Abstract
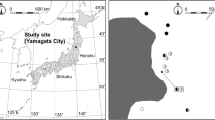
Mycorrhizal symbiosis often displays low specificity, except for mycoheterotrophic plants that obtain carbon from their mycorrhizal fungi and often have higher specificity to certain fungal taxa. Partially mycoheterotrophic (or mixotrophic, MX) plant species tend to have a larger diversity of fungal partners, e.g., in the genus Pyrola (Monotropoideae, Ericaceae). Preliminary evidence however showed that the Japanese Pyrola japonica has preference for russulacean fungi based on direct sequencing of the fungal internal transcribed spacer (ITS) region from a single site. The present study challenges this conclusion using (1) sampling of P. japonica in different Japanese regions and forest types and (2) fungal identification by ITS cloning. Plants were sampled from eight sites in three regions, in one of which the fungal community on tree ectomycorrhizal (ECM) tips surrounding P. japonica was also analyzed. In all, 1512 clone sequences were obtained successfully from 35 P. japonica plants and 137 sequences from ECM communities. These sequences were collectively divided into 74 molecular operational taxonomic units (MOTUs) (51 and 33 MOTUs, respectively). MOTUs from P. japonica involved 36 ECM taxa (96 % of all clones), and 17 of these were Russula spp. (76.2 % of all clones), which colonized 33 of the 35 sampled plants. The MOTU composition significantly differed between P. japonica and ECM tips, although shared species represented 26.3 % of the ECM tips community in abundance. This suggests that P. japonica has a preference for russulacean fungi.




Similar content being viewed by others
References
Abadie JC, Puttsepp U, Gebauer G, Faccio A, Bonfante P, Selosse MA (2006) Cephalanthera longifolia (Neottieae, Orchidaceae) is mixotrophic: a comparative study between green and nonphotosynthetic individuals. Can J Bot 84:1462–1477
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bellino A, Alfani A, Selosse MA, Guerrieri R, Borghetti M, Baldantoni D (2014) Nutritional regulation in mixotrophic plants: new insights from Limodorum abortivum. Oecol 175:875–885
Bidartondo M, Kretzer AM, Bruns TD (2000) High root concentration and uneven ectomycorrhizal diversity near Sarcodes sanguinea (Ericaceae): a cheater that stimulates its victims? Am J Bot 87:1783–1788
Bidartondo MI, Bruns TD (2001) Extreme specificity in epiparasitic Monotropoideae (Ericaceae): widespread phylogenetic and geographical structure. Mol Ecol 10:2285–2295
Björkman E (1960) Monotropa hypopitys L.—an epiparasite on tree roots. Physiol Plantarum 13:308–327
Colwell RK (2013) EstimateS: statistical estimation of species richness and shared species from samples, ver. 9.1. http://purl.oclc.org/estimates. Accessed 15th January 2016
Dowie NJ, Hemenway JJ, Miller SL (2012) Identity, genetic lineages and putative hybrids of an obligate mycobiont associated with the mycoheterotrophic plant Pterospora andromedea in the south-central Rocky mountains. Fungal Ecol 5:137–146
Dray S, Legendre P, Peres-Neto PR (2006) Spatial modelling: a comprehensive framework for principal coordinate analysis of neighbour matrices (PCNM). Ecol Model 196:483–493
Gebauer G, Meyer M (2003) 15N and 13C natural abundance of autotrophic and myco-heterotrophic orchids provides insight into nitrogen and carbon gain from fungal association. New Phytol 160:209–223
Girlanda M, Selosse MA, Cafasso D, Brilli F, Delfine S, Fabbian R, Ghignone S, Pinelli P, Segreto R, Loreto F, Cozzolino S, Perotto S (2006) Inefficient photosynthesis in the Mediterranean orchid Limodorum abortivum is mirrored by specific association to ectomycorrhizal Russulaceae. Mol Ecol 15:491–504
Gonneau C, Jersáková J, Tredern E, Till-Bottraud I, Sauve M, Saarinen K, Roy M, Hájek T, Selosse MA (2014) Photosynthesis in perennial mixotrophic Epipactis spp. (Orchidaceae) contributes more to shoot and fruit biomass than to hypogeous survival. J Ecol 102:1183–1194
Hashimoto Y, Fukukawa S, Kunishi A, Suga H, Richard F, Sauve M, Selosse MA (2012) Mycoheterotrophic germination of Pyrola asarifolia dust seeds reveals convergences with germination in orchids. New Phytol 195:620–630
