Abstract
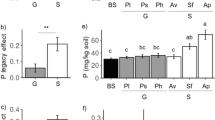
Interaction with orchid mycorrhizal fungi (OMF) is essential to all members of the Orchidaceae, yet we know little about whether or how OMF abundances in substrates shape orchid populations. While root-associated OMF diversity is catalogued frequently, technological constraints have impeded the assessments of OMF communities in substrates until recently, thereby limiting the ability to link OMF communities in a habitat to population responses. Furthermore, there is some evidence that edaphic and microclimatic conditions impact OMF in soil, yet we lack an understanding of the coupled influences of abiotic environment and OMF structure on orchid population dynamics. To discover the linkages between abiotic environment, OMF community structure, and population size, we characterized the microclimatic conditions, soil physicochemistry, and OMF communities hosted by roots and soil across large and small populations of a terrestrial orchid endemic to California Floristic Province in North America. By using high-throughput sequencing of the ITS2 region of nrDNA amplified from root and soil DNAs, we determined that both roots and soil of larger populations, which were high in phosphorus but low in zinc, organic matter, and silt, were dominated by Tulasnellaceae OTUs. In comparison, roots and soil from smaller populations of the orchid hosted higher relative abundances of the Ceratobasidiaceae. In this multiyear, range-wide study that simultaneously measured habitat environmental conditions, and soil and root OMF communities, our results suggest that soil chemistry is clearly linked to soil and root OMF communities, which then likely alter and shape orchid populations.




Similar content being viewed by others
Data accessibility
The raw sequences from root and soil samples were deposited in the NCBI SRA database under BioProject PRJNA609930.
References
Ackerman JD, Lauri RK (2018) Piperia cooperi, in Jepson Flora Project (eds.) Jepson eFlora
Alvarado P, Teixeira MDM, Andrews L, Fernandez A, Santander G, Doyle A, Perez M, Yegres F, Barker BM (2018) Detection of Coccidioides posadasii from xerophytic environments in Venezuela reveals risk of naturally acquired coccidioidomycosis infections. Emerg Microbes Infect 7:1–13.
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46. https://doi.org/10.1046/j.1442-9993.2001.01070.x
Aronesty E (2013) Comparison of sequencing utility programs. Open Bioinforma J 7:1–8. https://doi.org/10.2174/1875036201307010001
Bateman RM, Hollingsworth PM, Preston J, Yi-Bo L, Pridgeon AM, Chase MW (2003) Molecular phylogenetics and evolution of Orchidinae and selected Habenariinae (Orchidaceae). Bot J Linn Soc 142:1–40. https://doi.org/10.1046/j.1095-8339.2003.00157.x
Berry D, Ben Mahfoudh K, Wagner M, Loy A, Ben MK (2011) Barcoded primers used in multiplex amplicon pyrosequencing bias amplification. Appl Environ Microbiol 77:612–612. https://doi.org/10.1128/AEM.05220-11
Bidartondo MI, Read DJ (2008) Fungal specificity bottlenecks during orchid germination and development. Mol Ecol 17:3707–3716. https://doi.org/10.1111/j.1365-294X.2008.03848.x
Bunch WD, Cowden CC, Wurzburger N, Shefferson RP (2013) Geography and soil chemistry drive the distribution of fungal associations in lady’s slipper orchid, Cypripedium acaule. Botany 91:850–856. https://doi.org/10.1139/cjb-2013-0079
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pẽa AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335. https://doi.org/10.1038/nmeth.f.303
Coates F, Lunt ID, Tremblay RL (2006) Effects of disturbance on population dynamics of the threatened orchid Prasophyllum correctum D.L. Jones and implications for grassland management in south-eastern Australia. Biol Conserv 129:59–69. https://doi.org/10.1016/j.biocon.2005.06.037
