Abstract
Key Message
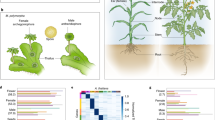
The first record of gene expression during seed development within the Nymphaeales provides evidence for a variety of biological processes, including dynamic epigenetic patterning during sexual reproduction in the water lily Nymphaea thermarum.
Abstract
Studies of gene expression during seed development have been performed for a growing collection of species from a phylogenetically broad sampling of flowering plants (angiosperms). However, angiosperm lineages whose origins predate the divergence of monocots and eudicots have been largely overlooked. In order to provide a new resource for understanding the early evolution of seed development in flowering plants, we sequenced transcriptomes of whole ovules and seeds from three key stages of reproductive development in the waterlily Nymphaea thermarum, an experimentally tractable member of the Nymphaeales. We first explore patterns of gene expression, beginning with mature ovules and continuing through fertilization into early- and mid-stages of seed development. We find patterns of gene expression that corroborate histological/morphological observations of seed development in this species, such as expression of genes involved in starch synthesis and transcription factors that have been associated with embryo and endosperm development in other species. We also find evidence for processes that were previously not known to be occurring during seed development in this species, such as epigenetic modification. We then examine the expression of genes associated with patterning DNA and histone methylation—processes that are essential for seed development in distantly related and structurally diverse monocots and eudicots. Around 89% of transcripts putatively homologous to DNA and histone methylation modifiers are expressed during seed development in N. thermarum, including homologs of genes known to pattern imprinting-related epigenetic modifications. Our results suggest that dynamic epigenetic patterning is a deeply conserved aspect of angiosperm seed development.








Similar content being viewed by others
Data availability
Raw sequence data of N. thermarum have been submitted to the National Center for Biotechnology Information (NCBI) database under BioProject PRJNA718528. Genome assembly and annotation (Povilus et al. 2020) is available as part of NCBI BioProject PRJNA508901. Biological material and all other data are available as Supplemental Data, or from the corresponding authors upon request.
References
Amborella Genome Project (2013) The Amborella genome and the evolution of flowering plants. Science 342:6165. https://doi.org/10.1126/science.1241089
Ausin I, Greenberg MVC, Simanshu DK, Hale CJ, Vashinsht AA, Simon SA, Lee T, Feng S, Espánola SD, Meyers BC, Wohlschlegel JA, Patel DJ, Jacobsen SE (2012) INVOLVED IN DE NOVO 2-containing complex involved in RNA-directed DNA methylation in Arabidopsis. PNAS 109(22):8374–8381. https://doi.org/10.1073/pnas.1206638109
Belmonte MF, Kirkbride RC, Stone SL, Pelletier JM, Bui AQ, Yeung EC, Hashimoto M, Fei J, Harada CM, Munoz MD, Le BH, Drews GN, Brady SM, Goldberg RB, Harada JJ (2013) Comprehensive developmental profiles of gene activity in regions and subregions of the Arabidopsis seed. PNAS 110(5):E435–E444. https://doi.org/10.1073/pnas.1222061110