van der Heijden MGA, Martin F, Selosse M-A, Sanders IR (2015) Mycorrhizal ecology and evolution: the past, the present, and the future. New Phytol 205:1406–1423
Henry C, Raivoarisoa JF, Razafimamonjy A, Ramanankierana H, Andrianaivomahefa P, Selosse MA, Ducousso M (2015) Asteropeia mcphersonii, a potential mycorrhizal facilitator for ecological restoration in Madagascar wet tropical rainforests. For Ecol Manag 358:202–211
Hynson NA, Bruns TD (2009) Evidence of a myco-heterotroph in the plant family Ericaceae that lacks mycorrhizal specificity. Proc Roy Soc Lond B 276:4053–4059
Hynson NA, Preiss K, Gebauer G, Bruns TD (2009) Isotopic evidence of full and partial myco-heterotrophy in the plant tribe Pyroleae (Ericaceae). New Phytol 182:719–726
Hynson NA, Madsen TP, Selosse MA, Adam IKU, Ogura-Tsujita Y, Roy M, Gebauer G (2013) The physiological ecology of mycoheterotrophy. In: Mycohetertrophy. Merckx VSFT, Springer, New York, pp 297–342
Hynson NA, Bidartondo MI, Read DJ (2015) Are there geographic mosaics of mycorrhizal specificity and partial mycoheterotrophy? A case study in Moneses uniflora (Ericaceae). New Phytol 208:1003–1007
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Kennedy AH, Taylor DL, Watson LE (2011) Mycorrhizal specificity in the fully mycoheterotrophic Hexalectris Raf. (Orchidaceae: Epidendroideae). Mol Ecol 20:1303–1316
Liu ZW, Zhou J, Liu ED, Peng H (2010) A molecular phylogeny and a new classification of Pyrola (Pyroleae, Ericaceae). Taxon 59:1690–1700
Massicotte HB, Melville LH, Tackaberry LE, Peterson RL (2008) A comparative study of mycorrhizas in several genera of Pyroleae (Ericaceae) from western Canada. Botany 86:610–622
Matsuda Y, Hijii N (2004) Ectomycorrhizal fungal communities in an Abies firma forest, with special reference to ectomycorrhizal associations between seedlings and mature trees. Can J Bot 82:822–829
Matsuda Y, Okochi S, Katayama T, Yamada A, Ito SI (2011) Mycorrhizal fungi associated with Monotropastrum humile (Ericaceae) in central Japan. Mycorrhiza 21:569–576
Matsuda Y, Shimizu S, Mori M, Ito SI, Selosse MA (2012) Seasonal and environmental changes of mycorrhizal associations and heterotrophy levels in mixotrophic Pyrola japonica (Ericaceae) growing under different light environments. Am J Bot 99:1177–1188
Miyamoto Y, Nakano T, Hattori M, Nara K (2014) The mid-domain effect in ectomycorrhizal fungi: range overlap along an elevation gradient on Mount Fuji, Japan. ISME J 8:1739–1746
VSFT Merckx (2013) Mycoheterotrophy: an introduction. In: VSFT Merckx (ed) Mycoheterotrophy. Springer, New York, pp 1–17
VSFT Merckx, Freudenstein JV, Kissling J, Christenhusz MJM, Stotler RE, Crandall-Stotler B, Wickett N, Rudall PJ, Maas-van de Kamer H, Maas PJM (2013) Taxonomy and classification. In: VSFT Merckx (ed) Mycoheterotrophy. Springer, New York, pp 19–101
Nara K (2015) The role of ectomycorrhizal networks in seedling establishment and primary succession. In: Horton TR (ed) Mycorrhizal networks. Springer, Berlin Heidelberg, pp 177–201
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2013) Vegan: community ecology package. R package version 2.0-10. http://CRAN.R-project.org/package=vegan
Peterson RL, Massicotte HB, Melville LH (2004) Mycorrhizas: anatomy and cell biology. NRC Research Press, Ottawa
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, http://www.R-project.org/
Robertson DC, Robertson JA (1985) Ultrastructural aspects of Pyrola mycorrhizae. Can J Bot 63:1089–1098
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Selosse MA, Richard F, He X, Simard S (2006) Mycorrhizal networks: les liaisons dangereuses. Trends Ecol Evol 11:621–628
Selosse MA, Roy M (2009) Green plants that feed on fungi: facts and questions about mixotrophy. Trends Plant Sci 14:64–70
Selosse MA, Bocayuva MF, Kasuya MCM, Courty PE (2016) Mixotrophy in mycorrhizal plants: extracting C from mycorrhizal networks. In Martin F (ed) Mycorrhizal ecology. Springer, Berlin-Heidelberg.
