Abstract
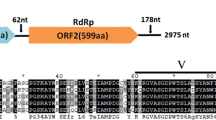
The complete genome of a novel mycovirus, Colletotrichum curcumae narnavirus 1 (CcNV1), derived from the phytopathogenic fungus Colletotrichum curcumae strain 780-2T, was sequenced and analyzed. The full sequence of CcNV1 is 3,374 nucleotides in length and contains a single large open reading frame (ORF) encoding an RNA-dependent RNA polymerase (RdRp) of 1,087 amino acids with a molecular mass of 124.2 kDa that shares the closest similarity with that of Monilinia narnavirus H (53.02% identity). RdRp phylogeny analysis showed that CcNV1 is a new member of the proposed genus "Betanarnavirus" within the family Narnaviridae. This is the first report of a novel narnavirus infecting the phytopathogenic fungus C. curcumae, the causal agent of leaf blight of Curcuma wenyujin.


Similar content being viewed by others
References
Ghabrial SA, Jiang D, Nibert ML, Suzuki N (2015) 50-plus years of fungal viruses. Virology 479:356–368
Ghabrial SA, Suzuki N (2009) Viruses of plant pathogenic fungi. Annu Rev Phytopathol 47(1):353–384
Pearson MN, Beever RE, Boine B, Arthur K (2009) Mycoviruses of filamentous fungi and their relevance to plant pathology. Mol Plant Pathol 10:115–128
Xie J, Jiang D (2014) New insights into mycoviruses and exploration for the biological control of crop fungal diseases. Annu Rev Phytopathol 52(1):45–68
Ghabrial SA, Suzuki N (2009) Viruses of plant pathogenic fungi. Annu Rev Phytopathol 47:353–384
Suzuki A, Kobayashi F, Abe M, Uchiumi T, Higashi S (2001) Cloning and expression of a down-regulated gene (TrEnodDR1) of white clover responded by the nod genes derived from Rhizobium leguminosarum bv. trifolii strain 4S. Gene 266(1–2):77–84
Kanematsu S, Sasaki A, Onoue M, Oikawa Y, Ito T (2010) Extending the fungal host range of a partitivirus and a mycoreovirus from Rosellinia necatrix by inoculation of protoplasts with virus particles. Phytopathology 100(9):922–930
Hillman BI, Cai G (2013) The family Narnaviridae: simplest of RNA viruses. Adv Virus Res 86:149–176
Wickner RB, Fujimura T, Esteban R (2013) Viruses and prions of Saccharomyces cerevisiae. Adv Virus Res 86:1–36
Vainio EJ (2019) Mitoviruses in the conifer root rot pathogens Heterobasidion annosum and H. parviporum. Virus Res 271:1–5
Ong JWL, Li H, Sivasithamparam K, Dixon KW, Jones MGK, Wylie SJ (2016) Novel Endorna-like viruses, including three with two open reading frames, challenge the membership criteria and taxonomy of the Endornaviridae. Virology 499:203–211
Zhang R, Hisano S, Tani A, Kondo H, Kanematsu S, Suzuki N (2016) A capsidless ssRNA virus hosted by an unrelated dsRNA virus. Nat Microbiol 1:15001
Jia H, Dong K, Zhou L, Wang G, Hong N, Jiang D (2017) A dsRNA virus with filamentous viral particles. Nat Commun 8:168
Wickner RB, Fujimura T, Esteban R (2013) Viruses and prions of Saccharomyces cerevisiae. Adv Virus Res 86:1–36
Klumpp K, Ruigrok RW, Baudin F (1997) Roles of the influenza virus polymerase and nucleoprotein in forming a functional RNP structure. EMBO J 16(6):1248–1257
Area E, Martín-Benito J, Gastaminza P, Torreira E, Valpuesta JM, Carrascosa JL, Ortín J (2004) 3D structure of the influenza virus polymerase complex: localization of subunit domains. Proc Natl Acad Sci USA 101(1):308–313
Göertz GP, Miesen P, Overheul GJ, Van Rij RP, Van Oers MM, Pijlman GP (2019) Mosquito small RNA responses to West Nile and insect-specific virus infections in Aedes and Culex mosquito cells. Viruses 11(3):E271
Wang Y, Liu S, Zhu HJ, Zhong J (2019) Molecular characterization of a novel mycovirus from the plant pathogenic fungus Colletotrichum gloeosporioides. Arch Virol 164(11):2859–2863