Bewick AJ, Niederhuth CE, Ji L, Rohr NA, Griffin PT, Leebens-Mack J, Schmitz RJ (2017) The evolution of CHROMOMETHYLASES and gene body DNA methylation in plants. Genome Biol 18:65. https://doi.org/10.1186/s13059-017-1195-1
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34:525–527. https://doi.org/10.1038/nbt.3519
Chen J, Zeng M, Xie S, Wang G, Hauck A, Lai J (2014) Dynamic transcriptome landscape of maize embryo and endosperm development. Plant Physiol 166:252–264. https://doi.org/10.1104/pp.114.240689
Desvoyes B, Sanchez MP, Ramirez-Parra E, Gutierrez C (2010) Impact of nucleosome dynamics and istone modifications on cell proliferation during Arabidopsis development. Heredity 105:80–91. https://doi.org/10.1038/hdy.2010.50
Dubos C, Stracke R, Grotewold E, Weisshaae B, Martin C, Lepiniec L (2010) MYB transcription factors in Arabidopsis. Trends Plant Sci 15(10):573–581. https://doi.org/10.1016/j.tplants.2010.06.005
Eddy SR (2011) Accelerated profile HMM searches. PLoS Comput Biol 7:e1002195. https://doi.org/10.1371/journal.pcbi.1002195
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797. https://doi.org/10.1093/nar/gkh340
Fischer E, Magdalena-Rodriguez C (2010) Nymphaea thermarum (Nymphaeaceae). Curtis Botanical Magazine 27:318–327
Furihata HY, Suenaga K, Kawanabe T, Yoshida T, Kawabe A (2016) Gene duplication, silencing, and expression alteration govern the molecular evolution of PRC2 genes in plants. Genes Genet Syst 91(2):85–95. https://doi.org/10.1266/ggs.15-00055
Gao Y, Xu H, Shen Y, Wang J (2013) Transcriptome analysis of rice (Oryza sativa) developing endosperm using the RNA-Seq technique. Plant Mol Biol 81(4–5):363–378. https://doi.org/10.1007/s11103-013-0009-4
Gao J, Yu X, Ma F, Li J (2014) RNA-seq analysis of transcriptome and glucosinolate metabolism in seeds and sprouts of broccoli (Brassica oleracea var. italic). PLoS ONE 9(2): e88804. DOI: https://doi.org/10.1371/journal.pone.0088804
Garg R, Singh VK, Rajkumar MS, Kumar V, Jain M (2017) Global transcriptome and coexpression network analysis reveals cultivar-specific molecular signatures associated with seed development and seed size-weight determination in chickpea. Plant J 91(6):1088–1107. https://doi.org/10.1111/tpj.13621
Gehring M, Satyaki PR (2017) Endosperm and imprinting, inextricably linked. Plant Physiol 173:143–154. https://doi.org/10.1104/pp.16.01353
Girke T, Todd J, Ruuska S, White J, Benning C, Ohirogge J (2000) Microarray analysis of developing Arabidopsis seeds. Plant Physiol 124:1570–1581. https://doi.org/10.1104/pp.124.4.1570
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40(Database issue): D1178–1186. https://doi.org/10.1093/nar/gkr944
Groth M, Moissiard G, Wirtz M, Wang H, Garcia-Salinas C, Ramos-Parra PA, Bischof S, Feng S, Cokus SJ, John A, Smith DC, Zhai J, Hale CJ, Long JA, Hell R, Díaz de la Garza RI, Jacobsen SE (2016) MTHFD1 controls DNA methylation in Arabidopsis. Nat Commun 7:11640. https://doi.org/10.1038/ncomms11640
Haig D (2013) Kin conflict in seed development: an interdependent but fractious collective. Annu Rev Cell Dev Biol 29:189–211. https://doi.org/10.1146/annurev-cellbio-101512-122324