Simard SW, Beiler KJ, Bingham MA, Deslippe JR, Philip LJ, Teste FP (2012) Mycorrhizal networks: mechanisms, ecology and modelling. Fungal Biol Rev 26:39–60
Smith SE, Read DJ (2008) Mycorrhizal symbiosis, 3rd edn. Academic, London
Takahashi H (1991) Distribution of Japanese Pyrola and Orthilia, and a distribution pattern with a gap in the central to southern Tohoku district. Acta Phytotax Geobot 42:23–43
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Taylor LD, Bruns TD (1999) Population, habitat and genetic correlates of mycorrhizal specialization in the ‘cheating’orchids Corallorhiza maculata and C. mertensiana. Mol Ecol 8:1719–1732
Tedersoo L, Pellet P, Kõljalg U, Selosse MA (2007) Parallel evolutionary paths to mycoheterotrophy in understorey Ericaceae and Orchidaceae: ecological evidence for mixotrophy in Pyroleae. Oecol 151:206–217
Tedersoo L, Jairus T, Horton BM, Abarenkov K, Suvi T, Saar I, Kõljalg U (2008) Strong host preference of ectomycorrhizal fungi in a Tasmanian wet sclerophyll forest as revealed by DNA barcoding and taxon-specific primers. New Phytol 180:479–490
Thomson D, Henry R (1995) Single-step protocol for preparation of plant tissue for analysis by PCR. Biotechniques 19:394–397
Toftegaard T, Iason GR, Alexander IJ, Rosendahl S, Taylor AFS (2010) The threatened plant intermediate wintergreen (Pyrola media) associates with a wide range of biotrophic fungi in native Scottish pine woods. Biodivers Conserv 19:3963–3971
Vincenot L, Tedersoo L, Richard F, Horcine H, Kõljalg U, Selosse MA (2008) Fungal associates of Pyrola rotundifolia, a mixotrophic Ericaceae, from two Estonian boreal forests. Mycorrhiza 19:15–25
Vincenot L, Nara K, Sthultz C, Labbe J, Dubois M-P, Tedersoo L, Martin F, Selosse MA (2012) Extensive gene flow over Europe and possible speciation over Eurasia in the ectomycorrhizal basidiomycete Laccaria amethystina complex. Mol Ecol 21:281–299
Waterman RJ, Klooster MR, Hentrich H, Bidartondo MI (2013) Species interactions of mycoheterotrophic plants: specialization and its potential consequences. In: VSFT Merckx (ed) Mycoheterotrophy. Springer, New York, pp 267–296
Zimmer K, Hynson NA, Gebauer G, Allen EB, Allen MF, Read DJ (2007) Wide geographical and ecological distribution of nitrogen and carbon gains from fungi in pyroloids and monotropoids (Ericaceae) and in orchids. New Phytol 175:166–175
Acknowledgments
We thank Mr. S Iio for habitat information regarding the Hashirida shrine, Iyama shrine, and Makiba Mizunosaka West Hill and the Aichi prefecture government office for permission to access the sites. We also thank T. Chikada, Life Science Research Center, Center for Molecular Biology and Genetics, Mie University, for helping out with DNA analyses, the members of the Laboratory of Forest Pathology and Mycology, Graduate School of Bioresources, Mie University, for their help in field sampling, and D. Marsh for correcting our English. This study was partly supported by KAKENHI (22688011, 23658124, 25304026) to YM and the Fondation de France (fond Ars Cuttoli & Appel) to M-AS.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Table S1
BLAST results for ITS consensus sequences of each MOTU. (DOC 92 kb)
Table S2
