Key message
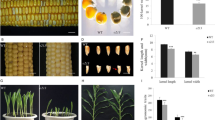
Genes influencing seed size.
Abstract
The designation emp (empty pericarp) refers to a group of defective kernel mutants that exhibit a drastic reduction in endosperm tissue production. They allow the isolation of genes controlling seed development and affecting seed size. Nine independently isolated emp mutants have been analyzed in this study and in all cases longitudinal sections of mature seeds revealed the absence of morphogenesis in the embryo proper, an observation that correlates with their failure to germinate. Complementation tests with the nine emp mutants, crossed inter se in all pairwise combinations, identified complementing and non-complementing pairs in the F1 progenies. Data were then validated in the F2/F3 generations. Mutant chromosomal location was also established. Overall our study has identified two novel emp genes and a novel allele at the previously identified emp4 gene. The introgression of single emp mutants in a different genetic background revealed the existence of a cryptic genetic variation (CGV) recognizable as a variable increase in the endosperm tissue. The unmasking of CGV by introducing single mutants in different genetic backgrounds is the result of the interaction of the emp mutants with a suppressor that has no obvious phenotype of its own and is present in the genetic background of the inbred lines into which the emp mutants were transferred. On the basis of these results, emp mutants could be used as tools for the detection of genetic factors that enhance the amount of endosperm tissue in the maize kernel and which could thus become valuable targets to exploit in future breeding programs.



Similar content being viewed by others
References
Alonso-Blanco C, Mendez-Vigo B, Koornneef M (2005) From phenotypic to molecular polymorphisms involved in naturally occurring variation of plant development. Int J Dev Biol 49:717–732
Arthur K, Vejlupkova Z, Meeley R, Fowler J (2003) Maize ROP2 GTPase provides a competitive advantage to the male gametophyte. Genetics 165:2137–2151
Cheng WH, Taliercio EW, Chourey PS (1996) The Miniature1 seed locus of maize encodes a cell wall invertase required for normal development of endosperm and maternal cells in the pedicel. Plant Cell 8:971–983
Chettoor AM, Yi G, Gomez E, Hueros G, Meeley RB, Becraft PW (2014) A putative plant organelle RNA recognition protein gene is essential for maize kernel development. J Integr Plant Biol 57:236–246
Consonni G, Aspesi C, Barbante A, Dolfini S, Giuliani C, Giulini A, Hansen S, Brettschneider R, Pilu R, Gavazzi G (2003) Analysis of four maize mutants arrested in early embryogenesis reveals an irregular pattern of cell division. Sex Plant Reprod 15:281–290
de Meaux J, Koornneef M (2008) The cause and consequences of natural variation: the genome era takes off! Curr Opin Plant Biol 11:99–102
Flint-Garcia SA, Thuillet AC, Yu J, Pressoir G, Romero SM, Mitchell SE, DoeblyJ Kresovich S, Goodman MM, Buckler ES (2005) Maize association population: a high resolution platform for quantitative trait locus dissection. Plant J 44:1054–1064
Forestan C, Varotto S (2013) Auxin immunolocalization in plant tissues. Methods Mol Biol 959:223–233
Fu S, Scanlon MJ (2004) Clonal analysis of EMPTY PERICARP2 reveals non redundant functions of the duplicated HEAT SHOCK FACTOR BINDING PROTEINs during maize shoot development. Genetics 167:1381–1394
Fu S, Meeley R, Scanlon MJ (2002) empty pericarp2 encodes a negative regulator of the heat shock response and is required for maize embryogenesis. Plant Cell 14:3119–3132
Garcia D, Fitz Gerald JN, Berger F (2005) Maternal control of integument cell elongation and zygotic control of endosperm growth are coordinated to determine seed size in Arabidopsis. Plant Cell 17:52–60
Gibson G, Dworkin I (2004) Uncovering cryptic genetic variation. Nat Rev Genet 5:681–690
Giuliani C, Consonni G, Gavazzi G, Colombo M, Dolfini S (2002) Programmed cell death cells during embryogenesis in Maize. Ann Bot 90:287–292
Gutierrez-Marcos J, Dal Prà M, Giulini A, Costa LM, Gavazzi G, Cordelier S, Olivier S, Tatout C, Wyatt P, Perez P, Dickinson HG, Consonni G (2007) empty pericarp4 encodes a mitochondrion-targeted pentatricopeptide repeat protein necessary for seed development and plant growth in maize. Plant Cell 19:196–210
Gutiérrez-Marcos JF, Costa LM, Biderre-Petit C, Khbaya B, O'Sullivan DM, Wormald M, Perez P, Dickinson HG (2004) Maternally expressed gene1 is a novel maize endosperm transfer cell-specific gene with a maternal parent-of-origin pattern of expression. Plant Cell 16:1288–1301
Jiang W-B, Huang H-Y, Hu Y-W, Zhu S-W, Wang Z-Y et al (2013) Brassinosteroid regulates seed size and shape in Arabidopsis. Plant Physiol 162:19651977