Haig D, Westoby M (1991) Genomic imprinting in endosperm: its effects on seed development in crosses between species and between different ploidies of the same species, and its implications for the evolution of apomixis. Philos Trans R Soc B 333:1–13
Hsieh TF, Shin J, Uzawa R, Silva P, Cohen S, Bauer MJ, Hashimoto M, Kirkbride RD, Harada JJ, Ziberman D, Fischer RL (2011) Regulation of imprinted gene expression in Arabidopsis endosperm. PNAS 108(5):1755–1762. https://doi.org/10.1073/pnas.1019273108
Huang Y, Chen D, Liu BY, Shen WH, Ruan Y (2017) Conservation and diversification of polycomb repressive complex (PRC2) proteins in the green lineage. Brief Funct Genomics 16(2):106–119. https://doi.org/10.1093/bfgp/elw007
Ingouff M, Selles B, Michaud C, Vu TM, Berger F, Schorn AJ, Autran D, Van Durme M, Nowack MK, Marteienssen RA, Grimanelli D (2017) Live-cell analysis of DNA methylation during sexual reproduction in Arabidopsis reveals context and sex-specific dynamics controlled by noncanonical RdDM. Genes & Dev 31:72–83. https://doi.org/10.1101/gad.289397.116
Jin JP, Tian F, Yang DC, Meng YQ, Kong L, Luo JC and Gao G. (2017). PlantTFDB 4.0: toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res., 45(D1): D1040-D1045. https://doi.org/10.1093/nar/gkw982
Jones SI, Vodkin LO (2013) Using RNA-seq to profile soybean seed development from fertilization to maturity. PLoS ONE 8(3):e59270. https://doi.org/10.1371/journal.pone.0059270
Köhler C, Wolff P, Spillane C (2012) Epigenetic mechanisms underlying genomic imprinting in plants. Annu Rev Plant Biol 63:331–352. https://doi.org/10.1146/annurev-arplant-042811-105514
Lagacé M, Matton DP (2004) Characterization of a WRKY transcription factor expressed in late torpedo-stage embryos of Solanum chacoense. Planta 219(1):185–189. https://doi.org/10.1007/s00425-004-1253-2
Lai X, Chahtane H, Martin-Arevalillo R, Parcy ZC, F, (2020) Contrasted evolutionary trajectories of plant transcription factors. Curr Opin Plant Biol 54:101–107. https://doi.org/10.1016/j.pbi.2020.03.002
Lamesch P, Berardini TZ, Li D, Swarbreck D, Wilks C, Sasidharan R, Muller R, Dreher K, Alexander DL, Garcia-Hernandez M, Karthikeyan AS, Lee CH, Nelson WD, Ploetz L, Singh S, Wensel A, Huala E (2012) The Arabidopsis Information Resource (TAIR): improved gene annotation and new tools. Nucleic Acids Res. 40(Database issue): D1202-D1210. https://doi.org/10.1093/nar/gkr1090
Le BH, Cheng C, Bui AQ, Wagmaister JA, Henry KF, Pelletier J, Kwong L, Belmonte M, Kirkbride R, Horvath S, Drews GN, Fischer RL, Okamuro JK, Harada JJ, Goldberg RB (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. PNAS 107(18):8063–8070. https://doi.org/10.1073/pnas.1003530107
Li H, Luan S (2011) The cyclophilin AtCYP71 interacts with CAF-1 and LHP1 and functions in multiple chromatin remodeling processes. Mol Plant 4(4):748–758. https://doi.org/10.1093/mp/ssr036
Li G, Wang D, Yang R, Logan K, Zhang S, Skaggs M, Lloyd A, Burnette WJ, Laurie JD, Hunter BG, Dannenhoffer JM, Larkins BA, Drews GN, Wang X, Yadegari R (2014) Temporal patterns of gene expression in developing maize endosperm identified through transcriptome sequencing. PNAS 111(21):7582–7587. https://doi.org/10.1073/pnas.1406383111