Xu X, Hai D, Li J, Huang F, Wang Y (2022) Molecular characterization of a novel penoulivirus from the phytopathogenic fungus Colletotrichum camelliae. Arch Virol 167(2):641–644
Cao J, Xie F, Zhang Z, Zhu H, Zhou X (2022) Molecular characterization of a novel polymycovirus identified in the phytopathogenic fungus Colletotrichum gloeosporioides. Arch Virol 167(12):2805–2810
Guo J, Zhu JZ, Zhou XY, Zhong J, Li CH, Zhang ZG, Zhu HJ (2019) A novel ourmia-like mycovirus isolated from the plant pathogenic fungus Colletotrichum gloeosporioides. Arch Virol 164(10):2631–2635
Wang Y, Liu S, Zhu HJ, Zhong J (2019) Molecular characterization of a novel mycovirus from the plant pathogenic fungus Colletotrichum gloeosporioides. Arch Virol 164(11):2859–2863
Zhong J, Chen D, Lei XH, Zhu HJ, Zhu JZ, Gao BD (2014) Detection and characterization of a novel Gammapartitivirus in the phytopathogenic fungus Colletotrichum acutatum strain HNZJ001. Virus Res 190:104–109
Jia H, Dong K, Zhou L, Wang GP, Hong N, Jiang DH, Xu WX (2017) A dsRNA virus with filamentous viral particles. Nat Commun 8(1):168
Zhai L, Zhang M, Hong N, Hong N, Xiao F, Fu M, Xiang J, Wang G (2018) Identification and characterization of a novel hepta-segmented dsRNA virus from the phytopathogenic fungus Colletotrichum fructicola. Front Microbiol 9:754
Wang H, Liu H, Zhou Q (2021) The complete genome sequence of a new mitovirus from the phytopathogenic fungus Colletotrichum higginsianum. Arch Virol 166(5):1481–1484
Liu H, Wang H, Lu X, Xiao C, Peng B, Zhou Q (2021) Molecular characterization of a novel single-stranded RNA virus, ChRV1, isolated from the plant-pathogenic fungus Colletotrichum higginsianum. Arch Virol 166(6):1805–1809
Lima JS, Figueiredo JG, Gomes RG, Stringari D, Goulin EH, Adamoski D, Kava-Cordeiro V, Galli-Terasawa LV, Glienke C (2012) Genetic Diversity of Colletotrichum spp. an Endophytic Fungi in a Medicinal Plant. Brazilian Pepper Tree. Isrn Microbiology,215716
Komatsu K, Katayama Y, Omatsu T, Mizutani T, Fukuhara T, Kodama M, Arie T, Teraoka T, Moriyama H (2016) Genome sequence of a novel mitovirus identified in the phytopathogenic fungus Alternaria arborescens. Arch Virol 161:2627–2631
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23(21):2947–2948
Kamer G, Argos P (1984) Primary structural comparison of RNA-dependent RNA polymerase from plant, animal, and bacterial viruses. Nucleic Acid Res 12(18):7269–7282
Koonin EV, Choi GH, Nuss DL, Shapira R, Carrington JC (1991) Evidence for common ancestry of a chestnut blight hypovirulence associated double-stranded RNA and a group of positive-strand RNA plant viruses. Proc Natl Acad Sci USA 88:10647–10651
Ng KK, Arnold JJ, Cameron CE (2008) Structure-function relationships among RNA-dependent RNA polymerases. Curr Top Microbiol Immunol 320(1):137–156
Funding
This study was supported by the National Natural Science Foundation of China (no. 32260648), the Hainan Province Key R&D Project (no. ZDYF2022XDNY242), and the Scientific Research Foundation for Advanced Talents, Hainan University (no. KYQD(ZR)1873).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with animals or human participants performed by any of the authors.
Additional information
Communicated by Ioly Kotta-Loizou.
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yujia Fu and Tian Wang contributed equally to this work.
Electronic Supplementary Material
Below is the link to the electronic supplementary material
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, Y., Wang, T., Zhou, S. et al. A novel narnavirus isolated from Colletotrichum curcumae strain 780-2T. Arch Virol 168, 226 (2023). https://doi.org/10.1007/s00705-023-05847-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00705-023-05847-x