Csardi G, Nepusz T (2006) The igraph software package for complex network research. InterJournal Complex Syst 1695:1–9
Davis NM, Proctor D, Holmes SP, Relman DA, Callahan BJ (2017) Simple statistical identification and removal of contaminant sequences in marker-gene and metagenomics data. Microbiome 6:226. https://doi.org/10.1101/221499
Dearnaley JDW, Martos F, Selosse MA (2012) Orchid mycorrhizas: molecular ecology, physiology, evolution and conservation aspects. In: Fungal Associations, 2nd Edition
Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C (2011) T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res 39(suppl_2:W13–W17. https://doi.org/10.1093/nar/gkr245
Diez JM (2007) Hierarchical patterns of symbiotic orchid germination linked to adult proximity and environmental gradients. J Ecol 95:159–170. https://doi.org/10.1111/j.1365-2745.2006.01194.x
Dray S, Blanchet G, Borcard D, Guenard G, Jombart T, Larocque G, Legendre P, Madi N, Wagne H (2016) Adespatial: multivariate multiscale spatial analysis. R Packag version 00–9
Dressler RL (1993) Phylogeny and classification of the orchid family. Cambridge University Press
Edgar R (2016) SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv 074161. https://doi.org/10.1101/074161
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Fiedler PL, Keever ME, Grewell BJ, Partridge DJ (2007) Rare plants in the Golden Gate Estuary (California): the relationship between scale and understanding. Aust J Bot 55:206–220. https://doi.org/10.1071/BT06069
Girlanda M, Segreto R, Cafasso D, Liebel HT, Rodda M, Ercole E, Cozzolino S, Gebauer G, Perotto S (2011) Photosynthetic Mediterranean meadow orchids feature partial mycoheterotrophy and specific mycorrhizal associations1. Am J Bot 98:1148–1163. https://doi.org/10.3732/ajb.1000486
Hill MO (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54:427–432. https://doi.org/10.2307/1934352
Hsieh TC, Ma KH, Chao A (2016) iNEXT: an R package for rarefaction and extrapolation of species diversity (Hill numbers). Methods Ecol Evol 7:1451–1456. https://doi.org/10.1111/2041-210X.12613
Hutchings MJ (2010) The population biology of the early spider orchid Ophrys sphegodes Mill. III. Demography over three decades J Ecol 98:867–878. https://doi.org/10.1111/j.1365-2745.2010.01661.x
Jacquemyn H, Brys R, Jongejans E (2010) Size-dependent flowering and costs of reproduction affect population dynamics in a tuberous perennial woodland orchid. J Ecol 98:1204–1215. https://doi.org/10.1111/j.1365-2745.2010.01697.x
Jacquemyn H, Waud M, Merckx VSFT, Lievens B, Brys R (2015) Mycorrhizal diversity, seed germination and long-term changes in population size across nine populations of the terrestrial orchid Neottia ovata. Mol Ecol 24:3269–3280. https://doi.org/10.1111/mec.13236
Jost L (2006) Entropy and diversity. Oikos 113:363–375. https://doi.org/10.1111/j.2006.0030-1299.14714.x
Kaur J, Andrews L, Sharma J (2019) High specificity of a rare terrestrial orchid toward a rare fungus within the North American tallgrass prairie. Fungal Biol 123:895–904. https://doi.org/10.1016/j.funbio.2019.09.010
Kaur J, Poff KE, Sharma J (2017) A rare temperate terrestrial orchid selects similar Tulasnella taxa in ex situ and in situ environments. Plant Ecol 219:1–11. https://doi.org/10.1007/s11258-017-0776-0
Koljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M (2014) Towards a unified paradigm for sequence-based identification of fungi. Mol Ecol 22:5271–5277. https://doi.org/10.1111/mec.12481
Kuntal BK, Dutta A, Mande SS (2016) CompNet: A GUI based tool for comparison of multiple biological interaction networks. BMC Bioinformatics 17:185. https://doi.org/10.1186/s12859-016-1013-x