Kang BH, Xiong Y, Williams DS, Pozueta-Romero D, Chourey PS (2009) Miniature1-encoded cell wall invertase is essential for assembly and function of wall-in-growth in the maize endosperm transfer cell. Plant Physiol 151:1366–1376
Keurentjes JJ, Koornneef M, Vreugdenhil D (2008) Quantitative genetics in the age of omics. Curr Opin Plant Biol 11:123–128
Liu YJ, Xiu ZH, Meeley R, Tan BC (2013) Empty pericarp5 encodes a pentatricopeptide repeat protein that is required for mitochondrial RNA editing and seed development in maize. Plant Cell 25:868–883
Liu Y, Wang L, Sun C, Zhang Z, Zheng Y et al (2014) Genetic analysis and major QTL detection for maize kernel size and weight in multienvironments. Theor Appl Genet 127:1019–1037
Liu L, Tong H, Xiao Y, Che R, Xu F et al (2015) Activation of Big Grain1 significantly improves grain size by regulating auxin transport in rice. Proc Natl Acad Sci USA 112:11102–11107
Liu Z, Cook J, Melia-Hancock S, Guill K, Bottoms C et al (2016a) Expanding maize genetic resources with predomestication alleles: Maize–teosinte introgression populations. Plant Genome 9:1–11
Liu Z, Garcia A, Mc Mullen MD, Flint Garcia SA (2016b) Genetic analysis of kernel traits in maize-teosinte introgression populations. G3 Genes Genomes Genetics Early. doi:10.1534/g3.116.030155
Lopato S, Borisjuk N, Langridge P, Hrmova M (2014) Endosperm transfer cell-specific genes and proteins: structure, function and applications in biotechnology. Front Plant Sci 5:64
Maitz M, Santandrea G, Zhang Z, Lal S, Hannah LC, Salamini F, Thompson RD (2000) rgf1, a mutation reducing grain filling in maize through effects on basal endosperm and pedicel development. Plant J 23:29–42
Meyer RS, Purugganan MD (2013) Evolution of crop species: genetics of domestication and diversification. Nat Rev Genet 14:840–852
Neuffer MG, Sheridan WF (1980) Defective kernel mutants of maize. I. Genetic and lethality studies. Genetics 95:929–944
Queitsch C, Sangster TA, Lindquist S (2002) Hsp90 as a capacitor of phenotypic variation. Nature 417:618–624
Sangster TA, Salathia N, Lee HN, Watanabe E, Schellenberg K, Morneau K, Wang H, Undurraga S, Queitsch C, Lindquist S (2008) HSP90-buffered genetic variation is common in Arabidopsis thaliana. Proc Natl Acad Sci USA 105:2969–2974
Scanlon MJ, Stinard PS, James MG, Myers AM, Robertson DS (1994) Genetic analysis of 63 mutations affecting maize kernel development isolated from Mutator stock. Genetics 136:281–294
Sheridan WF, Neuffer MG (1980) Defective kernel mutants of maize. Genetic and lethality studies. Genetics 95:929–944
Sheridan WF, Neuffer MG (1981) Maize mutants altered in embryo development. In: Subtelney S, Abbott U (eds) Levels of genetic control in development, vol 18. Alan R. Liss Inc, New York, pp 137–156
Song X-J, Huang W, Shi M, Zhu M-Z, Lin H-X (2007) A QTL for rice grain width and weight encodes a previously unknown RING-type E3 ubiquitin ligase. Nat Genet 39:623–630
Sosso D, Luo D, Li QB, Sasse J, Yang J, Gendrot G, Suzuki M, Koch KE, McCarty DR, Chourey PS, Rogowsky PM, Ross-Ibarra J, Yang B, Frommer WB (2015) Seed filling in domesticated maize and rice depends on SWEET mediated hexose transport. Nat Genet 47:1489–1493
Sun F, Wang X, Bonnard G, Shen Y, Xiu Z, Li X, Gao D, Zhang Z, Tan BC (2015) Empty pericarp 7 encodes a mitochondrial E-subgroup pentatricopeptide repeat protein that is required for ccmFN editing, mitochondrial function and seed development in maize. Plant J 84(2):283–295
Vernoud V, Hajduch M, Kaled A-S, Depege N, Rogowsky P (2005) Maize embryogenesis. Maydica 50:469–483
Waddington CH (1952) Selection for the basis of an acquired character. Nature 169:278
Waddington CH (1953) Genetic assimilation of an acquired character. Evolution 7:118–126
Weng J, Gu S, Wan X, Gao H, Guo T et al (2008) Isolation and initial characterization of GW5, a major QTL associated with rice grain width and weight. Cell Res 18:1199–1209
Whitt SR, Buckler ES (2003) Using natural allelic diversity to evaluate gene function. Methods Mol Biol 236:123–140
Yu J, Holland JB, McMullen MD, Buckler ES (2008) Genetic design and statistical power of nested association mapping in maize. Genetics 178:539–551
Acknowledgements
We thank the Maize Coop (http://maizecoop.cropsci.uiuc.edu) for kindly providing the genetic stocks used in these study and Serena Varotto for her advice and Marta Dell'Orto for her technical assistance on the histological analysis.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Thomas Dresselhaus.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sangiorgio, S., Carabelli, L., Gabotti, D. et al. A mutational approach for the detection of genetic factors affecting seed size in maize. Plant Reprod 29, 301–310 (2016). https://doi.org/10.1007/s00497-016-0294-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00497-016-0294-6