenrod root compounds active against crop pathogenic fungi. J Agric Food Chem 69:12686–12694. https://doi.org/10.1021/acs.jafc.1c03676
Krüger S, Urmann O, Morlock GE (2013) Development of a planar chromatographic method for quantitation of anthocyanes in pomace, feed, juice and wine. J Chromatogr A 1289:105–118. https://doi.org/10.1016/j.chroma.2013.03.005
Acknowledgements
This work was funded by the National Research, Development and Innovation Office of Hungary (NKFIH K128921). Á.M. Móricz thanks the OECD for the scholarship JA00092484 that allowed her to stay at JLU Giessen. Instrumentation was partially funded by the Deutsche Forschungsgemeinschaft (INST 162/471-1 FUGG; INST 162/536-1 FUGG). We are grateful to Péter Csontos (Institute for Soil Sciences and Agricultural Chemistry, Centre for Agricultural Research, Hungary) and András Schmotzer (Directorate of Bükk National Park, Eger, Hungary) for collecting Solidago samples in Hungary, to Jana Posslova (Botanical Garden of Masaryk University, Brno, Czech Republic) for sending S. graminifolia seed, and to The National Botanical Garden, Vácrátót (Hungary) to provide S. graminifolia plants. The authors thank Tim Häbe and Imanuel Yüce for support regarding HRMS.
Funding
Open access funding provided by ELKH Centre for Agricultural Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Móricz, Á.M., Ott, P.G., Krüzselyi, D. et al. High-performance thin-layer chromatography–direct bioautography combined with chemometrics for the distinction of goldenrod species. JPC-J Planar Chromat 35, 339–344 (2022). https://doi.org/10.1007/s00764-022-00159-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00764-022-00159-3