Leitao RP, Zuanon J, Williams SE, Baraloto C, Fortunel C, Mendonc FP, Mouillot D (2016) Rare species contribute disproportionately to the functional structure of species assemblages. Proc R Soc B 283:20160084
Mandal S, Van Treuren W, White RA, Eggesbø M, Knight R, Peddada SD (2015) Analysis of composition of microbiomes: a novel method for studying microbial composition. Microb Ecol Health Dis 26:27663. https://doi.org/10.3402/mehd.v26.27663
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17:10–12
Mattila E, Kuitunen MT (2000) Nutrient versus pollination limitation in Platanthera bifolia and Dactylorhiza incarnata (Orchidaceae). Oikos 89:360–366. https://doi.org/10.1034/j.1600-0706.2000.890217.x
McCormick MK, Jacquemyn H (2014) What constrains the distribution of orchid populations? New Phytol 202:392–400. https://doi.org/10.1111/nph.12639
McCormick MK, Lee Taylor D, Juhaszova K, Burnett RK, Whigham DF, O’Neill JP (2012) Limitations on orchid recruitment: not a simple picture. Mol Ecol 21:1511–1523. https://doi.org/10.1111/j.1365-294X.2012.05468.x
McCormick MK, Taylor DL, Whigham DF, Burnett RK (2016) Germination patterns in three terrestrial orchids relate to abundance of mycorrhizal fungi. J Ecol 104:744–754. https://doi.org/10.1111/1365-2745.12556
McCormick MK, Whigham DF, Canchani-Viruet A (2018) Mycorrhizal fungi affect orchid distribution and population dynamics. New Phytol 219:1207–1215. https://doi.org/10.1111/nph.15223
Mccormick MK, Whigham DF, O’Neill JP, Becker JJ, Sarah W, Rasmussen HN, Bruns And TD, Taylor DL (2009) Abundance and distribution of Corallorhiza odontorhiza reflect variations in climate and ectomycorrhizae. Ecol Monogr. https://doi.org/10.1890/08-0729.1
McCormick MK, Whigham DF, Sloan D, O’Malley K, Hodkinson B (2006) Orchid-fungus fidelity: a marriage meant to last? Ecology 87:903–911
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Mujica MI, Saez N, Cisternas M, Manzano M, Armesto JJ, Pérez F (2016) Relationship between soil nutrients and mycorrhizal associations of two Bipinnula species (Orchidaceae) from central Chile. Ann Bot 118:149–158. https://doi.org/10.1093/aob/mcw082
Newman MEJ (2006) Modularity and community structure in networks. Proc Natl Acad Sci 103:8577–8582. https://doi.org/10.1073/pnas.0601602103
Nurfadilah S, Swarts ND, Dixon KW, Lambers H, Merritt DJ (2013) Variation in nutrient-acquisition patterns by mycorrhizal fungi of rare and common orchids explains diversification in a global biodiversity hotspot. Ann Bot 111:1233–1241. https://doi.org/10.1093/aob/mct064
Oja J, Vahtra J, Bahram M, Kohout P, Kull T, Rannap R, Kõljalg U, Tedersoo L (2017) Local-scale spatial structure and community composition of orchid mycorrhizal fungi in semi-natural grasslands. Mycorrhiza 27:355–367. https://doi.org/10.1007/s00572-016-0755-7
Otero JT, Bayman P, Ackerman JD (2005) Variation in mycorrhizal performance in the epiphytic orchid Tolumnia variegata in vitro: the potential for natural selection. Evol Ecol 19:29–43. https://doi.org/10.1007/s10682-004-5441-0
Pandey M, Sharma J, Taylor DL, Yadon VL (2013) A narrowly endemic photosynthetic orchid is non-specific in its mycorrhizal associations. Mol Ecol 22:2341–2354. https://doi.org/10.1111/mec.12249
R Core Team (2019) R: a language and environment for statistical computing. R Found Stat Comput Vienna, Austria URL https://wwwR-project.org/
Rambaut A (2012) FigTree v1.4. https://tree.bio.ed.ac.uk/software/figtree/
Rock-Blake R, McCormick MK, Brooks HEA, Jones CS, Whigham DF (2017) Symbiont abundance can affect host plant population dynamics. Am J Bot 104:72–82. https://doi.org/10.3732/ajb.1600334
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rohland N, Reich D (2012) Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res 22:939–946. https://doi.org/10.1101/gr.128124.111