Liu F, Zhang X, Lu C, Li Y, Fu D, Wu G (2015) Non-specific lipid transfer proteins in plants: presenting new advances and an integrated functional analysis. J Exp Bot 66(9):5663–5681. https://doi.org/10.1093/jxb/erv313
Losada JM, Herrero M, Hormaza JI, Friedman WE (2014) Arabinogalactan proteins mark stigmatic receptivity in the protogynous flowers of Magnolia virginiana (Magnoliaceae). Am J Bot 101:1963–1975. https://doi.org/10.3732/ajb.1400280
Luo M, Dennis ES, Berger F, Peacock WJ, Chaudhury A (2005) MINISEED2 (MINI3), a WRKY family gene, and HAIKU2 (IKU2), a leucine-rich repeat (LRR) KINASE gene, are regulators of seed size in Arabidopsis. PNAS 102(48):17531–17536. https://doi.org/10.1073/pnas.0508418102
Montgomery S, Berger F (2021) The evolution of imprinting in plants: beyond the seed. Plant Reprod. https://doi.org/10.1007/s00497-021-00410-7
Nguyen HT, Silva JE, Podicheti R, Macrander J, Yang W, Nazerenus TJ, Nam JW, Jaworski JG, Lu C, Scheffler BE, Mackaitis K, Cahoon EB (2013) Camelina seed transcriptome: a tool for meal and oil improvement and translational research. Plant Biotechnol J 11(6):759–769
Palovaara J, Siaga S, Wendrich JR, van’t Wout Hofland N, van Schayck JP, Hater F, Mutte S, Sjollema J, Boekschoten M, Hooiveld GJ, Weijers D (2017) Transcriptome dynamics revealed by a gene expression atlas of the early Arabidopsis embryo. Nat Plants 3:894–904. https://doi.org/10.1038/s41477-017-0035-3
Pereira AM, Lopes AL, Coimbra S (2016) Arabinogalactan proteins as interactors along the crosstalk between the pollen tube and the female tissues. Front Plant Sci 7:1895. https://doi.org/10.3389/fpls.2016.01895
Picard CL, Povilus RA, Williams BP, Gehring M (2021) Transcriptional and imprinting complexity in Arabidopsis seeds at single-nucleus resolution. Nat Plants 7:730–738. https://doi.org/10.1038/s41477-021-00922-0
Pimentel HJ, Bray N, Puente S, Melsted P, Pachter L (2017) Differential analysis of RNA-Seq incorporating quantification uncertainty. Nat Methods 14:687–690. https://doi.org/10.1038/nmeth.4324
Povilus RA, Losada JM, Friedman WE (2015) Floral biology and ovule and seed ontogeny of Nymphaea thermarum, a water lily at the brink of extinction with potential as a model system for basal angiosperms. Ann Bot 115:211–226. https://doi.org/10.1093/aob/mcu235
Povilus RA, Diggle PK, Friedman WE (2018) Evidence for parent-of-origin effects and interparental conflict in seeds of an ancient flowering plant lineage. Proc R Soc Lond [biol] 285:20172491. https://doi.org/10.1098/rspb.2017.2491
Povilus RA, DaCosta JM, Grassa C, Satyaki PRV, Moeglein M, Jaenisch J, Xi Z, Mathews S, Gehring M, Davis CC, Friedman WE (2020) Water lily (Nymphaea thermarum) genome reveals variable genomic signatures of ancient vascular cambium losses. PNAS 17(15):8649–8656. https://doi.org/10.1073/pnas.1922873117
Qi Z, Zhang Z, Wang Z, Yu J, Qin H, Mao X, Jiang H, Xin D, Yin Z, Zhu R, Liu C, Yu W, Hu Z, Wu X, Liu J, Chen Q (2018) Meta-analysis and transcriptome profiling reveal hub genes for soybean seed storage compoisition during seed development. Plant Cell Environ 41:2109–2127. https://doi.org/10.1111/pce.13175
R Core Team (2017). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/.