Fungal taxa mycorrhizal on Pyrola japonica. Putative trophic status: ECM, ectomycorrhizal; ERM, ericoid; END, endophytic; SAP, saprotrophic; NEM, nematophagous. Asterisk indicates MOTUs that were detected from Pyrola japonica and tree roots but not identical soil cores. (DOC 130 kb)
Table S3
Fungal taxa detected from Pyrola japonica and tree roots in the same cores. MOTUs shared between Pyrola japonica and ECM tips in one core are enclosed. Hosts, Py: Pyrola japonica, Q: Quercus glauca, P: Pinus densiflora, C: Castanopsis cuspidata. (DOC 130 kb)
Fig. S1
Images of each study site: (a) Aichi, (b) Mie and (c) Kyoto, (d) flowering Pyrola japonica, (e) root fragment, (f) light and (g) fluorescent microscopic views of P. japonica roots made by hand cross section. Arrows in (e), (f) and (g) indicate hyphal coils, and no fungal mantles were formed on epidermal cells. Double arrowheads in (e) and (f) indicate plant cells that contained no hyphal coils. Bars 1 mm for (e) and (f) and 100 μm for (g). (DOC 728 kb)
Fig. S2
Rarefaction curves of (a) MOTUs in Pyrola japonica collected at all study sites (white circle), Aichi (black square) and Mie (black up-pointing triangle) and (b) Simpson’s diversity of clones derived from each plant collected. Plants collected at Kyoto were not analyzed owing to the lack of replicates (n = 3). Plant ID is shown with the combination of site codes, overstory trees (P for Pinus and Q for Quercus) and replicates. (DOC 81 kb)
Fig. S3
A non-metric multidimensional scaling plot of the mycorrhizal community of Pyrola japonica. Clone abundance data of each site were used for ordination with Horn dissimilarity index (a). White up-pointing triangle, Aichi sites A1 to A3; black square, Mie site M1 to M4; white circle, Kyoto unique site K1. Grouping according to region was not significant (PERMANOVA, p = 0.1). Eight variables were tested for significant correlation with mycorrhizal community dissimilarities (b). Significantly correlated variables are shown in the plot as arrows. The length of the arrow is proportional to the strength of the correlation with the ordination (*p < 0.05). PCNM is the principal coordinates of neighborhood matrix and see for “Materials and methods” in details. (DOC 61 kb)
Fig. S4
Rarefaction curves of MOTUs in Pyrola japonica (white circle) and its surrounding ectomycorrhizal roots (black circle) collected in Aichi. (DOC 43 kb)
Fig. S5
Neighbor-joining phylogeny of russulacean MOTUs recovered in this study. Bootstrap tests (1000 replicates) larger than 90 % are shown next to the branches. The tree is drawn to scale, with branch lengths in the same units as those of the evolutionary distances used to infer the phylogenetic tree. The evolutionary distances were computed using the Maximum Composite Likelihood method. MOTUs detected in this study are in bold, and when they were derived from ECM tips denoted by an ‘e’ following the MOTU names. (DOC 103 kb)
Rights and permissions
About this article
Cite this article
Uesugi, T., Nakano, M., Selosse, MA. et al. Pyrola japonica, a partially mycoheterotrophic Ericaceae, has mycorrhizal preference for russulacean fungi in central Japan. Mycorrhiza 26, 819–829 (2016). https://doi.org/10.1007/s00572-016-0715-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00572-016-0715-2