Sharma J, Zettler L, Sambeek J (2003) A survey of mycobionts of federally threatened Platanthera praeclara (Orchidaceae). Symbiosis 34:145–155
Shefferson RP, Kull T, Hutchings MJ, Selosse MA, Jacquemyn H, Kellett KM, Menges ES, Primack RB, Tuomi J, Alahuhta K, Hurskainen S, Alexander HM, Anderson DS, Brys R, Brzosko E, Dostálik S, Gregg K, Ipser Z, Jäkäläniemi A, Jersáková J, Dean Kettle W, Mccormick MK, Mendoza A, Miller MT, Moen A, Øien DI, Püttsepp Ü, Roy M, Sather N, Sletvold N, Štípková Z, Tali K, Warren RJ, Whigham DF (2018) Drivers of vegetative dormancy across herbaceous perennial plant species. Ecol Lett 21:724–733. https://doi.org/10.1111/ele.12940
Shefferson RP, Warren RJ, Pulliam RH (2014) Life-history costs make perfect sprouting maladaptive in two herbaceous perennials. J Ecol 102:1318–1328. https://doi.org/10.1111/1365-2745.12281
Stamatakis A (2006) RAxML-VI-HPC: Maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. https://doi.org/10.1093/bioinformatics/btl446
Stanton-Geddes J, Tiffin P, Shaw RG (2012) Role of climate and competitors in limiting fitness across range edges of an annual plant. Ecology 93:1604–1613
Swarts ND, Dixon KW (2009a) Perspectives on orchid conservation in botanic gardens. Trends Plant Sci 14:590–598. https://doi.org/10.1016/j.tplants.2009.07.008
Swarts ND, Dixon KW (2009b) Terrestrial orchid conservation in the age of extinction. Ann Bot 104:543–556. https://doi.org/10.1093/aob/mcp025
Swarts ND, Sinclair EA, Francis A, Dixon KW (2010) Ecological specialization in mycorrhizal symbiosis leads to rarity in an endangered orchid. Mol Ecol 19:3226–3242. https://doi.org/10.1111/j.1365-294X.2010.04736.x
Taylor DL, Walters WA, Lennon NJ, Bochicchio J, Krohn A, Caporaso JG, Pennanen T (2016) Accurate estimation of fungal diversity and abundance through improved lineage-specific primers optimized for Illumina amplicon sequencing. Appl Environ Microbiol 82:7217–7226. https://doi.org/10.1128/AEM.02576-16
Voyron S, Ercole E, Ghignone S, Perotto S, Girlanda M (2017) Fine-scale spatial distribution of orchid mycorrhizal fungi in the soil of host-rich grasslands. New Phytol 213:1428–1439. https://doi.org/10.1111/nph.14286
Wamelink GWW, Goedhart PW, Frissel JY (2014) Why some plant species are rare. PLoS One 9. https://doi.org/10.1371/journal.pone.0102674
Waud M, Brys R, Van Landuyt W, Lievens B, Jacquemyn H (2017) Mycorrhizal specificity does not limit the distribution of an endangered orchid species. Mol Ecol 26:1687–1701. https://doi.org/10.1111/mec.14014
Waud M, Busschaert P, Ruyters S, Jacquemyn H, Lievens B (2014) Impact of primer choice on characterization of orchid mycorrhizal communities using 454 pyrosequencing. Mol Ecol Resour 14:679–699. https://doi.org/10.1111/1755-0998.12229
Acknowledgments
We specifically thank Lisa Markovchick, Andrew Wastell, Michelle Maley, Chris Gillespie, and Jose Roldan for their assistance with project management at NBPL. We also thank San Diego County Orchid Society and California Native Plant Society for providing additional research funding, and Jardin Botanico San Quintin (Josue Campos), Santa Catalina Island Conservancy, and Cleveland National Forest for providing research permits and for assisting in the field.
Funding
We received support from the Naval Base Point Loma (NBPL; Office of Naval Research, U.S. Department of Defense; Cooperative Agreement Nos. N62473-13-2-4907 and N62473-15-2-0011).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kaur, J., Phillips, C. & Sharma, J. Host population size is linked to orchid mycorrhizal fungal communities in roots and soil, which are shaped by microenvironment. Mycorrhiza 31, 17–30 (2021). https://doi.org/10.1007/s00572-020-00993-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-020-00993-5