Satyaki PRV, Gehring M (2017) DNA methylation and imprinting in plants: machinery and mechanisms. Crit Rev Biochem Mol Biol 52(2):163–175. https://doi.org/10.1080/10409238.2017.1279119
Shu J, Chen C, Li C, Cui Y (2020) The complexity of PRC2 catalysts CLF and SWN in plants. Biochem Soc Trans 48(6):2779–2789. https://doi.org/10.1042/BST20200660
Stamatakis A (2014) RAxML Version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30(9):1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Strejčková B, Čegan R, Pecinka A, Milec Z, Šafář J (2020) Identification of polycomb repressive complex 1 and 2 core components in hexaploid bread wheat. BMC Plant Biol 20:175. https://doi.org/10.1186/s12870-020-02384-6
Swaminathan K, Peterson K, Jack T (2008) The plant B3 superfamily. Trends Plant Sci 13(12):647–655. https://doi.org/10.1016/j.tplants.2008.09.006
Tian T, Liu T, Yan H, You Q, Yi X, Du H, Xu W, Su Z (2017) agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res 45(W1):W122–W129. https://doi.org/10.1093/nar/gkx382
Tirot L, Jullien PE, Ingouff M (2021) Evolution of CG methylation maintenance machinery in plants. Epigenomes 5(19):1–11. https://doi.org/10.3390/epigenomes5030019
Veiseth SV, Rahman MA, Yap KL, Fischer A, Egge-Jacobsen W, Reuter G, Zhou M, Aalen RG, Thorstensen T (2011) The SUVR4 histone lysine methyltransferase binds ubiquitin and converts H3K9me1 to H3K9me3 on transposon chromatin in Arabidopsis. PLoS Genetics 7(3):e1001325. https://doi.org/10.1371/journal.pgen.1001325
Wang G, Wang G, Zhang X, Wang F, Song R (2012) Isolation of high quality RNA from cereal seeds containing high levels of starch. Phytochem Anal 23(2):159–163. https://doi.org/10.1002/pca.1337
Wickramasuriya AM, Dunwell JM (2015) Global scale transcriptome analysis of Arabidopsis embryogenesis in vitro. BMC Genomics 16(1):301. https://doi.org/10.1186/s12864-015-1504-6
Xu H, Gao Y, Wang J (2012) Transcriptome analysis of rice (Oryza sativa) developing embryos using the RNA-Seq technique. PLoS ONE 7(2):e30646. https://doi.org/10.1371/journal.pone.0030646
Yi F, Gu J, Chen J, Song N, Gao X, Zhang X, Zhou Y, Ma X, Song W, Zhao H, Esteban E, Pasha A, Provart N, Lai J (2019) High temporal-resolution transcriptome landscape of early maize seed development. Plant Cell 31(5):974–992. https://doi.org/10.1105/tpc.18.00961
Zemach A, Kim MY, Silva P, Rodrigues JA, Dotson B, Brooks MD, Zilberman D (2010) Local DNA hypomethylation activates genes in rice endosperm. PNAS 107(43):18729–18734. https://doi.org/10.1073/pnas.1009695107
Zhang J, Shan L, Duan J, Wang J, Chen S, Cheng Z, Zhang Q, Liang X, Li Y (2011) De novo assembly and characterization of the transcriptome during seed development, and generation of genic-SSR markers in peanut (Arachis hypogaea L.). BMC Genomics 13:90–96. https://doi.org/10.1186/1471-2164-13-90
Zhou F, Pickersky E (2020) More is better: the diversity of terpene metabolism in plants. Curr Opin Plant Biol 55:1–10. https://doi.org/10.1016/j.pbi.2020.01.005
Ziegler DJ, Khan D, Kalichuk JL, Becker MG, Belmonte MF (2019) Transcriptome landscape of the early Brassica napus seed. J Integr Plant Biol 61:639–650. https://doi.org/10.1111/jipb.12812
Zilberman D, Gehring M, Tran RK, Ballinger T, Henikoff S (2006) Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nature Genet 39:61–69. https://doi.org/10.1038/ng1929
Acknowledgements
We thank the Botanische Gärten der Universität Bonn for providing original plant material for propagation.
Funding
We acknowledge support from the National Science Foundation: IOS-0919986 awarded to W.E.F., and DEB-1500963 and IOS-1812116 awarded to R.A.P.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest or competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
All authors have given consent to publish this work.
Additional information
Communicated by Claudia Köhler.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Povilus, R.A., Friedman, W.E. Transcriptomes across fertilization and seed development in the water lily Nymphaea thermarum (Nymphaeales): evidence for epigenetic patterning during reproduction. Plant Reprod 35, 161–178 (2022). https://doi.org/10.1007/s00497-022-00438-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-022-00438-3